

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
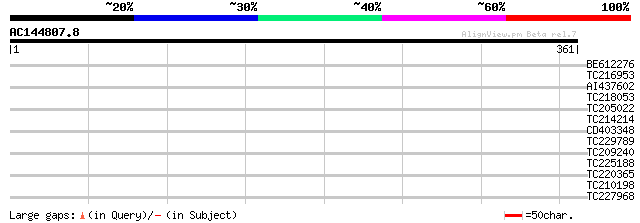
Query= AC144807.8 - phase: 0
(361 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BE612276 31 0.76
TC216953 similar to UP|PIRL_LYCES (Q9SEE4) Pirin-like protein, p... 29 2.9
AI437602 28 5.0
TC218053 weakly similar to UP|Q6I621 (Q6I621) Protein phosphatas... 28 6.5
TC205022 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, pa... 28 6.5
TC214214 weakly similar to UP|PRP1_SOYBN (P08012) Repetitive pro... 28 8.5
CD403348 28 8.5
TC229789 homologue to UP|Q9FVK4 (Q9FVK4) Resistance protein LM17... 28 8.5
TC209240 similar to UP|Q9LYS2 (Q9LYS2) ABC transporter-like prot... 28 8.5
TC225188 similar to UP|Q9FYB7 (Q9FYB7) Splicing factor-like prot... 28 8.5
TC220365 weakly similar to UP|Q6NLS5 (Q6NLS5) At5g16800, partial... 28 8.5
TC210198 28 8.5
TC227968 similar to UP|Q9M1C7 (Q9M1C7) Multi resistance protein ... 28 8.5
>BE612276
Length = 400
Score = 31.2 bits (69), Expect = 0.76
Identities = 19/66 (28%), Positives = 27/66 (40%)
Frame = +3
Query: 284 GPWLRYNHFGRRIMDYKDRKFSSNPTKCKNYGHFMPPIPPKMLEEMAKMSMLNHIKDHPQ 343
GPW YNHF + + SSNP +N IPP + S H +++
Sbjct: 24 GPWFLYNHFPVSQIFWPSILQSSNPVHLQNTSFNSIAIPPNANVPCSSESESRHKQNNLI 203
Query: 344 NHNTNE 349
N N +
Sbjct: 204 NDNRTQ 221
>TC216953 similar to UP|PIRL_LYCES (Q9SEE4) Pirin-like protein, partial (60%)
Length = 943
Score = 29.3 bits (64), Expect = 2.9
Identities = 19/58 (32%), Positives = 22/58 (37%)
Frame = +3
Query: 254 FACGLIGHTEANCKLHDTRTNAESRNKNVLGPWLRYNHFGRRIMDYKDRKFSSNPTKC 311
FAC C LHD R + S + V+ YN GR I K S P C
Sbjct: 213 FACVHTNTNHVPCFLHDARNSVASEDSRVVECLCLYN*RGRSIRVSKFIPNSGTPCPC 386
>AI437602
Length = 277
Score = 28.5 bits (62), Expect = 5.0
Identities = 14/42 (33%), Positives = 21/42 (49%)
Frame = +2
Query: 5 SPNNPYPTQNSTQINNKLMKNYLCSLPLMKMHHQNAITSQDP 46
SPN+ T+ ++ + Y SLPL +HH + QDP
Sbjct: 29 SPNHQNQTKKKWKLLCQNSSYYSSSLPLRLLHHLPQVEPQDP 154
>TC218053 weakly similar to UP|Q6I621 (Q6I621) Protein phosphatase 2A B'kappa
subunit, partial (74%)
Length = 2136
Score = 28.1 bits (61), Expect = 6.5
Identities = 14/39 (35%), Positives = 24/39 (60%)
Frame = +2
Query: 70 YDDSDIQERLEECQNSILGKIVSEKAIHRNSIQNALSNI 108
+D D +ER +C +IL ++ + IHR I+ ++SNI
Sbjct: 1022 FDSEDPRER--DCLKTILHRVYGKFMIHRPFIRKSVSNI 1132
>TC205022 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, partial (69%)
Length = 1623
Score = 28.1 bits (61), Expect = 6.5
Identities = 16/44 (36%), Positives = 20/44 (45%), Gaps = 1/44 (2%)
Frame = +2
Query: 225 PIKAGIYIGSAKDGAHWI-DFRYENLPQFCFACGLIGHTEANCK 267
P +G DG HW D + + C+ CG GH E NCK
Sbjct: 296 PPGSGRCFNCGLDG-HWARDCKAGDWKNKCYRCGERGHIERNCK 424
>TC214214 weakly similar to UP|PRP1_SOYBN (P08012) Repetitive proline-rich
cell wall protein 1 precursor, partial (53%)
Length = 857
Score = 27.7 bits (60), Expect = 8.5
Identities = 15/45 (33%), Positives = 22/45 (48%)
Frame = -3
Query: 256 CGLIGHTEANCKLHDTRTNAESRNKNVLGPWLRYNHFGRRIMDYK 300
C L+G+ + CKL D K VL + Y R+++DYK
Sbjct: 180 CNLVGYRKVTCKLVD--------YKMVLCNLVHYRKVTRKLVDYK 70
>CD403348
Length = 597
Score = 27.7 bits (60), Expect = 8.5
Identities = 11/33 (33%), Positives = 19/33 (57%)
Frame = -3
Query: 276 ESRNKNVLGPWLRYNHFGRRIMDYKDRKFSSNP 308
+SR+ G W + +H GRR + +K +S+P
Sbjct: 415 KSRDNQGRGEWRKNDHHGRRASSSRSKKAASHP 317
>TC229789 homologue to UP|Q9FVK4 (Q9FVK4) Resistance protein LM17 (Fragment),
partial (94%)
Length = 2104
Score = 27.7 bits (60), Expect = 8.5
Identities = 11/25 (44%), Positives = 17/25 (68%)
Frame = -3
Query: 3 PISPNNPYPTQNSTQINNKLMKNYL 27
PISP +P NS+ ++ +L+ NYL
Sbjct: 527 PISPTSPCYALNSSHVSPRLLHNYL 453
>TC209240 similar to UP|Q9LYS2 (Q9LYS2) ABC transporter-like protein, partial
(14%)
Length = 872
Score = 27.7 bits (60), Expect = 8.5
Identities = 19/68 (27%), Positives = 30/68 (43%)
Frame = +2
Query: 27 LCSLPLMKMHHQNAITSQDPAIENDPTEETVTEDTAETDSFIYYDDSDIQERLEECQNSI 86
+CS+ L + + I QDP + N + D + D +I E L +CQ
Sbjct: 53 ICSIGLHDLRSRFGIIPQDPTLFNGTVRYNM-------DPLSQHSDKEIWEVLRKCQ--- 202
Query: 87 LGKIVSEK 94
L ++V EK
Sbjct: 203 LREVVEEK 226
>TC225188 similar to UP|Q9FYB7 (Q9FYB7) Splicing factor-like protein, partial
(62%)
Length = 1131
Score = 27.7 bits (60), Expect = 8.5
Identities = 12/29 (41%), Positives = 15/29 (51%), Gaps = 1/29 (3%)
Frame = +1
Query: 240 HWI-DFRYENLPQFCFACGLIGHTEANCK 267
HW D + + C+ CG GH E NCK
Sbjct: 388 HWARDCKAGDWKNKCYRCGERGHIEKNCK 474
>TC220365 weakly similar to UP|Q6NLS5 (Q6NLS5) At5g16800, partial (45%)
Length = 1028
Score = 27.7 bits (60), Expect = 8.5
Identities = 11/22 (50%), Positives = 13/22 (59%)
Frame = +3
Query: 238 GAHWIDFRYENLPQFCFACGLI 259
G H I F+Y NLP AC L+
Sbjct: 366 GGHKICFKYSNLPSCLLACNLL 431
>TC210198
Length = 508
Score = 27.7 bits (60), Expect = 8.5
Identities = 13/36 (36%), Positives = 21/36 (58%)
Frame = +1
Query: 25 NYLCSLPLMKMHHQNAITSQDPAIENDPTEETVTED 60
N+LCS P + H+Q + S+D P ET++E+
Sbjct: 16 NFLCSTPEXRKHYQMLVKSKDQVW--SPKLETISEN 117
>TC227968 similar to UP|Q9M1C7 (Q9M1C7) Multi resistance protein homolog,
partial (21%)
Length = 1587
Score = 27.7 bits (60), Expect = 8.5
Identities = 21/68 (30%), Positives = 32/68 (46%)
Frame = +1
Query: 27 LCSLPLMKMHHQNAITSQDPAIENDPTEETVTEDTAETDSFIYYDDSDIQERLEECQNSI 86
+C + L + + +I QDPA+ E TV D Y D ++ E L++CQ
Sbjct: 430 ICKIGLHDLRSRLSIIPQDPAL----FEGTVR---GNLDPLQKYSDIEVWEALDKCQ--- 579
Query: 87 LGKIVSEK 94
LG +V K
Sbjct: 580 LGHLVRAK 603
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.135 0.425
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,353,937
Number of Sequences: 63676
Number of extensions: 304546
Number of successful extensions: 1731
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 1710
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1731
length of query: 361
length of database: 12,639,632
effective HSP length: 98
effective length of query: 263
effective length of database: 6,399,384
effective search space: 1683037992
effective search space used: 1683037992
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 59 (27.3 bits)
Medicago: description of AC144807.8


