

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
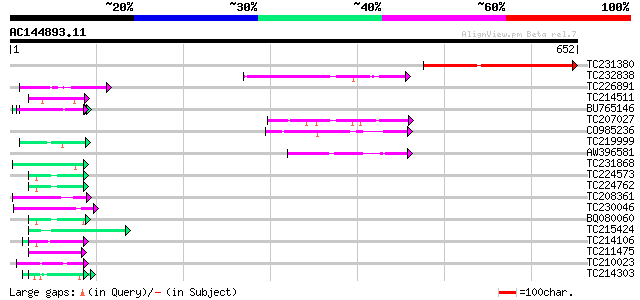
Query= AC144893.11 + phase: 0 /pseudo/partial
(652 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC231380 263 1e-70
TC232838 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {A... 54 2e-07
TC226891 similar to GB|AAM61419.1|21537078|AY084855 putaive DNA-... 53 5e-07
TC214511 weakly similar to PIR|C85356|C85356 glycine-rich protei... 51 1e-06
BU765146 similar to GP|21322711|e pherophorin-dz1 protein {Volvo... 50 3e-06
TC207027 weakly similar to UP|Q9LW95 (Q9LW95) KED, partial (17%) 49 7e-06
CO985236 49 9e-06
TC219999 weakly similar to UP|Q7T9Y7 (Q7T9Y7) Odv-e66, partial (5%) 48 1e-05
AW396581 47 3e-05
TC231868 similar to UP|Q93424 (Q93424) C. elegans GRL-23 protein... 47 3e-05
TC224573 homologue to UP|Q41707 (Q41707) Extensin class 1 protei... 46 5e-05
TC224762 homologue to UP|Q09085 (Q09085) Hydroxyproline-rich gly... 45 8e-05
TC208361 weakly similar to UP|O64835 (O64835) Expressed protein ... 44 2e-04
TC230046 similar to UP|Q9LU79 (Q9LU79) Gb|AAF21150.1, partial (28%) 44 2e-04
BQ080060 homologue to PIR|C29356|C29 hydroxyproline-rich glycopr... 43 4e-04
TC215424 similar to UP|Q9MA04 (Q9MA04) F20B17.16, partial (26%) 43 4e-04
TC214106 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 pr... 43 5e-04
TC211475 similar to UP|P79122 (P79122) Pinin, partial (5%) 43 5e-04
TC210023 weakly similar to UP|Q868B4 (Q868B4) C. elegans GRL-25 ... 43 5e-04
TC214303 similar to UP|Q9T0K5 (Q9T0K5) Extensin-like protein, pa... 42 7e-04
>TC231380
Length = 1013
Score = 263 bits (673), Expect = 1e-70
Identities = 129/177 (72%), Positives = 151/177 (84%), Gaps = 1/177 (0%)
Frame = +1
Query: 477 MPSGTGSSIQAMEAVQMPFGSGIHLHDSSVGDFLSARDDPQMIHGSSLFGNGHKRD-IGL 535
M SG G IQ MEA QMPFGSGI L D+ VGDFLS+RD+PQMI GSSLFGNGHKRD +GL
Sbjct: 1 MSSGNGGLIQVMEAGQMPFGSGIDLRDNPVGDFLSSRDEPQMISGSSLFGNGHKRDNLGL 180
Query: 536 DNHNSHHTLNGSNKRLRSDSPWSSQPIDFEGCMEQMQHFMEKARMMYASKDQAAEESAAN 595
DNH H+LNGSNKRLR DSPW+S+P+DFE C+EQM+H+M KARMM+A+KDQA EES N
Sbjct: 181 DNH---HSLNGSNKRLRGDSPWNSKPMDFETCIEQMEHWMGKARMMFATKDQACEESTMN 351
Query: 596 QQVLLNELQRRDEMVAHLQKARIDQTQKTQIEVYRLEKELYMMQSLVEGYRKALKET 652
QQ+L+NELQ+RD M+ HL KA++++ QK QIEVYR EKELYMMQSLV+GYRKALKET
Sbjct: 352 QQLLINELQKRDNMIEHLHKAKLEEAQKRQIEVYRFEKELYMMQSLVDGYRKALKET 522
>TC232838 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {Arabidopsis
thaliana;} , partial (46%)
Length = 902
Score = 53.9 bits (128), Expect = 2e-07
Identities = 50/204 (24%), Positives = 96/204 (46%), Gaps = 11/204 (5%)
Frame = +2
Query: 269 LMWSMLEKELKAT-HLEECYYASHLQHLIKSQHKELFEETLVVEVEGEGEEGVVKDEEEE 327
++W L+ ++KAT L E + ++ L++ +E VV+ + E+ D+++E
Sbjct: 197 VLW-FLDSKMKATKELSEQVEENEVKVLVEEDDEEYESVDEVVDDAEDDEDEEEDDDDDE 373
Query: 328 GEAKDEEEEE--DGVVVKDEVDG--SGDVKMGGVEENQVQELEEHNIELSLGQDKVETLP 383
G+ D+EE++ DG D+ D GDV+ GG ++ + ++ + D+ E
Sbjct: 374 GDDNDDEEDDAPDGGDDDDDEDDEEEGDVQRGGEPDDDDNDSDDDS-----DDDEDEDEE 538
Query: 384 VEKEQGEGEQM------MDFEQSKKEETEMWFLGQKNYVGEPSLRPCHNRDRKGIDCEQV 437
E+EQGE E + E +++EE F ++N GE D DCE+
Sbjct: 539 DEEEQGEEEDLGTEYLIRPLETAEEEEASSDFEPEEN--GEE-----EEEDVDDEDCEKA 697
Query: 438 KEDEGEEEEHEQEEEEEDDVEEDE 461
+ + + + +++D E+DE
Sbjct: 698 EAPPKRKRSDKDDSDDDDGGEDDE 769
Score = 33.5 bits (75), Expect = 0.31
Identities = 19/74 (25%), Positives = 33/74 (43%)
Frame = +2
Query: 392 EQMMDFEQSKKEETEMWFLGQKNYVGEPSLRPCHNRDRKGIDCEQVKEDEGEEEEHEQEE 451
E +S+ E +WFL K + + K + E +E E +E + E
Sbjct: 155 EAAQRISKSRGERNVLWFLDSKMKATKELSEQVEENEVKVLVEEDDEEYESVDEVVDDAE 334
Query: 452 EEEDDVEEDEHDVG 465
++ED+ E+D+ D G
Sbjct: 335 DDEDEEEDDDDDEG 376
Score = 30.4 bits (67), Expect = 2.6
Identities = 25/116 (21%), Positives = 54/116 (46%), Gaps = 3/116 (2%)
Frame = +2
Query: 351 DVKMGGVEENQVQELEEHNIELSLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFL 410
D KM +E +++EE+ +++ + +D E V++ + E D E+ ++
Sbjct: 212 DSKMKATKELS-EQVEENEVKVLVEEDDEEYESVDEVVDDAEDDEDEEEDDDDDE----- 373
Query: 411 GQKNYVGEPSLRPCHNRDRKGIDCEQVKEDEG---EEEEHEQEEEEEDDVEEDEHD 463
G N E + D D E+ G ++++++ +++ +DD +EDE D
Sbjct: 374 GDDNDDEEDDAPDGGDDDDDEDDEEEGDVQRGGEPDDDDNDSDDDSDDDEDEDEED 541
>TC226891 similar to GB|AAM61419.1|21537078|AY084855 putaive DNA-binding
protein {Arabidopsis thaliana;} , partial (41%)
Length = 1187
Score = 52.8 bits (125), Expect = 5e-07
Identities = 35/108 (32%), Positives = 50/108 (45%), Gaps = 2/108 (1%)
Frame = +3
Query: 12 TLNPITTEQCPEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLMD 71
T NP T + PPPPPPP P P S T + + ++P P P+H +
Sbjct: 393 TQNPATKTRL*PSPPPPPPPSRPPSPTTPASPTARS-ALSSLPPP----PSH------LS 539
Query: 72 EDPTHPQIEDDPEPEPPSPAATTVRRGHK--RKKFGGKRTAAQKRKSH 117
PT P PP+P AT R+G+ ++FG +R A + + SH
Sbjct: 540 PPPTGKSAATLPPHSPPNPPATAPRKGNHA*NERFGCRR*AQRSQISH 683
>TC214511 weakly similar to PIR|C85356|C85356 glycine-rich protein [imported]
- Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(88%)
Length = 1085
Score = 51.2 bits (121), Expect = 1e-06
Identities = 30/78 (38%), Positives = 38/78 (48%), Gaps = 8/78 (10%)
Frame = -3
Query: 22 PEQPPPPPPPPPNPQ-------PDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLMDEDP 74
PE PPPPPPPPP P PDP + P + +P P + ED +E
Sbjct: 765 PEPPPPPPPPPPPPPPPLP*LLPDP---DPAREPAYEPASEPAPEPPEPELED---EEAE 604
Query: 75 THPQIEDDPEPEP-PSPA 91
++E +PEPEP P PA
Sbjct: 603 LEAELELEPEPEPEPDPA 550
Score = 48.5 bits (114), Expect = 9e-06
Identities = 30/76 (39%), Positives = 37/76 (48%), Gaps = 6/76 (7%)
Frame = -3
Query: 22 PEQPPPPPPPPPNPQPDPHFSETLENPQIQTIP--DPDSTNPNHDQEDTLMDED---PTH 76
P PPPPPPPP P P P L +P P +P S E L DE+
Sbjct: 771 P*PEPPPPPPPPPPPPPPPLP*LLPDPDPAREPAYEPASEPAPEPPEPELEDEEAELEAE 592
Query: 77 PQIEDDPEPEP-PSPA 91
++E +PEPEP P+PA
Sbjct: 591 LELEPEPEPEPDPAPA 544
Score = 32.7 bits (73), Expect = 0.52
Identities = 26/72 (36%), Positives = 28/72 (38%), Gaps = 3/72 (4%)
Frame = -3
Query: 22 PEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLMDEDPT-HPQIE 80
P PPP P P P P P P P+ P P P L D DP P E
Sbjct: 828 P*SPPP*P*PSPWP*PSP-----*P*PEPPPPPPPPPPPPPPPLP*LLPDPDPAREPAYE 664
Query: 81 --DDPEPEPPSP 90
+P PEPP P
Sbjct: 663 PASEPAPEPPEP 628
Score = 28.5 bits (62), Expect = 9.8
Identities = 19/59 (32%), Positives = 27/59 (45%), Gaps = 13/59 (22%)
Frame = -3
Query: 25 PPPPPPPPPNPQPDPHFSETLENPQIQ-----------TIPDPDSTNPN--HDQEDTLM 70
P P P PPP P+P P S T P+ + TI D +N + HDQ+ ++
Sbjct: 543 PAPEPLPPPIPRPKPSPSPTPNPPRSRPKSFLATLLSDTIVDIKRSNASR*HDQDVAIL 367
>BU765146 similar to GP|21322711|e pherophorin-dz1 protein {Volvox carteri f.
nagariensis}, partial (25%)
Length = 407
Score = 50.1 bits (118), Expect = 3e-06
Identities = 28/82 (34%), Positives = 32/82 (38%)
Frame = -3
Query: 9 PNETLNPITTEQCPEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDT 68
P+ L P+ P PPPPPPPPP+P P P P P P P
Sbjct: 396 PSSLLPPLPPPPPPPPPPPPPPPPPSPPPPP--PPPPPPPPPPPPPPPPPLPPLPXSLSP 223
Query: 69 LMDEDPTHPQIEDDPEPEPPSP 90
L + P P P P PP P
Sbjct: 222 LKNPPPPPPPPPPPPPPPPPPP 157
Score = 48.1 bits (113), Expect = 1e-05
Identities = 28/83 (33%), Positives = 34/83 (40%)
Frame = -2
Query: 8 TPNETLNPITTEQCPEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQED 67
+P L P + P PPPPPPPPP+P P P P P P + P
Sbjct: 406 SPPPLLPPPSPPPPPPPPPPPPPPPPSPLPPPPPPXPPPPPPPPPPPPPPPSPP--PPXX 233
Query: 68 TLMDEDPTHPQIEDDPEPEPPSP 90
+L + P P P P PP P
Sbjct: 232 SLPPKKPPPPPPPPPPPPPPPPP 164
Score = 47.4 bits (111), Expect = 2e-05
Identities = 29/88 (32%), Positives = 32/88 (35%)
Frame = -2
Query: 4 PIADTPNETLNPITTEQCPEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNH 63
P P L P P PPPPPPPPP P P P +L + P P P
Sbjct: 349 PPPPPPPSPLPPPPPPXPPPPPPPPPPPPPPPSPPPP-XXSLPPKKPPPPPPPPPPPPPP 173
Query: 64 DQEDTLMDEDPTHPQIEDDPEPEPPSPA 91
L P P P P PP P+
Sbjct: 172 PPPPPLPPPPPPPPPPPPPPPPPPPPPS 89
Score = 45.8 bits (107), Expect = 6e-05
Identities = 26/79 (32%), Positives = 29/79 (35%)
Frame = -3
Query: 12 TLNPITTEQCPEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLMD 71
+L+P+ P PPPPPPPPP P P P P P P L
Sbjct: 234 SLSPLKNPPPPPPPPPPPPPPPPPPPFPP-------------PPPPPPPPPPPPPPPLPP 94
Query: 72 EDPTHPQIEDDPEPEPPSP 90
P P P P PP P
Sbjct: 93 PPPPPPPPPPPPPPPPPPP 37
Score = 45.1 bits (105), Expect = 1e-04
Identities = 30/92 (32%), Positives = 33/92 (35%), Gaps = 1/92 (1%)
Frame = -1
Query: 4 PIADTPNETLNPI-TTEQCPEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPN 62
P P+ L+P T P PPPPPPPPP P P P P P P P
Sbjct: 260 PPLSPPSLXLSPP*KTPPPPPPPPPPPPPPPPPPPSP--------PPPPPPPPPPPPPPP 105
Query: 63 HDQEDTLMDEDPTHPQIEDDPEPEPPSPAATT 94
P P P P PP P T+
Sbjct: 104 PSPPLPPPPPPPPPPPPPPPPPPPPPPPGQTS 9
Score = 41.2 bits (95), Expect = 0.001
Identities = 28/92 (30%), Positives = 30/92 (32%)
Frame = -3
Query: 1 MTNPIADTPNETLNPITTEQCPEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTN 60
+ NP P P P PPPPPPPPP P P P +P P
Sbjct: 222 LKNPPPPPPPPPPPPPPPPPPPFPPPPPPPPPPPPPPPP------------PLPPPPPPP 79
Query: 61 PNHDQEDTLMDEDPTHPQIEDDPEPEPPSPAA 92
P P P P P PP P A
Sbjct: 78 P-----------PPPPPPPPPPPPPPPPPPRA 16
Score = 41.2 bits (95), Expect = 0.001
Identities = 26/88 (29%), Positives = 31/88 (34%)
Frame = -1
Query: 4 PIADTPNETLNPITTEQCPEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNH 63
P + P+ P P PPPPPPPP P P P P P P + P+
Sbjct: 392 PPSSLPSPPPPPPPPPPPPPPLPPPPPPPPXPPPPP--------PPPPPPPPPPLSPPSL 237
Query: 64 DQEDTLMDEDPTHPQIEDDPEPEPPSPA 91
P P P P PP P+
Sbjct: 236 XLSPP*KTPPPPPPPPPPPPPPPPPPPS 153
Score = 40.4 bits (93), Expect = 0.003
Identities = 27/87 (31%), Positives = 29/87 (33%)
Frame = -1
Query: 4 PIADTPNETLNPITTEQCPEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNH 63
P P+ +P P PPPP PPPP P P P P P P P
Sbjct: 404 PPPPPPSSLPSPPPPPPPPPPPPPPLPPPPPPPPXP--------PPPPPPPPPPPPPPLS 249
Query: 64 DQEDTLMDEDPTHPQIEDDPEPEPPSP 90
L T P P P PP P
Sbjct: 248 PPSLXLSPP*KTPPPPPPPPPPPPPPP 168
Score = 38.1 bits (87), Expect = 0.012
Identities = 28/103 (27%), Positives = 30/103 (28%), Gaps = 15/103 (14%)
Frame = -2
Query: 3 NPIADTPNETLNPITTEQCPEQPPP---------------PPPPPPNPQPDPHFSETLEN 47
+P+ P P P PPP PPPPPP P P P L
Sbjct: 328 SPLPPPPPPXPPPPPPPPPPPPPPPSPPPPXXSLPPKKPPPPPPPPPPPPPPPPPPPLPP 149
Query: 48 PQIQTIPDPDSTNPNHDQEDTLMDEDPTHPQIEDDPEPEPPSP 90
P P P P P P P P PP P
Sbjct: 148 PPPPPPPPPPPPPPPPPPPSPPPPPPPPPPPPPPPPPPPPPPP 20
Score = 37.7 bits (86), Expect = 0.016
Identities = 15/27 (55%), Positives = 15/27 (55%)
Frame = -3
Query: 22 PEQPPPPPPPPPNPQPDPHFSETLENP 48
P PPPPPPPPP P P P L P
Sbjct: 81 PPPPPPPPPPPPPPPPPPPPRANLSPP 1
Score = 35.0 bits (79), Expect = 0.11
Identities = 17/40 (42%), Positives = 18/40 (44%)
Frame = -1
Query: 22 PEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNP 61
P P PPPPPPP P P P P P P T+P
Sbjct: 104 PSPPLPPPPPPPPPPPPP-------PPPPPPPPPPGQTSP 6
Score = 28.5 bits (62), Expect = 9.8
Identities = 13/34 (38%), Positives = 14/34 (40%)
Frame = -3
Query: 4 PIADTPNETLNPITTEQCPEQPPPPPPPPPNPQP 37
P+ P P P PPPPPPP N P
Sbjct: 105 PLPPPPPPPPPPPPPPPPPPPPPPPPPPRANLSP 4
>TC207027 weakly similar to UP|Q9LW95 (Q9LW95) KED, partial (17%)
Length = 1005
Score = 48.9 bits (115), Expect = 7e-06
Identities = 50/192 (26%), Positives = 93/192 (48%), Gaps = 24/192 (12%)
Frame = +3
Query: 297 KSQHKELFEETLVVEVEGEGEEGVVKDEEEEGEAKDEEEEEDG-----VVVKDEVDGSGD 351
K ++K+ ++T V +V+G+ +G DEE++ + K E++++DG + K E DG D
Sbjct: 51 KKENKDKGDKTDVEKVKGKENDGE-DDEEKKEKKKKEKKDKDGKTDAEKIKKKEDDGKDD 227
Query: 352 -------VKMGGVEENQVQELEEHNIELSLGQDKVETLPVEKEQGEGE-----QMMDFEQ 399
K ++ +V+ E+ G+D+ +KE+G+G+ + D E+
Sbjct: 228 EEKKEKKKKYDKIDAXKVKGKEDD------GKDEGNKEKKDKEKGDGDGEEKKEKKDKEK 389
Query: 400 SKK-------EETEMWFLGQKNYVGEPSLRPCHNRDRKGIDCEQVKEDEGEEEEHEQEEE 452
KK EET+ KN GE N+ +K E+ K+ + E+++ E+ E+
Sbjct: 390 EKKKEKKDKDEETDTLKEKGKNDEGEDDE---GNKKKKKDKKEKEKDHKKEKKDKEEGEK 560
Query: 453 EEDDVEEDEHDV 464
E+ VE D+
Sbjct: 561 EDSKVEVSVRDI 596
Score = 37.4 bits (85), Expect = 0.021
Identities = 41/178 (23%), Positives = 87/178 (48%), Gaps = 3/178 (1%)
Frame = +3
Query: 297 KSQHKELFEETLVVEVEGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDVKMGG 356
K + K+ +++ +V+G+ ++ KDE + + E+ + DG K++ D + K
Sbjct: 234 KKEKKKKYDKIDAXKVKGKEDD--GKDEGNKEKKDKEKGDGDGEEKKEKKDKEKEKKKEK 407
Query: 357 VEENQVQELEEHNIELSLGQDKVETLPVEKEQGEGEQMMDFEQSKKEETEMWFLGQKNYV 416
++++ + + + G+D +K++ E E+ E+ KEE E +++
Sbjct: 408 KDKDEETDTLKEKGKNDEGEDDEGNKKKKKDKKEKEKDHKKEKKDKEEGE-----KEDSK 572
Query: 417 GEPSLRPCHNRDRKGIDCEQVKEDEGEE---EEHEQEEEEEDDVEEDEHDVGFHFSTK 471
E S+R + D + I E KED+G++ E E++++E+ D +E + V TK
Sbjct: 573 VEVSVR---DIDIEEIKKEGEKEDKGKDGGKEVKEKKKKEDKDKKEKKKKVTGKDKTK 737
Score = 33.5 bits (75), Expect = 0.31
Identities = 40/175 (22%), Positives = 77/175 (43%), Gaps = 9/175 (5%)
Frame = +3
Query: 297 KSQHKELFEETLVVEVEGEGEEGVVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDVKMGG 356
K + K+ EET ++ +G+ +EG DE + + KD++E+E KD D + G
Sbjct: 396 KKEKKDKDEETDTLKEKGKNDEG-EDDEGNKKKKKDKKEKE-----KDHKKEKKDKEEG- 554
Query: 357 VEENQVQELEEHNIELSLGQDKVETLPVEKEQGEGEQM-----MDFEQSKKEETEMWFLG 411
E E+ +E+S+ +E + K++GE E + ++ KK+E +
Sbjct: 555 -------EKEDSKVEVSVRDIDIEEI---KKEGEKEDKGKDGGKEVKEKKKKEDKDKKEK 704
Query: 412 QKNYVGEPSLRPCHNRDRK----GIDCEQVKEDEGEEEEHEQEEEEEDDVEEDEH 462
+K G+ + +K + + E + + E +E E E V +E+
Sbjct: 705 KKKVTGKDKTKDLSTLKQKLEKINGKIQPLLEKKADIERQIKEVEAEGHVVNEEN 869
>CO985236
Length = 466
Score = 48.5 bits (114), Expect = 9e-06
Identities = 48/175 (27%), Positives = 87/175 (49%), Gaps = 6/175 (3%)
Frame = -1
Query: 295 LIKSQHKELFEETLVVEVEGEGEEGVVKDEEEEGEAKDEEEEE-DGVVVKDEVDGSGDV- 352
L++ +H+E EE + E +G E V +D +E+G + +E++E DGV EV+ G+V
Sbjct: 457 LVRIRHEEGVEEATEDKKE-DGVEEVNEDRKEDGVGEVKEDKEVDGVSCVKEVEEDGEVD 281
Query: 353 ---KMGGVEENQVQELEEHNIELSLGQ-DKVETLPVEKEQGEGEQMMDFEQSKKEETEMW 408
K+ + ++++E+EE + + + DK++ KE EGE ++ KEE+E
Sbjct: 280 GVKKVEDKKGDELKEIEEDKKDDHISENDKMDEDTEIKETMEGE-----PENGKEESE-- 122
Query: 409 FLGQKNYVGEPSLRPCHNRDRKGIDCEQVKEDEGEEEEHEQEEEEEDDVEEDEHD 463
+ +D +V+ E+EE ++ E+ E +EEDE D
Sbjct: 121 --------------------KPVVDAMEVEGGIKEKEESKENEKVEVVMEEDEKD 17
>TC219999 weakly similar to UP|Q7T9Y7 (Q7T9Y7) Odv-e66, partial (5%)
Length = 641
Score = 48.1 bits (113), Expect = 1e-05
Identities = 29/92 (31%), Positives = 36/92 (38%), Gaps = 10/92 (10%)
Frame = -3
Query: 12 TLNPITTEQCPEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDST----------NP 61
T +P T P PPPPPPPPP+P P P + P T P P S+ NP
Sbjct: 279 TPSPSPTTSSPSYPPPPPPPPPHPSPPPSSTP----PPTATCPTPSSSAKSNPLPPPFNP 112
Query: 62 NHDQEDTLMDEDPTHPQIEDDPEPEPPSPAAT 93
+H P P SP+A+
Sbjct: 111 SHLSPKATWHSSSPLPPFTSPYSIFPSSPSAS 16
>AW396581
Length = 401
Score = 47.0 bits (110), Expect = 3e-05
Identities = 36/146 (24%), Positives = 76/146 (51%), Gaps = 2/146 (1%)
Frame = -1
Query: 320 VVKDEEEEGEAKDEEEEEDGVVVKDEVDGSGDVKMGGVEENQVQELEEHNIELSLGQDKV 379
++K +E +GE +E++E+ KD+ D + K+ G E+ + ++ E+ + DK
Sbjct: 377 LIKVKEHDGEDAEEKKEKKKKEKKDKDDKTDVEKVKGKEDKEKKKKEK-----KVKDDKT 213
Query: 380 ETLPVEKEQGEGEQMMDFEQSKKEETEMWFLGQKNYVGEPSLRPCHNRDRKGIDCEQVK- 438
+ V+ ++ +GE + E+ KK++ + +D K ID E+VK
Sbjct: 212 DAEKVKGKEDDGEDNEEKEEKKKKKKK-------------------EKDDK-IDVEKVKG 93
Query: 439 -EDEGEEEEHEQEEEEEDDVEEDEHD 463
ED+GE+E ++++++E +++E D
Sbjct: 92 KEDDGEDEGKKEKKKKEKKDKDEETD 15
>TC231868 similar to UP|Q93424 (Q93424) C. elegans GRL-23 protein
(Corresponding sequence E02A10.2), partial (23%)
Length = 790
Score = 46.6 bits (109), Expect = 3e-05
Identities = 28/93 (30%), Positives = 33/93 (35%), Gaps = 6/93 (6%)
Frame = -3
Query: 4 PIADTPNETLNPITTEQCPEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNP-- 61
P P L+ + P+ P PPP PPP P P P P Q P P P
Sbjct: 452 PPPQPPQPLLSNLFPHPPPQPPQPPPQPPPQPAPQPPLLIKFPQPPPQPPPQPPQPPPLI 273
Query: 62 NHDQEDTLMDEDP----THPQIEDDPEPEPPSP 90
Q + P PQ P P+PP P
Sbjct: 272 KFPQPPPQPEPQPPLLIIFPQPPPQPAPQPPQP 174
Score = 40.0 bits (92), Expect = 0.003
Identities = 24/80 (30%), Positives = 30/80 (37%), Gaps = 13/80 (16%)
Frame = -3
Query: 22 PEQPPP-------------PPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDT 68
P QPPP PPP PP P P P + P + P P P +
Sbjct: 461 PPQPPPQPPQPLLSNLFPHPPPQPPQPPPQPPPQPAPQPPLLIKFPQPPPQPPPQPPQPP 282
Query: 69 LMDEDPTHPQIEDDPEPEPP 88
+ + P P PEP+PP
Sbjct: 281 PLIKFPQPP---PQPEPQPP 231
Score = 34.3 bits (77), Expect = 0.18
Identities = 19/62 (30%), Positives = 24/62 (38%)
Frame = -3
Query: 29 PPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLMDEDPTHPQIEDDPEPEPP 88
PP PP P P S +P Q P P + L+ + P P P P+PP
Sbjct: 461 PPQPPPQPPQPLLSNLFPHPPPQPPQPPPQPPPQPAPQPPLLIKFPQPP---PQPPPQPP 291
Query: 89 SP 90
P
Sbjct: 290 QP 285
>TC224573 homologue to UP|Q41707 (Q41707) Extensin class 1 protein precursor,
partial (44%)
Length = 1112
Score = 46.2 bits (108), Expect = 5e-05
Identities = 25/74 (33%), Positives = 30/74 (39%), Gaps = 5/74 (6%)
Frame = +3
Query: 22 PEQPPPPP-----PPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLMDEDPTH 76
P+ PPPP PPPP+P P PH+ P PDP P + + P
Sbjct: 282 PDPSPPPPYYYKSPPPPSPSPPPHYYYKSPPP-----PDPSPPPPYYYRSPPPPSPSPPP 446
Query: 77 PQIEDDPEPEPPSP 90
P P P PSP
Sbjct: 447 PYYYKSPPPPSPSP 488
Score = 37.4 bits (85), Expect = 0.021
Identities = 25/90 (27%), Positives = 32/90 (34%), Gaps = 8/90 (8%)
Frame = +3
Query: 9 PNETLNPITTEQCPEQPPPPP--------PPPPNPQPDPHFSETLENPQIQTIPDPDSTN 60
P+ + P + P P P P PPPP+P P P + P P P
Sbjct: 42 PSPSPPPPYYYKSPPPPDPSPPPPYYYKSPPPPSPSPPPPYYYKSPPP-----PSPSPPP 206
Query: 61 PNHDQEDTLMDEDPTHPQIEDDPEPEPPSP 90
P + + D P P P P PSP
Sbjct: 207 PYYYKSPPPPDPSPPPPYYYKSPPPPDPSP 296
Score = 37.0 bits (84), Expect = 0.028
Identities = 26/89 (29%), Positives = 35/89 (39%), Gaps = 6/89 (6%)
Frame = +3
Query: 9 PNETLNPITTEQCPEQPPPPPPP------PPNPQPDPHFSETLENPQIQTIPDPDSTNPN 62
P+ + P + P P P PPP PP P P P ++P P P + P
Sbjct: 186 PSPSPPPPYYYKSPPPPDPSPPPPYYYKSPPPPDPSPPPPYYYKSP-----PPPSPSPPP 350
Query: 63 HDQEDTLMDEDPTHPQIEDDPEPEPPSPA 91
H + DP+ P P PPSP+
Sbjct: 351 HYYYKSPPPPDPSPPPPYYYRSPPPPSPS 437
Score = 35.0 bits (79), Expect = 0.11
Identities = 22/79 (27%), Positives = 27/79 (33%), Gaps = 11/79 (13%)
Frame = +3
Query: 23 EQPPPPPP-----------PPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLMD 71
+ PPPP P PPP+P P P + P P P P + +
Sbjct: 27 KSPPPPSPSPPPPYYYKSPPPPDPSPPPPYYYKSPPP-----PSPSPPPPYYYKSPPPPS 191
Query: 72 EDPTHPQIEDDPEPEPPSP 90
P P P P PSP
Sbjct: 192 PSPPPPYYYKSPPPPDPSP 248
Score = 30.8 bits (68), Expect = 2.0
Identities = 23/90 (25%), Positives = 30/90 (32%), Gaps = 8/90 (8%)
Frame = +3
Query: 9 PNETLNPITTEQCPEQPPPPP--------PPPPNPQPDPHFSETLENPQIQTIPDPDSTN 60
P+ + P + P P P P PPPP+P P P + P P P
Sbjct: 330 PSPSPPPHYYYKSPPPPDPSPPPPYYYRSPPPPSPSPPPPYYYKSPPP-----PSPSPPP 494
Query: 61 PNHDQEDTLMDEDPTHPQIEDDPEPEPPSP 90
P + + P P P PSP
Sbjct: 495 PYYYKSPPPPSPVSHPPYYYKSPPPSTPSP 584
Score = 28.9 bits (63), Expect = 7.5
Identities = 14/40 (35%), Positives = 17/40 (42%), Gaps = 5/40 (12%)
Frame = +3
Query: 22 PEQPPPPP-----PPPPNPQPDPHFSETLENPQIQTIPDP 56
P PPPP PPPP+P P + P + P P
Sbjct: 474 PSPSPPPPYYYKSPPPPSPVSHPPYYYKSPPPSTPSPPPP 593
>TC224762 homologue to UP|Q09085 (Q09085) Hydroxyproline-rich glycoprotein
(HRGP) (Fragment), partial (38%)
Length = 422
Score = 45.4 bits (106), Expect = 8e-05
Identities = 25/74 (33%), Positives = 29/74 (38%), Gaps = 5/74 (6%)
Frame = +2
Query: 22 PEQPPPPP-----PPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLMDEDPTH 76
P PPPP PPPP+P P PH+ P PDP P + + P
Sbjct: 137 PSPSPPPPYYYKSPPPPSPSPPPHYYYKSPPP-----PDPSPPPPYYYRSPPPPSPSPPP 301
Query: 77 PQIEDDPEPEPPSP 90
P P P PSP
Sbjct: 302 PYYYKSPPPPSPSP 343
Score = 40.0 bits (92), Expect = 0.003
Identities = 23/70 (32%), Positives = 27/70 (37%), Gaps = 5/70 (7%)
Frame = +2
Query: 26 PPPP-----PPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLMDEDPTHPQIE 80
PPPP PPPP+P P P + P PDP P + + P P
Sbjct: 5 PPPPYYYKSPPPPDPSPPPSYYYKSPPP-----PDPSPPPPYYYKSPPPPSPSPPPPYYY 169
Query: 81 DDPEPEPPSP 90
P P PSP
Sbjct: 170KSPPPPSPSP 199
Score = 37.4 bits (85), Expect = 0.021
Identities = 26/89 (29%), Positives = 35/89 (39%), Gaps = 6/89 (6%)
Frame = +2
Query: 9 PNETLNPITTEQCPEQPPPPPPP------PPNPQPDPHFSETLENPQIQTIPDPDSTNPN 62
P+ + P + P P P PPP PP P P P ++P P P + P
Sbjct: 41 PDPSPPPSYYYKSPPPPDPSPPPPYYYKSPPPPSPSPPPPYYYKSP-----PPPSPSPPP 205
Query: 63 HDQEDTLMDEDPTHPQIEDDPEPEPPSPA 91
H + DP+ P P PPSP+
Sbjct: 206 HYYYKSPPPPDPSPPPPYYYRSPPPPSPS 292
>TC208361 weakly similar to UP|O64835 (O64835) Expressed protein
(At2g23670/F26B6.32), partial (72%)
Length = 636
Score = 44.3 bits (103), Expect = 2e-04
Identities = 25/91 (27%), Positives = 38/91 (41%)
Frame = -2
Query: 4 PIADTPNETLNPITTEQCPEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNH 63
P A TP + +P++ P P PPPP P P P P+ + Q +P P S+
Sbjct: 413 PPAPTPQNSPSPLSPSPLPLSPSPPPPTPSPPPPIPYPQPQIPLSQSPPLPPPPSS---- 246
Query: 64 DQEDTLMDEDPTHPQIEDDPEPEPPSPAATT 94
D + + + P P P P+ +T
Sbjct: 245 ---------DSSVSRTRECPAPRTPLPSPST 180
Score = 36.6 bits (83), Expect = 0.036
Identities = 23/83 (27%), Positives = 35/83 (41%)
Frame = -2
Query: 14 NPITTEQCPEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLMDED 73
+P+ + ++PP PPP PP P P S +P + P P P+ +
Sbjct: 467 SPLPLQPKSQKPPLPPPLPPAPTPQNSPSPLSPSP-LPLSPSPPPPTPS--PPPPIPYPQ 297
Query: 74 PTHPQIEDDPEPEPPSPAATTVR 96
P P + P P PPS ++ R
Sbjct: 296 PQIPLSQSPPLPPPPSSDSSVSR 228
>TC230046 similar to UP|Q9LU79 (Q9LU79) Gb|AAF21150.1, partial (28%)
Length = 1798
Score = 43.9 bits (102), Expect = 2e-04
Identities = 27/100 (27%), Positives = 42/100 (42%), Gaps = 2/100 (2%)
Frame = +3
Query: 5 IADTPNET--LNPITTEQCPEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPN 62
+ D P +T + P PPPPPPPPP P + + +++ + + T
Sbjct: 729 VKDIPVDTFEIRPSPPPVKSTSPPPPPPPPPPPPESARHNTRRTHRKVERERESEITVEL 908
Query: 63 HDQEDTLMDEDPTHPQIEDDPEPEPPSPAATTVRRGHKRK 102
D E T + P P P+PPS + R+ +RK
Sbjct: 909 DDHEFTTIRSPPPAP----PTPPQPPSVKTRSERKSDRRK 1016
Score = 35.0 bits (79), Expect = 0.11
Identities = 29/99 (29%), Positives = 36/99 (36%)
Frame = +3
Query: 26 PPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLMDEDPTHPQIEDDPEP 85
PPPPPPPP + E PQ + P N D ++ P I P P
Sbjct: 1266 PPPPPPPPPSKRKSQTPPPPEPPQRRNSGRPPLPNRTVTFNDETLNAGNQSPLI---PVP 1436
Query: 86 EPPSPAATTVRRGHKRKKFGGKRTAAQKRKSHEKFEVII 124
PP P + R F R+ R S + E II
Sbjct: 1437 PPPPPFKMKAMKFVVRGDFVKIRSNQSSRCSSPEREDII 1553
Score = 32.7 bits (73), Expect = 0.52
Identities = 21/66 (31%), Positives = 30/66 (44%), Gaps = 2/66 (3%)
Frame = +3
Query: 25 PPPPPPPPPNPQP-DPHFSETL-ENPQIQTIPDPDSTNPNHDQEDTLMDEDPTHPQIEDD 82
PPPPPPPPP P F + L ++ +I ++ P P P+ + +
Sbjct: 1167 PPPPPPPPPLPSVFHSLFRKGLGKSKKIHSVSAPPPPPP----------PPPSKRKSQTP 1316
Query: 83 PEPEPP 88
P PEPP
Sbjct: 1317 PPPEPP 1334
>BQ080060 homologue to PIR|C29356|C29 hydroxyproline-rich glycoprotein (clone
Hyp2.13) - kidney bean (fragment), partial (38%)
Length = 422
Score = 43.1 bits (100), Expect = 4e-04
Identities = 26/81 (32%), Positives = 31/81 (38%), Gaps = 10/81 (12%)
Frame = +1
Query: 22 PEQPPPPP-----PPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLMDEDPTH 76
P PPPP PPPP+P P PH+ P PDP P + + P
Sbjct: 148 PSPSPPPPYYYKSPPPPSPSPPPHYYYKSPPP-----PDPSPPPPYYYRSPPPPSPSPPP 312
Query: 77 PQIEDDP-----EPEPPSPAA 92
P P P PPSP +
Sbjct: 313 PYYYKSPPYYYKSPPPPSPVS 375
Score = 37.0 bits (84), Expect = 0.028
Identities = 26/89 (29%), Positives = 35/89 (39%), Gaps = 6/89 (6%)
Frame = +1
Query: 9 PNETLNPITTEQCPEQPPPPPPP------PPNPQPDPHFSETLENPQIQTIPDPDSTNPN 62
P+ + P + P P P PPP PP P P P ++P P P + P
Sbjct: 52 PSPSPPPPYYYKSPPPPDPSPPPPYYYKSPPPPSPSPPPPYYYKSP-----PPPSPSPPP 216
Query: 63 HDQEDTLMDEDPTHPQIEDDPEPEPPSPA 91
H + DP+ P P PPSP+
Sbjct: 217 HYYYKSPPPPDPSPPPPYYYRSPPPPSPS 303
Score = 35.0 bits (79), Expect = 0.11
Identities = 22/74 (29%), Positives = 26/74 (34%), Gaps = 8/74 (10%)
Frame = +1
Query: 25 PPPPPP--------PPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLMDEDPTH 76
P P PP PPP+P P P + P PDP P + + P
Sbjct: 4 PSPSPPPP*YYKSPPPPSPSPPPPYYYKSPPP-----PDPSPPPPYYYKSPPPPSPSPPP 168
Query: 77 PQIEDDPEPEPPSP 90
P P P PSP
Sbjct: 169PYYYKSPPPPSPSP 210
>TC215424 similar to UP|Q9MA04 (Q9MA04) F20B17.16, partial (26%)
Length = 1030
Score = 43.1 bits (100), Expect = 4e-04
Identities = 29/120 (24%), Positives = 45/120 (37%), Gaps = 2/120 (1%)
Frame = +3
Query: 22 PEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLMDEDPTHPQIED 81
P P PPPPPPP P+ Q P P + + ++ P P
Sbjct: 123 PHYPYPPPPPPPPPE-----------ASYQPPPPPPPAPMYYPSNNQYSNQPPPPPPPLS 269
Query: 82 DPEPEPP--SPAATTVRRGHKRKKFGGKRTAAQKRKSHEKFEVIIQTLKPIPFVPDKVLD 139
P P PP P V + ++ + F T+ ++ H + + K P VP K ++
Sbjct: 270 PPPPPPPVSPPPPPPVTQNNEERHFKEPSTSGRREYEHSNHGIAHKQHKQQPPVPVKKMN 449
Score = 34.7 bits (78), Expect = 0.14
Identities = 28/112 (25%), Positives = 41/112 (36%), Gaps = 9/112 (8%)
Frame = +3
Query: 17 TTEQCPEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHD--QEDTLMDEDP 74
+ Q QPPPPPPP P P P S P Q + P+ +E +
Sbjct: 219 SNNQYSNQPPPPPPPLSPPPPPPPVSPPPPPPVTQNNEERHFKEPSTSGRREYEHSNHGI 398
Query: 75 THPQIEDDP-------EPEPPSPAATTVRRGHKRKKFGGKRTAAQKRKSHEK 119
H Q + P PP A T + ++K+ K+ +K + K
Sbjct: 399 AHKQHKQQPPVPVKKMNNGPPGRAETDEEKRLRKKREFEKQRQEEKHRQQLK 554
>TC214106 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 protein
precursor, partial (22%)
Length = 1296
Score = 42.7 bits (99), Expect = 5e-04
Identities = 25/78 (32%), Positives = 29/78 (37%), Gaps = 2/78 (2%)
Frame = +1
Query: 15 PITTEQCPEQPPPPPPPPPNPQPDPH--FSETLENPQIQTIPDPDSTNPNHDQEDTLMDE 72
P + C PPPP PPPP+P P P FS + P P +P
Sbjct: 82 PPSPTYCVRSPPPPSPPPPSPPPPPPPVFSPPPPVQYYYSSPPPPQHSP----------- 228
Query: 73 DPTHPQIEDDPEPEPPSP 90
P P P P PP P
Sbjct: 229 PPPPPHSPPPPPPSPPPP 282
Score = 42.4 bits (98), Expect = 7e-04
Identities = 26/80 (32%), Positives = 29/80 (35%), Gaps = 4/80 (5%)
Frame = +1
Query: 15 PITTEQCPEQPPPPP----PPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLM 70
P + C E PPPPP PPPP P P P + P P S +P
Sbjct: 361 PPSPPPCIEPPPPPPPCIEPPPPPPSPPPCEEHS---------PPPPSPHPAPYHPPPSP 513
Query: 71 DEDPTHPQIEDDPEPEPPSP 90
P Q P P PP P
Sbjct: 514 SPPPPPVQYNSPPPPSPPPP 573
Score = 42.0 bits (97), Expect = 9e-04
Identities = 24/69 (34%), Positives = 29/69 (41%)
Frame = +1
Query: 22 PEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLMDEDPTHPQIED 81
P PPPPPP P P P+ S P + + P P + P E P P
Sbjct: 238 PPHSPPPPPPSPPPPVYPYLSPP-PPPPVHSPPPPVYSPPPPPPSPPPCIEPPPPPPPCI 414
Query: 82 DPEPEPPSP 90
+P P PPSP
Sbjct: 415 EPPPPPPSP 441
Score = 36.6 bits (83), Expect = 0.036
Identities = 19/64 (29%), Positives = 24/64 (36%)
Frame = +1
Query: 25 PPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLMDEDPTHPQIEDDPE 84
PPPP PPPP+P P + + P P + P + P P P
Sbjct: 4 PPPPSPPPPSPP-----------PPVYSPPPPPPSPPPPSPTYCVRSPPPPSPPPPSPPP 150
Query: 85 PEPP 88
P PP
Sbjct: 151PPPP 162
Score = 36.2 bits (82), Expect = 0.047
Identities = 23/76 (30%), Positives = 26/76 (33%), Gaps = 10/76 (13%)
Frame = +1
Query: 25 PPPPPP----------PPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLMDEDP 74
PPPPPP PPP P P E P P P +P +E + P
Sbjct: 301 PPPPPPVHSPPPPVYSPPPPPPSPPPCIEPPPPPPPCIEPPPPPPSPPPCEEHSPPPPSP 480
Query: 75 THPQIEDDPEPEPPSP 90
P P PP P
Sbjct: 481 HPAPYHPPPSPSPPPP 528
Score = 34.3 bits (77), Expect = 0.18
Identities = 29/99 (29%), Positives = 32/99 (32%), Gaps = 11/99 (11%)
Frame = +1
Query: 3 NPIADTPNETLNPITTEQCPEQPPPPPPP--------PPNPQPDPHFSETLENPQIQTIP 54
+P +P P P PP PPPP PP P P P P P
Sbjct: 1 SPPPPSPPPPSPPPPVYSPPPPPPSPPPPSPTYCVRSPPPPSPPP--------PSPPPPP 156
Query: 55 DPDSTNPNHDQEDTLMDEDPTH---PQIEDDPEPEPPSP 90
P + P Q P H P P P PPSP
Sbjct: 157PPVFSPPPPVQYYYSSPPPPQHSPPPPPPHSPPPPPPSP 273
Score = 32.3 bits (72), Expect = 0.68
Identities = 25/91 (27%), Positives = 30/91 (32%), Gaps = 7/91 (7%)
Frame = +1
Query: 15 PITTEQCPEQPPPPPP-------PPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQED 67
P + C E PPPP PPP+P P P P +Q P + P
Sbjct: 430 PPSPPPCEEHSPPPPSPHPAPYHPPPSPSPPP--------PPVQYNSPPPPSPP------ 567
Query: 68 TLMDEDPTHPQIEDDPEPEPPSPAATTVRRG 98
P P + P P P T V G
Sbjct: 568 ------PPTPVYHYNSPPPPSFPPPTPVYEG 642
Score = 29.3 bits (64), Expect = 5.8
Identities = 24/87 (27%), Positives = 29/87 (32%), Gaps = 5/87 (5%)
Frame = +1
Query: 9 PNETLNPITTEQCPEQPPPPPP-----PPPNPQPDPHFSETLENPQIQTIPDPDSTNPNH 63
P + +P P PPPPP PPP P P +P + P P
Sbjct: 466 PPPSPHPAPYHPPPSPSPPPPPVQYNSPPPPSPPPPTPVYHYNSPPPPSFPPP------- 624
Query: 64 DQEDTLMDEDPTHPQIEDDPEPEPPSP 90
T + E P P I PP P
Sbjct: 625 ----TPVYEGPLPPVIGVSYASPPPPP 693
>TC211475 similar to UP|P79122 (P79122) Pinin, partial (5%)
Length = 436
Score = 42.7 bits (99), Expect = 5e-04
Identities = 22/67 (32%), Positives = 29/67 (42%)
Frame = +2
Query: 22 PEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLMDEDPTHPQIED 81
P PPPPPP NPQP P SE+ +T +P T + + P+ +
Sbjct: 230 PISKNPPPPPPSNPQPQPTSSESETESGSETESEPHPTPVKVKPLASKPMDQAQKPKAQP 409
Query: 82 DPEPEPP 88
P P PP
Sbjct: 410 SPAPPPP 430
>TC210023 weakly similar to UP|Q868B4 (Q868B4) C. elegans GRL-25 protein
(Corresponding sequence ZK643.8), partial (12%)
Length = 819
Score = 42.7 bits (99), Expect = 5e-04
Identities = 25/83 (30%), Positives = 34/83 (40%)
Frame = -3
Query: 8 TPNETLNPITTEQCPEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQED 67
+P + P+ + P PP PP PPP P P P +P Q P P P +
Sbjct: 466 SPPQPPQPLLSNLFPHPPPQPPQPPPQP-PQPPLFNIDPHPPPQPPPQPAPQPP----QP 302
Query: 68 TLMDEDPTHPQIEDDPEPEPPSP 90
L+ + P P P P+ P P
Sbjct: 301 PLLIKFPQPPPQPPQPAPQRPQP 233
Score = 36.2 bits (82), Expect = 0.047
Identities = 26/87 (29%), Positives = 29/87 (32%)
Frame = -3
Query: 4 PIADTPNETLNPITTEQCPEQPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNH 63
P P L I P+ PP P P PP P F Q P P P
Sbjct: 397 PPPQPPQPPLFNIDPHPPPQPPPQPAPQPPQPPLLIKFP--------QPPPQPPQPAPQR 242
Query: 64 DQEDTLMDEDPTHPQIEDDPEPEPPSP 90
Q L+ PQ P P+PP P
Sbjct: 241 PQPPLLI----KFPQPPPQPAPQPPQP 173
Score = 35.8 bits (81), Expect = 0.062
Identities = 25/93 (26%), Positives = 35/93 (36%), Gaps = 8/93 (8%)
Frame = -3
Query: 9 PNETLNPITTEQCPEQPPPPPPPPP--------NPQPDPHFSETLENPQIQTIPDPDSTN 60
P ++P Q P QP P PP PP PQP + + P + P P
Sbjct: 373 PLFNIDPHPPPQPPPQPAPQPPQPPLLIKFPQPPPQPPQPAPQRPQPPLLIKFPQPPPQP 194
Query: 61 PNHDQEDTLMDEDPTHPQIEDDPEPEPPSPAAT 93
+ L+ + P P P+P P P +T
Sbjct: 193 APQPPQPPLLIKFPQPP-----PQPAPQPPLST 110
Score = 33.5 bits (75), Expect = 0.31
Identities = 22/69 (31%), Positives = 25/69 (35%)
Frame = -3
Query: 24 QPPPPPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLMDEDPTHPQIEDDP 83
QPPP PP P +P P P Q P P Q L+ PQ P
Sbjct: 280 QPPPQPPQPAPQRPQPPLLIKFPQPPPQPAPQP-------PQPPLLI----KFPQPPPQP 134
Query: 84 EPEPPSPAA 92
P+PP A
Sbjct: 133 APQPPLSTA 107
Score = 31.2 bits (69), Expect = 1.5
Identities = 13/38 (34%), Positives = 17/38 (44%), Gaps = 5/38 (13%)
Frame = -3
Query: 19 EQCPEQPPPP-----PPPPPNPQPDPHFSETLENPQIQ 51
+ P+ P PP P PPP P P P S P ++
Sbjct: 199 QPAPQPPQPPLLIKFPQPPPQPAPQPPLSTAFPQPPME 86
>TC214303 similar to UP|Q9T0K5 (Q9T0K5) Extensin-like protein, partial (18%)
Length = 1014
Score = 42.4 bits (98), Expect = 7e-04
Identities = 27/76 (35%), Positives = 30/76 (38%), Gaps = 7/76 (9%)
Frame = +2
Query: 22 PEQPPP---PPPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLMDEDPTHPQ 78
P PPP PPPPPP+P P PH P + P P H + P P
Sbjct: 29 PPAPPPTYSPPPPPPSP-PPPH--SPPPPPPVYPYLSPPPPPPVHSPPPPVYSPPPPPPS 199
Query: 79 ----IEDDPEPEPPSP 90
IE P P PP P
Sbjct: 200 PPPCIESPPPPPPPPP 247
Score = 42.0 bits (97), Expect = 9e-04
Identities = 28/89 (31%), Positives = 31/89 (34%), Gaps = 5/89 (5%)
Frame = +2
Query: 15 PITTEQCPEQPPPPPPPPP-----NPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTL 69
P + C E PPPPPPPPP P P PH + P P P N
Sbjct: 191 PPSPPPCIESPPPPPPPPPCEEHSPPPPSPHSAPYHPPPSPSPPPPPIQYN-----SPPP 355
Query: 70 MDEDPTHPQIEDDPEPEPPSPAATTVRRG 98
P P + P P SP V G
Sbjct: 356 PSPPPPTPVYHYNSPPPPSSPPPAPVYEG 442
Score = 37.0 bits (84), Expect = 0.028
Identities = 24/80 (30%), Positives = 28/80 (35%), Gaps = 4/80 (5%)
Frame = +2
Query: 15 PITTEQCPEQPPPP----PPPPPNPQPDPHFSETLENPQIQTIPDPDSTNPNHDQEDTLM 70
P+ + P PPP PPPPP P P S +P P S +P
Sbjct: 167 PVYSPPPPPPSPPPCIESPPPPPPPPPCEEHSPPPPSPHSAPYHPPPSPSP--------- 319
Query: 71 DEDPTHPQIEDDPEPEPPSP 90
P Q P P PP P
Sbjct: 320 --PPPPIQYNSPPPPSPPPP 373
Score = 30.8 bits (68), Expect = 2.0
Identities = 18/58 (31%), Positives = 20/58 (34%), Gaps = 5/58 (8%)
Frame = +2
Query: 9 PNETLNPITTEQCPEQPPPP-----PPPPPNPQPDPHFSETLENPQIQTIPDPDSTNP 61
P+ P P PPPP PPPP P P P + P P P P
Sbjct: 272 PSPHSAPYHPPPSPSPPPPPIQYNSPPPPSPPPPTPVYHYNSPPPPSSPPPAPVYEGP 445
Score = 28.5 bits (62), Expect = 9.8
Identities = 13/32 (40%), Positives = 15/32 (46%), Gaps = 8/32 (25%)
Frame = +2
Query: 22 PEQPPPP--------PPPPPNPQPDPHFSETL 45
P PPPP PPPP +P P P + L
Sbjct: 353 PPSPPPPTPVYHYNSPPPPSSPPPAPVYEGPL 448
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.311 0.131 0.377
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 29,002,528
Number of Sequences: 63676
Number of extensions: 580932
Number of successful extensions: 31042
Number of sequences better than 10.0: 1228
Number of HSP's better than 10.0 without gapping: 8057
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 16810
length of query: 652
length of database: 12,639,632
effective HSP length: 103
effective length of query: 549
effective length of database: 6,081,004
effective search space: 3338471196
effective search space used: 3338471196
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 62 (28.5 bits)
Medicago: description of AC144893.11


