

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144931.10 + phase: 0 /pseudo
(416 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
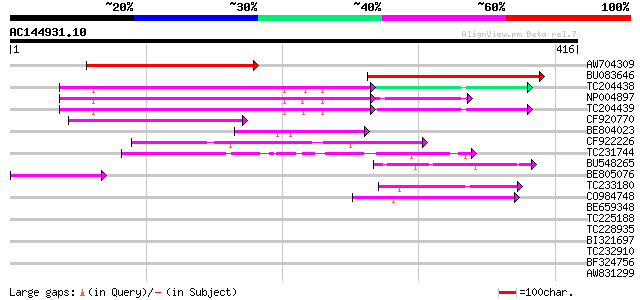
Score E
Sequences producing significant alignments: (bits) Value
AW704309 weakly similar to GP|6642775|gb| gag-pol polyprotein {V... 134 8e-32
BU083646 weakly similar to GP|6642775|gb| gag-pol polyprotein {V... 120 9e-28
TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, co... 96 4e-20
NP004897 gag-protease polyprotein 94 9e-20
TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete 93 3e-19
CF920770 74 9e-14
BE804023 71 1e-12
CF922226 61 1e-09
TC231744 49 4e-06
BU548265 homologue to GP|18000935|gb| cytochrome b {Mabuya delal... 49 5e-06
BE805076 weakly similar to GP|9294121|dbj copia-like retrotransp... 47 1e-05
TC233180 similar to UP|Q944K0 (Q944K0) At1g18030/T10F20_3, parti... 47 2e-05
CO984748 46 4e-05
BE659348 weakly similar to PIR|JC7809|JC78 sulfakinin receptor p... 39 0.004
TC225188 similar to UP|Q9FYB7 (Q9FYB7) Splicing factor-like prot... 35 0.062
TC228935 similar to UP|Q9FHC2 (Q9FHC2) Arabidopsis thaliana geno... 35 0.062
BI321697 35 0.062
TC232910 similar to UP|Q6I8N0 (Q6I8N0) Pol polypeptide, partial ... 30 0.26
BF324756 similar to GP|29423270|gb gag-pol polyprotein {Glycine ... 32 0.53
AW831299 32 0.53
>AW704309 weakly similar to GP|6642775|gb| gag-pol polyprotein {Vitis
vinifera}, partial (23%)
Length = 423
Score = 134 bits (337), Expect = 8e-32
Identities = 64/126 (50%), Positives = 96/126 (75%)
Frame = -1
Query: 57 IFIKILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKETETVREFSDRISK 116
IFI+I+ L+ K W+ L EEY G +R + M+VLNLRREFE +MKE+ET++E+S+++
Sbjct: 387 IFIRIMTLKSPKAIWDCLKEEYAGDDRIRSMQVLNLRREFEL*RMKESETIKEYSNKLLG 208
Query: 117 VITQIRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNFSEITVPELVNALQASEQR 176
++ +I+LLG D +D R+VEK LV +PE +EA I+SLE K+ S+IT+ E+++ALQA EQR
Sbjct: 207 IVNKIKLLGSDFADSRIVEKNLVTVPERYEAFIASLENTKDLSKITLAEVLHALQAQEQR 28
Query: 177 RSLRME 182
R +R +
Sbjct: 27 RLMRQD 10
>BU083646 weakly similar to GP|6642775|gb| gag-pol polyprotein {Vitis
vinifera}, partial (19%)
Length = 428
Score = 120 bits (302), Expect = 9e-28
Identities = 62/130 (47%), Positives = 85/130 (64%)
Frame = +1
Query: 263 QEDEEQLFKASCYLACSSKETWLIDSGCTNHMTNNVSFFKELDESFYSKVVFGNGQHVKV 322
Q EEQLF SC+ A SS +WLIDSGCTNHMT + F ELDE +SKV N ++ V
Sbjct: 10 QSQEEQLFVVSCFAASSST*SWLIDSGCTNHMTYDRELFTELDEVVFSKVKIRNEAYIDV 189
Query: 323 KGKGVVAVETLLGIKYILDVLFVPEINQSLLSVGQMMEKNYSLHFKNMKCTIFDPYGSKL 382
KGK VA+ G+K I +VL+V EI+Q+LLSV Q+++K Y + F++ C I D ++
Sbjct: 190 KGKETVAI*GHTGLKLISNVLYVSEISQNLLSVPQLLKKGYKVLFEDKNCMIKDSESREV 369
Query: 383 MIVEMRGKSF 392
++M+G SF
Sbjct: 370 FNIQMKGMSF 399
>TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, complete
Length = 4734
Score = 95.5 bits (236), Expect = 4e-20
Identities = 68/265 (25%), Positives = 121/265 (45%), Gaps = 33/265 (12%)
Frame = +1
Query: 37 DEVAK-EGRALAIIHAALHDDIFIKILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRRE 95
DE+A +AL + + +IF I +AK+AW L ++G+ + K ++ L +
Sbjct: 220 DELALGNSKALNALFNGVDKNIFRLINTCTVAKDAWEILKTTHEGTSKVKMSRLQLLATK 399
Query: 96 FEALKMKETETVREFSDRISKVITQIRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEEN 155
FE LKMKE E + +F I ++ LGE ++D+++V KIL LP+ F+ K++++EE
Sbjct: 400 FENLKMKEEECIHDFHMNILEIANACTALGERMTDEKLVRKILRSLPKRFDMKVTAIEEA 579
Query: 156 KNFSEITVPELVNALQASEQRRSLRMEENVEG-AFLASNKGKNQSF-------------- 200
++ + V EL+ +LQ E S R E+ + AF+++++G+ +
Sbjct: 580 QDICNMRVDELIGSLQTFELGLSDRTEKKSKNLAFVSNDEGEEDEYDLDTDEGLTNAVVL 759
Query: 201 ------KSFGEKKFPPCPHCK-------KDTHLDKFCWYRP----GVKCRACNQLGHVEK 243
K PH + K + K +P G++C C GH++
Sbjct: 760 LGKQFNKVLNRMDRRQKPHVRNIPFDIRKGSEYQKKSDEKPSHSKGIQCHGCEGYGHIKA 939
Query: 244 VCKNKTNQQEQQARVVEHHQEDEEQ 268
C +Q + V + EQ
Sbjct: 940 ECPTHLKKQRKGLSVCRSDDTESEQ 1014
Score = 71.6 bits (174), Expect = 6e-13
Identities = 75/351 (21%), Positives = 141/351 (39%), Gaps = 4/351 (1%)
Frame = +1
Query: 37 DEVAKEGRALAI-IHAALHDDIFIK--ILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLR 93
DE+A R L I L + +K I NLE KEA + + E +G K+ N+
Sbjct: 1105 DELAISYRELCIKSEKILQQEAQLKKVIANLEAEKEAHEEEISELKGEVGFLNSKLENMT 1284
Query: 94 REFEALKMKETETVREFSDRISKVITQIRLLGEDLSDQRVVEKILVCLPEMFEAKISSLE 153
+ + L ++ S ++ ++ LG+++ +QR +
Sbjct: 1285 KSIKML------------NKGSDMLDEVLQLGKNVGNQRGLGF----------------- 1377
Query: 154 ENKNFSEITVPELVNALQASEQRRSLRMEENVEGAFLASNKGKNQSFKSFGEKKFP-PCP 212
+K+ T+ E V A +N GA ++ ++ ++ + K+ C
Sbjct: 1378 NHKSAGRTTMTEFVPA-------------KNSTGATMSQHRSRHHGTQQKKSKRKKWRCH 1518
Query: 213 HCKKDTHLDKFCWYRPGVKCRACNQLGHVEKVCKNKTNQQEQQARVVEHHQEDEEQLFKA 272
+C K H+ FC++ G + Q R + + +
Sbjct: 1519 YCGKYGHIKPFCYHLHGHP---------------HHGTQSSSSGRKMMWVPKHKIVSLVV 1653
Query: 273 SCYLACSSKETWLIDSGCTNHMTNNVSFFKELDESFYSKVVFGNGQHVKVKGKGVVAVET 332
L S+KE W +DSGC+ HMT F ++ S V FG+G K+ G G + +
Sbjct: 1654 HTSLRASAKEDWYLDSGCSRHMTGVKEFLVNIEPCSTSYVTFGDGSKGKITGMGKLVHD- 1830
Query: 333 LLGIKYILDVLFVPEINQSLLSVGQMMEKNYSLHFKNMKCTIFDPYGSKLM 383
G+ + VL V + +L+S+ Q+ ++ ++++F +C + + LM
Sbjct: 1831 --GLPSLNKVLLVKGLTANLISISQLCDEGFNVNFTKSECLVTNEKSEVLM 1977
>NP004897 gag-protease polyprotein
Length = 1923
Score = 94.4 bits (233), Expect = 9e-20
Identities = 68/265 (25%), Positives = 120/265 (44%), Gaps = 33/265 (12%)
Frame = +1
Query: 37 DEVAK-EGRALAIIHAALHDDIFIKILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRRE 95
DE+A +AL + + +IF I +AK+AW L ++G+ + K ++ L +
Sbjct: 220 DELALGNSKALNALFNGVDKNIFRLINTCTVAKDAWEILKTTHEGTSKVKMSRLQLLATK 399
Query: 96 FEALKMKETETVREFSDRISKVITQIRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEEN 155
FE LKMKE E + +F I ++ LGE ++D+++V KIL LP+ F+ K++++EE
Sbjct: 400 FENLKMKEEECIHDFHMNILEIANACTALGERMTDEKLVRKILRSLPKRFDMKVTAIEEA 579
Query: 156 KNFSEITVPELVNALQASEQRRSLRMEENVEG-AFLASNKGKNQSF-------------- 200
++ + V EL+ +LQ E S R E+ + AF+++++G+ +
Sbjct: 580 QDICNLRVDELIGSLQTFELGLSDRTEKKSKNLAFVSNDEGEEDEYDLDTDEGLTNAVVL 759
Query: 201 ------KSFGEKKFPPCPH-------CKKDTHLDKFCWYRP----GVKCRACNQLGHVEK 243
K PH +K + K +P G +C C GH++
Sbjct: 760 LGKQFNKVLNRMDRRQKPHVRNIPFDIRKGSEYQKRSDEKPSHSKGFQCHGCEGYGHIKA 939
Query: 244 VCKNKTNQQEQQARVVEHHQEDEEQ 268
C +Q + V + EQ
Sbjct: 940 ECPTHLKKQRKGLSVCRSDDTESEQ 1014
Score = 55.8 bits (133), Expect = 3e-08
Identities = 67/307 (21%), Positives = 127/307 (40%), Gaps = 4/307 (1%)
Frame = +1
Query: 37 DEVAKEGRALAI-IHAALHDDIFIK--ILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLR 93
DE+A R L I L + +K I NLE KEA + + E +G K+ N+
Sbjct: 1105 DELATSYRELCIKSEKILQQEAQLKKVIANLEAEKEAHEEEISELKGEVGFLNSKLENMT 1284
Query: 94 REFEALKMKETETVREFSDRISKVITQIRLLGEDLSDQRVVEKILVCLPEMFEAKISSLE 153
+ + L ++ S ++ ++ LG+++ +QR +
Sbjct: 1285 KSIKML------------NKGSDMLDEVLQLGKNVGNQRGLGF----------------- 1377
Query: 154 ENKNFSEITVPELVNALQASEQRRSLRMEENVEGAFLASNKGKNQSFKSFGEKKFP-PCP 212
+K+ IT+ E V A ++ GA ++ ++ ++ + K+ C
Sbjct: 1378 NHKSAGRITMTEFVPAKIST-------------GATMSQHRSRHHGTQQKKSKRKKWRCH 1518
Query: 213 HCKKDTHLDKFCWYRPGVKCRACNQLGHVEKVCKNKTNQQEQQARVVEHHQEDEEQLFKA 272
+C K H+ FC++ G H ++ +++++ V H+ + +
Sbjct: 1519 YCGKYGHIKPFCYHLHG----------HPHHGTQSSSSRRKMMW--VPKHKIVSLVVHTS 1662
Query: 273 SCYLACSSKETWLIDSGCTNHMTNNVSFFKELDESFYSKVVFGNGQHVKVKGKGVVAVET 332
L S+KE W +DSGC+ HMT F ++ S V FG+G K+ G G + +
Sbjct: 1663 ---LRASAKEDWYLDSGCSRHMTGVKEFLVNIEPCSTSYVTFGDGSKGKITGMGKLVHD- 1830
Query: 333 LLGIKYI 339
G++Y+
Sbjct: 1831 --GLRYV 1845
>TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete
Length = 4731
Score = 92.8 bits (229), Expect = 3e-19
Identities = 67/265 (25%), Positives = 121/265 (45%), Gaps = 33/265 (12%)
Frame = +1
Query: 37 DEVAK-EGRALAIIHAALHDDIFIKILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRRE 95
DE+A +AL + + +IF I +AK+AW L ++G+ + K ++ L +
Sbjct: 220 DELALGNSKALNALFNGVDKNIFRLINTCTVAKDAWEILKITHEGTSKVKMSRLQLLATK 399
Query: 96 FEALKMKETETVREFSDRISKVITQIRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEEN 155
FE LKMKE E + +F I ++ LGE ++D+++V KIL LP+ F+ K++++EE
Sbjct: 400 FENLKMKEEECIHDFHMNILEIANACTALGERITDEKLVRKILRSLPKRFDMKVTAIEEA 579
Query: 156 KNFSEITVPELVNALQASEQRRSLRMEENVEG-AFLASNKGKNQSF-------------- 200
++ + V EL+ +LQ E S R E+ + AF+++++G+ +
Sbjct: 580 QDICNMRVDELIGSLQTFELGLSDRAEKKSKNLAFVSNDEGEEDEYDLDTDEGLTNAVVL 759
Query: 201 ------KSFGEKKFPPCPHC-------KKDTHLDKFCWYRP----GVKCRACNQLGHVEK 243
K PH +K + K +P G++C C GH+
Sbjct: 760 LGKQFNKVLNRMDKRQKPHVQNIPFDIRKGSKYQKRSDVKPSHSKGIQCHGCEGYGHIIA 939
Query: 244 VCKNKTNQQEQQARVVEHHQEDEEQ 268
C + + V + E E++
Sbjct: 940 ECPTHLKKHRKGLSVCQSDTESEQE 1014
Score = 72.4 bits (176), Expect = 4e-13
Identities = 78/352 (22%), Positives = 142/352 (40%), Gaps = 5/352 (1%)
Frame = +1
Query: 37 DEVAKEGRALAI-IHAALHDDIFIK--ILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLR 93
DE+A R L I L + +K I +LE KEA + + E +G K+ N+
Sbjct: 1102 DELAASYRKLCIKSEKILQQEAQLKKVIADLEAEKEAHKEEISELKGEVGFLNSKLENMT 1281
Query: 94 REFEALKMKETETVREFSDRISKVITQIRLLGEDLSDQRVVEKILVCLPEMFEAKISSLE 153
+ + L ++ S + ++ LLG++ +QR L
Sbjct: 1282 KSIKML------------NKGSDTLDEVLLLGKNAGNQR------------------GLG 1371
Query: 154 EN-KNFSEITVPELVNALQASEQRRSLRMEENVEGAFLASNKGKNQSFKSFGEKKFP-PC 211
N K+ T+ E V A +N GA ++ ++ ++ + K+ C
Sbjct: 1372 FNPKSAGRTTMTEFVPA-------------KNRTGATMSQHRSRHHGMQQKKSKRKKWRC 1512
Query: 212 PHCKKDTHLDKFCWYRPGVKCRACNQLGHVEKVCKNKTNQQEQQARVVEHHQEDEEQLFK 271
+C K H+ FC++ G +N +++ V +H
Sbjct: 1513 HYCGKYGHIKPFCYHLHGHPHHGTQS-----------SNSRKKMMWVPKHKAVS----LV 1647
Query: 272 ASCYLACSSKETWLIDSGCTNHMTNNVSFFKELDESFYSKVVFGNGQHVKVKGKGVVAVE 331
L S+KE W +DSGC+ HMT F ++ S V FG+G K+ G G + +
Sbjct: 1648 VHTSLRASAKEDWYLDSGCSRHMTGVKEFLLNIEPCSTSYVTFGDGSKGKIIGMGKLVHD 1827
Query: 332 TLLGIKYILDVLFVPEINQSLLSVGQMMEKNYSLHFKNMKCTIFDPYGSKLM 383
G+ + VL V + +L+S+ Q+ ++ ++++F +C + + LM
Sbjct: 1828 ---GLPSLNKVLLVKGLTANLISISQLCDEGFNVNFTKSECLVTNEKSEVLM 1974
>CF920770
Length = 581
Score = 74.3 bits (181), Expect = 9e-14
Identities = 33/131 (25%), Positives = 73/131 (55%)
Frame = -2
Query: 44 RALAIIHAALHDDIFIKILNLEIAKEAWNKLMEEYQGSERTKKMKVLNLRREFEALKMKE 103
+A II +AL D + ++ N + AKE W+ L ++G+ K+ ++ L E+E +M
Sbjct: 415 KAKNIITSALGMDEYFRVSNCKSAKEMWDTLRLTHEGTTDVKRSRINALTHEYELFRMNT 236
Query: 104 TETVREFSDRISKVITQIRLLGEDLSDQRVVEKILVCLPEMFEAKISSLEENKNFSEITV 163
E ++ R + ++ + LG++ ++ ++ K+L CL ++ K++++ E+++ S +++
Sbjct: 235 NENIQSMQKRFTHIVNHLAALGKEFQNEDLINKVLRCLSREWQPKVTAISESRDLSNMSL 56
Query: 164 PELVNALQASE 174
L LQ E
Sbjct: 55 ATLFGKLQEHE 23
>BE804023
Length = 407
Score = 70.9 bits (172), Expect = 1e-12
Identities = 38/116 (32%), Positives = 57/116 (48%), Gaps = 17/116 (14%)
Frame = -3
Query: 166 LVNALQASEQRRSLRMEENVEGAFLASNKG-----KNQSFKSFG------------EKKF 208
+++ LQA + RR +R E +EG A + KN++F G +K +
Sbjct: 405 VIHVLQAQD*RRLMRKESRIEGVLSARYQNATKYKKNKNFDGMGSSSTSTNAKGNWKKFY 226
Query: 209 PPCPHCKKDTHLDKFCWYRPGVKCRACNQLGHVEKVCKNKTNQQEQQARVVEHHQE 264
P C HC+K H C RP KC CNQ+GH +CKN+ + A+VV+ +E
Sbjct: 225 PSC*HCRKKGHPPFRC*KRPDAKCSQCNQMGHEVVICKNRNQSHNEAAKVVDPKEE 58
>CF922226
Length = 667
Score = 60.8 bits (146), Expect = 1e-09
Identities = 49/224 (21%), Positives = 97/224 (42%), Gaps = 7/224 (3%)
Frame = -3
Query: 90 LNLRREFEALKMKETETVREFSDRISKVITQIRLLGEDLSDQRVVEKILVCLPEMFEAKI 149
L ++ + KM E +V E D +K+I + + + D+ +L LP+ +
Sbjct: 641 LYXKQSLYSFKMHEDRSVGEQLDLFNKLILDLENIDVTIDDEDQALLLLCYLPKSY---- 474
Query: 150 SSLEENKNFSE--ITVPELVNALQASEQRRSLRMEENVEGAFLASNKGKNQSFKSFGEKK 207
S +E F +++ E+ AL + E + + G L + + F +KK
Sbjct: 473 SHFKETLLFGRDSVSLDEVQTALNSKELNERKEKKSSASGEGLTARGKTFKKDSEFDKKK 294
Query: 208 FPPCPHCKKDTHLDKFCWYRPGVKCRACNQLGHVEKVCKNK-----TNQQEQQARVVEHH 262
P + ++ K ++C C + GH KVC + +N +++ +
Sbjct: 293 QKPENQKNGEGNIFK-------IRCYHCKKEGHTRKVCPERQKNGGSNNRKKDSGNAAIV 135
Query: 263 QEDEEQLFKASCYLACSSKETWLIDSGCTNHMTNNVSFFKELDE 306
Q+D + +A + + W++DSGC+ HMT N S+F++ +
Sbjct: 134 QDDGYESAEALMVSEKNPETKWIMDSGCSWHMTPNKSWFEQFSD 3
>TC231744
Length = 794
Score = 48.9 bits (115), Expect = 4e-06
Identities = 64/266 (24%), Positives = 112/266 (42%), Gaps = 6/266 (2%)
Frame = +2
Query: 83 RTKKMKVLNLRREFEALKMKETETVREFSDRISKVITQIRLLGEDLSDQRVVEKILVCLP 142
+ +K + NL + ++K K +RE+ IS + ++++ L +L + V +L+ LP
Sbjct: 14 KNEKAETSNLLDKLISMKYKGKGNIREYIMEISNLASKLKSLKLELGEDLFVHLVLISLP 193
Query: 143 EMF-EAKISSLEENKNFSEITVPELVNALQASEQRRSLRMEENVEGAFLASNKGKNQSFK 201
F + K+S + +S + EL++ E+R E+ A NK K + K
Sbjct: 194 AHFGQFKVSYNTQKDKWS---LNELISHCVQEEERL*RDRTES------AHNK-KRKKTK 343
Query: 202 SFGEKKFPPCPHCKKDTHLDKFCWYRPGVKCRACNQLGHVEKVCKNKTNQQEQQARVVEH 261
EK KK ++F C C + H++K C + ++ + +
Sbjct: 344 DVAEKTS*Q----KKQQKDEEFT-------CYFCKKSRHMKKKCPKYAAWRVKKDKFLT- 487
Query: 262 HQEDEEQLFKASCYLACSSKETWLIDSGCTNH--MTNNVSFFKELDESFYSKVVFGNGQH 319
L + LA K+TW +DSG T H MT + L + G+G+
Sbjct: 488 -------LVCSEVNLAFVPKDTWWVDSGATTHISMTMQGCLWSRLPSDDERFIFMGDGKK 646
Query: 320 VKVKGKGVVAVETL---LGIKYILDV 342
V V+ A+ET L I + LD+
Sbjct: 647 VAVE-----AIETFRLQLKIGFYLDL 709
>BU548265 homologue to GP|18000935|gb| cytochrome b {Mabuya delalandii},
partial (5%)
Length = 667
Score = 48.5 bits (114), Expect = 5e-06
Identities = 34/126 (26%), Positives = 65/126 (50%), Gaps = 7/126 (5%)
Frame = -3
Query: 268 QLFKASCYLACSSKETWLIDSGCTNHMTN---NVSFFKELDESFYSKVVFGNGQHVKVKG 324
Q+F S A + ++W DSG ++H+TN N+ +E +++ GNGQ + +
Sbjct: 659 QIFLTSS--AATPSQSWYPDSGASHHVTNMSQNIQQVAPFEEPI--QIIIGNGQGLNINS 492
Query: 325 KGVVAVETLLGIKYIL---DVLFVPEINQSLLSVGQM-MEKNYSLHFKNMKCTIFDPYGS 380
G+ + + ++ L ++LFVP I ++L+ V Q + N F + C +F +
Sbjct: 491 SGLSTFSSPINPQFSLVLSNLLFVPTITKNLIRVSQFCKDNNVYFEFHSYVC-LFKS*DT 315
Query: 381 KLMIVE 386
KL+++E
Sbjct: 314 KLVLLE 297
>BE805076 weakly similar to GP|9294121|dbj copia-like retrotransposable
element {Arabidopsis thaliana}, partial (2%)
Length = 388
Score = 47.4 bits (111), Expect = 1e-05
Identities = 25/71 (35%), Positives = 37/71 (51%)
Frame = +1
Query: 1 MKTYLRAQSLWDVVEKGNNPPPLPNNPTLAQIRNHSDEVAKEGRALAIIHAALHDDIFIK 60
M+TYL +Q LWD+VE+G P + +Q + K + L + A D IF +
Sbjct: 175 METYLSSQDLWDIVEEGFTIPADTSALNASQEKELKKNKQKNSKTLFTLQQAETDPIFPR 354
Query: 61 ILNLEIAKEAW 71
I+ + AKEAW
Sbjct: 355 IMGAKTAKEAW 387
>TC233180 similar to UP|Q944K0 (Q944K0) At1g18030/T10F20_3, partial (11%)
Length = 916
Score = 47.0 bits (110), Expect = 2e-05
Identities = 31/111 (27%), Positives = 64/111 (56%), Gaps = 5/111 (4%)
Frame = +2
Query: 271 KASCYLACSSKETW---LIDSGCTNHMTNNVSFFKELDESFYSK-VVFGNGQHVKVKGKG 326
K+S +L+ ++ T +IDSG T+HMT + S+F ++ ++ NG H+ + G G
Sbjct: 404 KSSSFLSFNASGTENI*IIDSGVTDHMTPHSSYFSSYTFLIGNQHIIVANGSHIPIIGCG 583
Query: 327 VVAVETLLGIKYILDVLFVPEINQSLLSVGQMM-EKNYSLHFKNMKCTIFD 376
+ +++ L ++ +VL+VP+++ +LLS+ ++ + N + F + C D
Sbjct: 584 NIQLQSSL---HLNNVLYVPKLSNNLLSIHKIT*DLNCVVTFFHSHCVF*D 727
>CO984748
Length = 810
Score = 45.8 bits (107), Expect = 4e-05
Identities = 31/129 (24%), Positives = 58/129 (44%), Gaps = 6/129 (4%)
Frame = -2
Query: 252 QEQQARVVEHHQEDEEQLFKASCYLACSS----KETWLIDSGCTNHMT-NNVSFFKELDE 306
Q + + + + ++ YLA ++ + W I+ G T+H+T + + E+
Sbjct: 674 QRSNCQALNNKSQSQQNAQAPQAYLANANPTQQSQNWYINYGATHHVTASPQNLMNEVPT 495
Query: 307 SFYSKVVFGNGQHVKVKGKGVVAVETLLGIKYIL-DVLFVPEINQSLLSVGQMMEKNYSL 365
S +V GNGQ + + G V +K L ++L VP I ++L+SV + N
Sbjct: 494 SGNEQVFLGNGQGLPITGSTVFNSPFASNVKLTLNNLLHVPHITKNLVSVS*FAKDNVFF 315
Query: 366 HFKNMKCTI 374
F + C +
Sbjct: 314 EFHSSHCVV 288
>BE659348 weakly similar to PIR|JC7809|JC78 sulfakinin receptor protein
DSK-R1 - fruit fly (Drosophila melanogaster), partial
(6%)
Length = 770
Score = 38.9 bits (89), Expect = 0.004
Identities = 31/135 (22%), Positives = 60/135 (43%), Gaps = 16/135 (11%)
Frame = -1
Query: 183 ENVEGAFLASNKGK-----NQSFKSFGEKKFPPCPHCKKDTHLDKFCWYRPGVKCRACN- 236
+N+E + + SN+G NQ ++ G P C +CK+ H C+ G ++ N
Sbjct: 479 DNIETSVMVSNRGGQGGRGNQGGRA-GRGGRP*CSYCKRVGHTQDTCYSIHGFPGKSVNI 303
Query: 237 ------QLGHVE----KVCKNKTNQQEQQARVVEHHQEDEEQLFKASCYLACSSKETWLI 286
++ +E + + K ++ Q + V+ H A +++ W+I
Sbjct: 302 SKSETSEIKFLEADYQEYLQLKATKESQTSSVISGHNST------ACISQVGNNQSPWII 141
Query: 287 DSGCTNHMTNNVSFF 301
DSG ++H+ +N S F
Sbjct: 140 DSGASDHIASNSSLF 96
>TC225188 similar to UP|Q9FYB7 (Q9FYB7) Splicing factor-like protein, partial
(62%)
Length = 1131
Score = 35.0 bits (79), Expect = 0.062
Identities = 18/51 (35%), Positives = 24/51 (46%), Gaps = 4/51 (7%)
Frame = +1
Query: 211 CPHCKKDTHLDKFC----WYRPGVKCRACNQLGHVEKVCKNKTNQQEQQAR 257
C +C D H + C W KC C + GH+EK CKN + Q+ R
Sbjct: 364 CFNCGIDGHWARDCKAGDWKN---KCYRCGERGHIEKNCKNSPKKLSQRGR 507
>TC228935 similar to UP|Q9FHC2 (Q9FHC2) Arabidopsis thaliana genomic DNA,
chromosome 5, TAC clone:K24M7, partial (36%)
Length = 1224
Score = 35.0 bits (79), Expect = 0.062
Identities = 19/62 (30%), Positives = 30/62 (47%)
Frame = +1
Query: 194 KGKNQSFKSFGEKKFPPCPHCKKDTHLDKFCWYRPGVKCRACNQLGHVEKVCKNKTNQQE 253
K KN +FK K+ P P +K HL + +PG C C + H+ K+C K ++
Sbjct: 169 KNKNSAFK---RKRPEPKPGSRK-RHLLRVPGMKPGESCFICKAMDHIAKLCPEKAEWEK 336
Query: 254 QQ 255
+
Sbjct: 337 NK 342
Score = 28.1 bits (61), Expect = 7.6
Identities = 17/53 (32%), Positives = 23/53 (43%), Gaps = 6/53 (11%)
Frame = +1
Query: 204 GEKKFPPCPHCKKDTHLDKFCWY---RPGVK---CRACNQLGHVEKVCKNKTN 250
G K C +C ++ H C + G K C CNQ GH+ K C T+
Sbjct: 403 GAKDAKYCYNCGENGHALTQCLHPLQEGGTKFAECFVCNQRGHLSKNCPQNTH 561
>BI321697
Length = 368
Score = 35.0 bits (79), Expect = 0.062
Identities = 21/70 (30%), Positives = 37/70 (52%), Gaps = 1/70 (1%)
Frame = -3
Query: 306 ESFYSKVVFGNGQHVKVKGKGVVAVETLLGIKYILD-VLFVPEINQSLLSVGQMMEKNYS 364
ES V GNG V G+G V +E G +LD V + +I ++L+S ++++ Y
Sbjct: 261 ESSTRTVSMGNGSMAHVLGEGQVKLELSSGNFIVLDGVYHISDIRKNLISTSLLVQQGYK 82
Query: 365 LHFKNMKCTI 374
+ F++ + I
Sbjct: 81 VVFESNRVVI 52
>TC232910 similar to UP|Q6I8N0 (Q6I8N0) Pol polypeptide, partial (3%)
Length = 690
Score = 30.4 bits (67), Expect(2) = 0.26
Identities = 21/54 (38%), Positives = 28/54 (50%)
Frame = +2
Query: 301 FKELDESFYSKVVFGNGQHVKVKGKGVVAVETLLGIKYILDVLFVPEINQSLLS 354
F+ +D+ S V GN + G G V + G LDVLFVP I ++LLS
Sbjct: 215 FRPIDDG--SIVNMGNVATEPILGLGCVNLVFTSGKSLYLDVLFVPGIRKNLLS 370
Score = 21.2 bits (43), Expect(2) = 0.26
Identities = 10/41 (24%), Positives = 20/41 (48%), Gaps = 3/41 (7%)
Frame = +1
Query: 264 EDEEQLFKASCYLACSSKE---TWLIDSGCTNHMTNNVSFF 301
++ +L + + S+K+ W DSG T+H+ + F
Sbjct: 85 KERTRLVQVGLMILKSNKDDDVAWWFDSGATSHVCKDRRLF 207
>BF324756 similar to GP|29423270|gb gag-pol polyprotein {Glycine max},
partial (2%)
Length = 121
Score = 32.0 bits (71), Expect = 0.53
Identities = 13/25 (52%), Positives = 16/25 (64%)
Frame = -1
Query: 276 LACSSKETWLIDSGCTNHMTNNVSF 300
L S+K+ W IDSGC+ HMT F
Sbjct: 91 LRASAKQDWHIDSGCSRHMTGVKEF 17
>AW831299
Length = 334
Score = 32.0 bits (71), Expect = 0.53
Identities = 12/26 (46%), Positives = 17/26 (65%)
Frame = +2
Query: 279 SSKETWLIDSGCTNHMTNNVSFFKEL 304
S ++ W IDSGC+ HMT + S F +
Sbjct: 236 SLRKKWYIDSGCSKHMTGDASNFTHI 313
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.321 0.135 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,951,522
Number of Sequences: 63676
Number of extensions: 268837
Number of successful extensions: 1550
Number of sequences better than 10.0: 52
Number of HSP's better than 10.0 without gapping: 1518
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1545
length of query: 416
length of database: 12,639,632
effective HSP length: 100
effective length of query: 316
effective length of database: 6,272,032
effective search space: 1981962112
effective search space used: 1981962112
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC144931.10


