

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144893.5 - phase: 0 /pseudo
(966 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
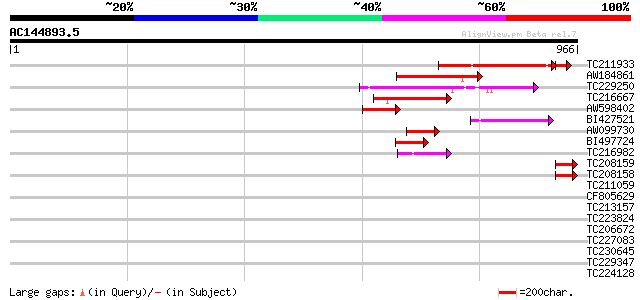
Score E
Sequences producing significant alignments: (bits) Value
TC211933 similar to UP|Q9LNC4 (Q9LNC4) F9P14.9 protein, partial ... 291 5e-83
AW184861 251 8e-67
TC229250 similar to UP|Q9AR00 (Q9AR00) PSTVd RNA-biding protein,... 222 5e-58
TC216667 weakly similar to GB|BAC06968.1|22093674|AP003815 kinas... 130 2e-30
AW598402 121 2e-27
BI427521 similar to GP|13186132|emb PSTVd RNA-biding protein Vi... 118 1e-26
AW099730 weakly similar to GP|29569106|gb| global transcription ... 74 2e-13
BI497724 similar to GP|16323117|gb| AT3g01770/F28J7_10 {Arabidop... 62 9e-10
TC216982 similar to UP|Q9AR19 (Q9AR19) Histone acetyltransferase... 55 1e-07
TC208159 similar to UP|Q39379 (Q39379) Non intermediate filament... 49 1e-05
TC208158 similar to UP|Q39379 (Q39379) Non intermediate filament... 49 1e-05
TC211059 similar to UP|Q8LRK9 (Q8LRK9) HAF1, partial (7%) 41 0.003
CF805629 30 0.032
TC213157 similar to UP|Q8CEM2 (Q8CEM2) Mus musculus adult male t... 37 0.032
TC223824 homologue to UP|Q99491 (Q99491) DAN15 protein (Fragment... 35 0.12
TC206672 weakly similar to UP|Q7UTL3 (Q7UTL3) 3-hydroxyisobutyra... 35 0.16
TC227083 35 0.16
TC230645 weakly similar to UP|Q9EQF1 (Q9EQF1) GABA-A epsilon sub... 33 0.61
TC229347 similar to UP|Q9FZ85 (Q9FZ85) 3-phosphoserine phosphata... 33 0.61
TC224128 weakly similar to PRF|1604369A.0|226743|1604369A sulfat... 29 8.7
>TC211933 similar to UP|Q9LNC4 (Q9LNC4) F9P14.9 protein, partial (15%)
Length = 710
Score = 291 bits (746), Expect(2) = 5e-83
Identities = 147/204 (72%), Positives = 170/204 (83%), Gaps = 3/204 (1%)
Frame = +2
Query: 731 YNPKGQDVHAMAEQLSKIFEDRWAIIESDYNREMRYGMDYGAPSPLSRRVPAFTPPPPLD 790
YNPKGQDVH MAEQLS IFE+RWAIIES+YNREM YG+DYGAPSP+SR+ P F PPP +D
Sbjct: 2 YNPKGQDVHIMAEQLSNIFEERWAIIESNYNREMTYGLDYGAPSPVSRKAPPFRPPP-ID 178
Query: 791 MRRILDRQEPFARTPRSMNNTPSSRTPAPKKPKAKDPNKRDMTYDEKQKLSTSLHSLPVE 850
MRRILDR E + P+ M TPSSRTPAPKKPKAKDP+KRDMTY+EKQKLST L SLP E
Sbjct: 179 MRRILDRSESMTQPPKIMGITPSSRTPAPKKPKAKDPHKRDMTYEEKQKLSTHLQSLPSE 358
Query: 851 KLDAVVQMMRKKNL*LNQQDDEIEVDFDAVDAEILWELDRFVLNYKKSLSKNKRKAE--- 907
KLDA+VQ+++K+N L+Q DDEIEVD D+VD E LWELDRFV NYKKSLSKNKRKAE
Sbjct: 359 KLDAIVQIIKKRNSALSQHDDEIEVDIDSVDTETLWELDRFVTNYKKSLSKNKRKAELAI 538
Query: 908 QARERAEALQNSVQSSRLPISVQM 931
ARERAE QN+ Q S+ P++V++
Sbjct: 539 LARERAE--QNAQQKSQAPVAVEI 604
Score = 36.2 bits (82), Expect(2) = 5e-83
Identities = 17/28 (60%), Positives = 20/28 (70%)
Frame = +3
Query: 930 QMKEMFPHPCLCKGEVKQIIRVDQVVQA 957
QMKEMFPHPC K + + I+ V VVQA
Sbjct: 621 QMKEMFPHPCPSKDKFQWIMGVRPVVQA 704
>AW184861
Length = 448
Score = 251 bits (642), Expect = 8e-67
Identities = 124/150 (82%), Positives = 132/150 (87%), Gaps = 4/150 (2%)
Frame = +2
Query: 660 SSLLEKLMEHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKEFAE 719
SSLLEKLM H++ WVFN+PVDV+ LGLHDYFTIIT+PMDLGTVKTRLNKNWYKSPKEFAE
Sbjct: 2 SSLLEKLMRHKHGWVFNSPVDVETLGLHDYFTIITHPMDLGTVKTRLNKNWYKSPKEFAE 181
Query: 720 DVRLTFHNAMTYNPKGQDVHAMAEQLSKIFEDRWAIIESDYNREMRYGMDY----GAPSP 775
DVRLTF NAMTYNP+GQDVH MAE LSKIFEDRWAIIESDYNREMRYG DY APSP
Sbjct: 182 DVRLTFRNAMTYNPQGQDVHIMAELLSKIFEDRWAIIESDYNREMRYGFDYRAAPPAPSP 361
Query: 776 LSRRVPAFTPPPPLDMRRILDRQEPFARTP 805
LSRRV AFT PPPLDMRRILDR + +TP
Sbjct: 362 LSRRVSAFT-PPPLDMRRILDRSDSMTQTP 448
>TC229250 similar to UP|Q9AR00 (Q9AR00) PSTVd RNA-biding protein, Virp1
(PSTVd RNA-binding protein Virp1a) (PSTVd RNA-binding
protein Virp1b) (PSTVd RNA-binding protein Virp1c)
(PSTVd RNA-binding protein Virp1d), partial (34%)
Length = 1450
Score = 222 bits (566), Expect = 5e-58
Identities = 134/329 (40%), Positives = 180/329 (53%), Gaps = 24/329 (7%)
Frame = +2
Query: 597 EGVEKEKRMPKANQFYHNSEFLLANDKFPPAESNKKSKLNWKKQGGGDMGXGLQMGSKFF 656
E ++K + +++F + L + K P + +K D+
Sbjct: 485 EQIQKLRNQIGSSEF-QPGQSLNGHPKKPSGKKISGNKRPLPSNSAKDLKRSHSEVGNLM 661
Query: 657 KSCSSLLEKLMEHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKE 716
K CS +L+KL++H++ WVF PVDV GL LHDY II PMDLGTVK+ L+KN Y +P +
Sbjct: 662 KCCSQVLQKLIKHKHGWVFKAPVDVVGLKLHDYCDIIKQPMDLGTVKSNLSKNVYATPAD 841
Query: 717 FAEDVRLTFHNAMTYNPKGQDVHAMAEQLSKIFEDRW---------AIIESDYNREMRYG 767
FA DVRLTF+NA+ YNPKG DV+ MAEQL FE+ + +I+ + E
Sbjct: 842 FASDVRLTFNNALAYNPKGHDVYTMAEQLLARFEELYRPVHEKFEGSIVHDRESEEELQA 1021
Query: 768 MDYGAPSPLSRRVPAFTPPPPLDMRRILDRQEPFARTPRSMNNTP----SSRTPAP---- 819
+ P R P PP + + EP + P S +N P RTP+P
Sbjct: 1022SSWSQVEP-ERVKKKENPIPPAKLHK-----EPPPQHPASSSNPPLVQSPVRTPSPMRAP 1183
Query: 820 -------KKPKAKDPNKRDMTYDEKQKLSTSLHSLPVEKLDAVVQMMRKKNL*LNQQDDE 872
KPKAKDPNKRDM+ +EK KL L SLP EK++ VVQ++R++N L Q DE
Sbjct: 1184PVKPLKQPKPKAKDPNKRDMSLEEKHKLGLGLQSLPPEKMEQVVQIIRRRNGHLKQDGDE 1363
Query: 873 IEVDFDAVDAEILWELDRFVLNYKKSLSK 901
IE+ +AVD E LWELDR V NYKK +SK
Sbjct: 1364IELHIEAVDTETLWELDRLVTNYKKMVSK 1450
>TC216667 weakly similar to GB|BAC06968.1|22093674|AP003815 kinase-like
protein {Oryza sativa (japonica cultivar-group);} ,
partial (21%)
Length = 892
Score = 130 bits (328), Expect = 2e-30
Identities = 68/139 (48%), Positives = 89/139 (63%), Gaps = 6/139 (4%)
Frame = +3
Query: 621 NDKFPPAESNKKSK-LNWKKQGG-----GDMGXGLQMGSKFFKSCSSLLEKLMEHQYAWV 674
NDK SN+ S LN K+G G ++ K C+++L+ LM H Y+WV
Sbjct: 366 NDKNRRCNSNENSSALNSNKRGPPVSVEGQKEKRQKIDRKGSMQCATILKSLMSHTYSWV 545
Query: 675 FNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKEFAEDVRLTFHNAMTYNPK 734
F+ PVD L + DYFTII++PMDLGT+K++L KN Y +EFA DVRLTF NAM YNP
Sbjct: 546 FSKPVDPIALSIPDYFTIISHPMDLGTIKSKLEKNIYSGTEEFAADVRLTFSNAMKYNPP 725
Query: 735 GQDVHAMAEQLSKIFEDRW 753
DVH MA++LSKIF+ +W
Sbjct: 726 SNDVHLMAKELSKIFDRKW 782
>AW598402
Length = 202
Score = 121 bits (303), Expect = 2e-27
Identities = 58/66 (87%), Positives = 59/66 (88%)
Frame = +3
Query: 601 KEKRMPKANQFYHNSEFLLANDKFPPAESNKKSKLNWKKQGGGDMGXGLQMGSKFFKSCS 660
KEKR PKANQFY NSEFLLA DKFP AESNKKSKLNWKKQGGG+MG G MGSKFFKSCS
Sbjct: 3 KEKRTPKANQFYRNSEFLLAKDKFPSAESNKKSKLNWKKQGGGEMGHGFGMGSKFFKSCS 182
Query: 661 SLLEKL 666
SLLEKL
Sbjct: 183 SLLEKL 200
>BI427521 similar to GP|13186132|emb PSTVd RNA-biding protein Virp1
{Lycopersicon esculentum}, partial (15%)
Length = 421
Score = 118 bits (295), Expect = 1e-26
Identities = 68/142 (47%), Positives = 84/142 (58%)
Frame = +1
Query: 785 PPPPLDMRRILDRQEPFARTPRSMNNTPSSRTPAPKKPKAKDPNKRDMTYDEKQKLSTSL 844
P PP Q P RTP M P PK PKAKDPNKRDM+ +EK KL L
Sbjct: 1 PQPPASSSNPPLLQSP-VRTPSPMRAPPVKPLKQPK-PKAKDPNKRDMSLEEKHKLGLGL 174
Query: 845 HSLPVEKLDAVVQMMRKKNL*LNQQDDEIEVDFDAVDAEILWELDRFVLNYKKSLSKNKR 904
SLP EK++ VVQ++R++N L Q DEIE+D +AVD E LWELDR V NYKK +SK KR
Sbjct: 175 QSLPPEKMEQVVQIIRRRNGHLKQDGDEIELDIEAVDTETLWELDRLVTNYKKMVSKIKR 354
Query: 905 KAEQARERAEALQNSVQSSRLP 926
+A + +Q + + LP
Sbjct: 355 QALMGNIYNDNVQANKGNEELP 420
>AW099730 weakly similar to GP|29569106|gb| global transcription factor group
E {Zea mays}, partial (8%)
Length = 177
Score = 74.3 bits (181), Expect = 2e-13
Identities = 34/56 (60%), Positives = 42/56 (74%)
Frame = +2
Query: 676 NTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPKEFAEDVRLTFHNAMTY 731
N PVDV GL L DY+ +I PMDLGTVK+ L+ N Y +P +FA DVRLTF+NA+ Y
Sbjct: 8 NAPVDVVGLQLTDYYDVIKQPMDLGTVKSNLSMNKYTTPSDFASDVRLTFNNALAY 175
>BI497724 similar to GP|16323117|gb| AT3g01770/F28J7_10 {Arabidopsis
thaliana}, partial (10%)
Length = 408
Score = 62.4 bits (150), Expect = 9e-10
Identities = 29/57 (50%), Positives = 40/57 (69%)
Frame = +1
Query: 657 KSCSSLLEKLMEHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKS 713
K C +LL+++M HQ+ VF+ PVD+ + DYFTII +PMDLGTVK++L Y S
Sbjct: 229 KQCETLLKRVMSHQFGKVFDKPVDIVKWNIPDYFTIIKHPMDLGTVKSKLISCEYTS 399
>TC216982 similar to UP|Q9AR19 (Q9AR19) Histone acetyltransferase GCN5
(At3g54610) (Expressed protein), partial (53%)
Length = 1203
Score = 55.5 bits (132), Expect = 1e-07
Identities = 32/93 (34%), Positives = 50/93 (53%), Gaps = 1/93 (1%)
Frame = +1
Query: 661 SLLEKLMEHQYAWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRL-NKNWYKSPKEFAE 719
SLL+ + +H AW F PVD + DY+ II +PMDL T+ R+ ++ +Y + + F
Sbjct: 595 SLLKSMFDHADAWPFKEPVDARDVP--DYYDIIKDPMDLKTMSKRVDSEQYYVTFEMFVA 768
Query: 720 DVRLTFHNAMTYNPKGQDVHAMAEQLSKIFEDR 752
D R F NA TYN + + +L F+ +
Sbjct: 769 DARRMFANARTYNSPETIYYKCSTRLEAHFQSK 867
>TC208159 similar to UP|Q39379 (Q39379) Non intermediate filament IFA binding
protein (Fragment), partial (19%)
Length = 652
Score = 48.9 bits (115), Expect = 1e-05
Identities = 25/40 (62%), Positives = 31/40 (77%), Gaps = 3/40 (7%)
Frame = +1
Query: 930 QMKEMFPHPCLCKGEVKQIIRVDQ---VVQAVILGLLQVV 966
QMK +FP+PCLCKGE+K+I+ V Q V AVIL LLQV+
Sbjct: 70 QMKGVFPNPCLCKGEIKRIMGVGQAVPVAPAVILVLLQVI 189
>TC208158 similar to UP|Q39379 (Q39379) Non intermediate filament IFA binding
protein (Fragment), partial (17%)
Length = 758
Score = 48.9 bits (115), Expect = 1e-05
Identities = 26/40 (65%), Positives = 31/40 (77%), Gaps = 3/40 (7%)
Frame = +2
Query: 930 QMKEMFPHPCLCKGEVKQIIRVDQVVQ---AVILGLLQVV 966
QMK +FP+PCL KGE+KQI+ V Q VQ AVIL LLQV+
Sbjct: 128 QMKGIFPNPCLRKGEIKQIMGVGQAVQAAPAVILDLLQVI 247
Score = 42.0 bits (97), Expect = 0.001
Identities = 27/57 (47%), Positives = 34/57 (59%), Gaps = 9/57 (15%)
Frame = +3
Query: 897 KSLSKNKRKAEQARERAEAL-QNSVQSSRLPISVQM--------KEMFPHPCLCKGE 944
KSLSKNKRKAE A+ RAEAL QN++Q S+ P ++ KE P KG+
Sbjct: 3 KSLSKNKRKAELAQARAEALQQNAIQKSQAPAMAEIPKGNTNR*KESSPTLACAKGK 173
>TC211059 similar to UP|Q8LRK9 (Q8LRK9) HAF1, partial (7%)
Length = 800
Score = 40.8 bits (94), Expect = 0.003
Identities = 34/137 (24%), Positives = 60/137 (42%), Gaps = 6/137 (4%)
Frame = +2
Query: 602 EKRMPKANQFYHNSEFL----LANDKFPPAESNKKSKLNWKKQGGGDMGXGLQMGSKFFK 657
+KRMP+ ++ + D PP + ++ GGG++G
Sbjct: 101 DKRMPERDRSGKRRSITELGKIGADYMPPTKR--------RRGGGGEVGLA--------N 232
Query: 658 SCSSLLEKLMEHQY--AWVFNTPVDVDGLGLHDYFTIITNPMDLGTVKTRLNKNWYKSPK 715
S+++ +++ +Y +++F PV DY II PMDL ++ R+ YKS +
Sbjct: 233 ILESVVDTIVKDRYDLSYLFLKPVSKKEAP--DYLDIIERPMDLSRIRERVRNMEYKSRE 406
Query: 716 EFAEDVRLTFHNAMTYN 732
+F D+ NA YN
Sbjct: 407 DFRHDMWQITFNAHKYN 457
>CF805629
Length = 556
Score = 30.0 bits (66), Expect(2) = 0.032
Identities = 13/22 (59%), Positives = 14/22 (63%)
Frame = +1
Query: 711 YKSPKEFAEDVRLTFHNAMTYN 732
YK +E DVRL F NAM YN
Sbjct: 52 YKHVREICADVRLVFKNAMKYN 117
Score = 26.2 bits (56), Expect(2) = 0.032
Identities = 27/128 (21%), Positives = 53/128 (41%), Gaps = 1/128 (0%)
Frame = +2
Query: 737 DVHAMAEQLSKIFEDRWAIIESDYNREMRYGMDYGAPSPLSRRVPAFTPPPPLDMRRILD 796
DVH MA+ L FE++W + E + A + L+ +V
Sbjct: 131 DVHVMAKTLLSKFEEKWLQLLPKVTEEETRREEEEAEAQLALQVA--------------- 265
Query: 797 RQEPFARTPRSMNNTPSSRTPAPKKPKAKDPNK-RDMTYDEKQKLSTSLHSLPVEKLDAV 855
++ A+ R ++N ++ + + R M+ +EK+KL +L L E L
Sbjct: 266 QEAAQAKMARDLSNELYEVDVILEELREMVVKRFRKMSTEEKRKLGDALTRLSPEDLSKA 445
Query: 856 VQMMRKKN 863
++++ + N
Sbjct: 446 LEIVAQNN 469
>TC213157 similar to UP|Q8CEM2 (Q8CEM2) Mus musculus adult male testis cDNA,
RIKEN full-length enriched library, clone:1700123E07
product:protamine 1, full insert sequence, partial (32%)
Length = 465
Score = 37.4 bits (85), Expect = 0.032
Identities = 28/87 (32%), Positives = 42/87 (48%), Gaps = 4/87 (4%)
Frame = +3
Query: 740 AMAEQLSKIFEDRWAIIESDYNREMRYGMDYGAPSPLSRRVPAFTPPPPLDMRRILDRQE 799
A A++L+++ + A + D + E G Y AP PLS + P P R +++
Sbjct: 141 AAAQRLAQVMASQTA--DDDDDDEDDLGFRYTAPPPLSLSRSSARSPSPALARNLVEESS 314
Query: 800 PFAR--TPRSMNNTP--SSRTPAPKKP 822
+ R P + NN P S RTPAP P
Sbjct: 315 MYLRPAPPAAGNNRPPLSLRTPAPVLP 395
>TC223824 homologue to UP|Q99491 (Q99491) DAN15 protein (Fragment), partial
(6%)
Length = 439
Score = 35.4 bits (80), Expect = 0.12
Identities = 26/85 (30%), Positives = 39/85 (45%), Gaps = 2/85 (2%)
Frame = -1
Query: 740 AMAEQLSKIFEDRWAIIESDYNR-EMRYGMDYGAPSPLSRRVPAFTPPPPLDMRRILDRQ 798
A A++L+++ + A+ +D + E G Y AP PLS + P P R +++
Sbjct: 262 AAAQRLAQVMASQTAVAAADDDDDEDDLGFRYTAPPPLSLSRGSAKSPSPALARNLVEES 83
Query: 799 EPFARTPRSMNNTP-SSRTPAPKKP 822
P N P S RTPAP P
Sbjct: 82 MYLRPAPTPGNRPPLSLRTPAPVPP 8
>TC206672 weakly similar to UP|Q7UTL3 (Q7UTL3) 3-hydroxyisobutyrate
dehydrogenase , partial (76%)
Length = 1230
Score = 35.0 bits (79), Expect = 0.16
Identities = 19/52 (36%), Positives = 25/52 (47%)
Frame = +2
Query: 773 PSPLSRRVPAFTPPPPLDMRRILDRQEPFARTPRSMNNTPSSRTPAPKKPKA 824
PSP S P +PPPP + P + TPR+ + S+ PAP P A
Sbjct: 191 PSPSSTSGPT-SPPPPTPSQPAPTSSSPSSATPRTSAQSSSTPPPAPSPPSA 343
>TC227083
Length = 1777
Score = 35.0 bits (79), Expect = 0.16
Identities = 21/75 (28%), Positives = 35/75 (46%)
Frame = +2
Query: 291 EDQNLAQTHVSSRTEDVNSPRPQNSTQKDGDSPQILENSRPEDASLTQPEVNSRLEDRSF 350
E ++ VSS ++ + P QN DG P + + + + + + +DRSF
Sbjct: 395 EGHPFGESKVSSNSKPITQPWHQNGRCPDGTIP-VRRTKKDDMLRASSVQHFGKKKDRSF 571
Query: 351 PQPELNSKLEDMTSQ 365
PQP+ L D+ SQ
Sbjct: 572 PQPKPAKPLPDIISQ 616
>TC230645 weakly similar to UP|Q9EQF1 (Q9EQF1) GABA-A epsilon subunit splice
variant (Fragment), partial (8%)
Length = 464
Score = 33.1 bits (74), Expect = 0.61
Identities = 28/116 (24%), Positives = 51/116 (43%), Gaps = 6/116 (5%)
Frame = +2
Query: 274 PPVSSRTD---DGDSAQQQVEDQNLAQTHVSSRTEDVNSPRPQNSTQKDGDSPQILENSR 330
P S+R+ D D ++ + Q+ + S ++ P QN ++D P+ +
Sbjct: 5 PSCSTRSSLLLDSDRSKALLCPQSSVSLSLKS*HSEMPEPESQNGHEQD---PEPQPETE 175
Query: 331 PEDASLTQPEVNSRLEDRSFPQPELNSKLED---MTSQQQDNSILEDGNSSQPQLN 383
P TQP++ + +S P+PE N + E T Q Q + G+ + P +N
Sbjct: 176 PVPTEQTQPQLEPK--SKSTPEPEPNPQPESEPVPTEQTQAQLEPKSGSEADPAVN 337
>TC229347 similar to UP|Q9FZ85 (Q9FZ85) 3-phosphoserine phosphatase, partial
(73%)
Length = 1278
Score = 33.1 bits (74), Expect = 0.61
Identities = 24/83 (28%), Positives = 29/83 (34%), Gaps = 2/83 (2%)
Frame = -2
Query: 410 QSVSDDLHINQQAEPSNLNVRQEDDRPSSPIHRQGAISDN--LHSHQQVEPSNPNVLLED 467
Q V LH + Q P + P+ + G S N L SHQQ +P V
Sbjct: 434 QQVHQSLHPSTQCYPHQSTQHHHFSKAQEPLWK-GVYSQNGPLWSHQQKQPQRNTVSSSS 258
Query: 468 DGPSSPIHRHKAVPSTEYRHSEN 490
P IH H E EN
Sbjct: 257 SQPQFQIHTHCYSSDAELMKEEN 189
>TC224128 weakly similar to PRF|1604369A.0|226743|1604369A sulfated surface
glycoprotein SSG185. {Volvox carteri;} , partial (9%)
Length = 633
Score = 29.3 bits (64), Expect = 8.7
Identities = 19/59 (32%), Positives = 23/59 (38%)
Frame = -1
Query: 771 GAPSPLSRRVPAFTPPPPLDMRRILDRQEPFARTPRSMNNTPSSRTPAPKKPKAKDPNK 829
G P P R P TPP P Q P + P + PS TP K + P+K
Sbjct: 183 GKPPPKQRNYP-HTPPTPPHTPSSSSPQSPPSPXPXPASPPPSPATP*AKTSPSPPPHK 10
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.310 0.128 0.357
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 36,323,778
Number of Sequences: 63676
Number of extensions: 510415
Number of successful extensions: 3288
Number of sequences better than 10.0: 100
Number of HSP's better than 10.0 without gapping: 3050
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3248
length of query: 966
length of database: 12,639,632
effective HSP length: 106
effective length of query: 860
effective length of database: 5,889,976
effective search space: 5065379360
effective search space used: 5065379360
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 64 (29.3 bits)
Medicago: description of AC144893.5


