

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144807.5 + phase: 0
(112 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
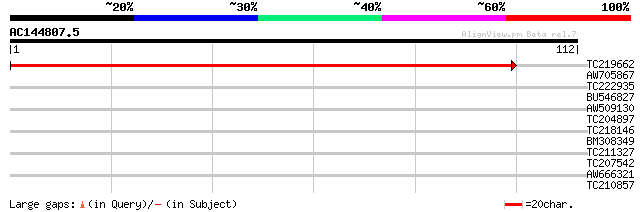
TC219662 weakly similar to UP|Q6NPH1 (Q6NPH1) At1g48200, partial... 136 1e-33
AW705867 similar to EGAD|27174|2799 cell death-inducing protein ... 27 1.1
TC222935 similar to UP|Q8RY64 (Q8RY64) AT4g37080/C7A10_280, part... 27 1.4
BU546827 26 2.5
AW509130 25 4.2
TC204897 similar to UP|Q8W419 (Q8W419) Twin LOV protein 1, parti... 25 4.2
TC218146 25 4.2
BM308349 25 5.5
TC211327 similar to UP|Q94JT2 (Q94JT2) AT5g17630/K10A8_110, part... 25 5.5
TC207542 similar to UP|Q6H976 (Q6H976) STYLOSA protein, partial ... 25 5.5
AW666321 25 5.5
TC210857 similar to UP|Q9M7I2 (Q9M7I2) WD-repeat cell cycle regu... 24 9.4
>TC219662 weakly similar to UP|Q6NPH1 (Q6NPH1) At1g48200, partial (64%)
Length = 800
Score = 136 bits (343), Expect = 1e-33
Identities = 71/100 (71%), Positives = 81/100 (81%)
Frame = +3
Query: 1 MEFVHKIVDIAKRASENNTVINVGLGAVFAILGARSYNQQKIIEALEAEKDSLTKTNKSI 60
MEF ++IVDIA RA +N VINV L A FA LG RS NQQK IEAL+ EK+SLTK+NKSI
Sbjct: 105 MEFSNRIVDIATRAVNSNAVINVCLLASFATLGVRSMNQQKTIEALQDEKESLTKSNKSI 284
Query: 61 RNTLWDWKQQLYAEAASDSAPVPLARLYEIYGEAAPPQQS 100
R TLWDWKQQLYAEA++DSA VPLARL IYGEA PP ++
Sbjct: 285 RKTLWDWKQQLYAEASADSAVVPLARLKAIYGEAPPPPRA 404
>AW705867 similar to EGAD|27174|2799 cell death-inducing protein Bik {Homo
sapiens}, partial (9%)
Length = 420
Score = 27.3 bits (59), Expect = 1.1
Identities = 14/49 (28%), Positives = 23/49 (46%)
Frame = +1
Query: 34 ARSYNQQKIIEALEAEKDSLTKTNKSIRNTLWDWKQQLYAEAASDSAPV 82
+ S +++ +++ K S K + R WDW+Q AE S S V
Sbjct: 130 SHSIKRRRFHDSIIHRKQSKKKKQQQ*REFTWDWEQLREAEEESSSKKV 276
>TC222935 similar to UP|Q8RY64 (Q8RY64) AT4g37080/C7A10_280, partial (17%)
Length = 312
Score = 26.9 bits (58), Expect = 1.4
Identities = 17/40 (42%), Positives = 19/40 (47%), Gaps = 7/40 (17%)
Frame = +1
Query: 35 RSYNQQKIIEALEAEKDS-------LTKTNKSIRNTLWDW 67
R Y K+ E LEA K +TKTNK I L DW
Sbjct: 16 RVYTASKVDEELEAAKRDYLHASVGITKTNKLIIPKLLDW 135
>BU546827
Length = 401
Score = 26.2 bits (56), Expect = 2.5
Identities = 17/59 (28%), Positives = 28/59 (46%)
Frame = +3
Query: 14 ASENNTVINVGLGAVFAILGARSYNQQKIIEALEAEKDSLTKTNKSIRNTLWDWKQQLY 72
A E++T+++ G V I+ Q + I +A T++S + TLW W Q Y
Sbjct: 120 APEHSTILSAGWKHVQVII-----TQTQSISGRKAAICRCQLTSESAQYTLWQWIQLAY 281
>AW509130
Length = 424
Score = 25.4 bits (54), Expect = 4.2
Identities = 13/42 (30%), Positives = 21/42 (49%)
Frame = -1
Query: 37 YNQQKIIEALEAEKDSLTKTNKSIRNTLWDWKQQLYAEAASD 78
+N+ +++ + D L K N R LWDW +Q Y + D
Sbjct: 409 WNRLRLLHGWDWLWDRLEKWN---RKWLWDWLEQWYRKWLGD 293
>TC204897 similar to UP|Q8W419 (Q8W419) Twin LOV protein 1, partial (59%)
Length = 1749
Score = 25.4 bits (54), Expect = 4.2
Identities = 12/44 (27%), Positives = 21/44 (47%)
Frame = +2
Query: 39 QQKIIEALEAEKDSLTKTNKSIRNTLWDWKQQLYAEAASDSAPV 82
+++ +++ K + K + R WDW+Q AE S S V
Sbjct: 191 RRRFHDSIIHRKQNNNKKQQQQREFTWDWEQPREAEEESSSKKV 322
>TC218146
Length = 509
Score = 25.4 bits (54), Expect = 4.2
Identities = 8/16 (50%), Positives = 12/16 (75%)
Frame = -3
Query: 81 PVPLARLYEIYGEAAP 96
P PLA ++ +YGE +P
Sbjct: 300 PTPLAHIFHLYGEKSP 253
>BM308349
Length = 429
Score = 25.0 bits (53), Expect = 5.5
Identities = 14/46 (30%), Positives = 24/46 (51%)
Frame = -3
Query: 15 SENNTVINVGLGAVFAILGARSYNQQKIIEALEAEKDSLTKTNKSI 60
S+ + + G F ILGA S + I++ ++ + L K +KSI
Sbjct: 259 SKKTKCL*ISKGKNFTILGAPSISNFIILKKIKKKNPKLNKWHKSI 122
>TC211327 similar to UP|Q94JT2 (Q94JT2) AT5g17630/K10A8_110, partial (72%)
Length = 1224
Score = 25.0 bits (53), Expect = 5.5
Identities = 16/38 (42%), Positives = 20/38 (52%), Gaps = 3/38 (7%)
Frame = +1
Query: 24 GLGAVFAILGARSYNQ---QKIIEALEAEKDSLTKTNK 58
GLG+ AILG Y+Q +K +E EK S K K
Sbjct: 916 GLGSAIAILGTFLYSQATSKKKAMKIEGEKSS*RKRRK 1029
>TC207542 similar to UP|Q6H976 (Q6H976) STYLOSA protein, partial (18%)
Length = 670
Score = 25.0 bits (53), Expect = 5.5
Identities = 16/52 (30%), Positives = 24/52 (45%), Gaps = 1/52 (1%)
Frame = +1
Query: 32 LGARSYNQQKIIEA-LEAEKDSLTKTNKSIRNTLWDWKQQLYAEAASDSAPV 82
L A + N I++ +A + SL KSI + WD + A + DS V
Sbjct: 70 LAAAAENVVSILDVETQASRYSLKGHTKSIHSVCWDPSGEFLASVSEDSVRV 225
>AW666321
Length = 422
Score = 25.0 bits (53), Expect = 5.5
Identities = 10/22 (45%), Positives = 13/22 (58%)
Frame = +3
Query: 46 LEAEKDSLTKTNKSIRNTLWDW 67
+ + S +TN SIRNT W W
Sbjct: 42 IRGRQGSRWETNISIRNTCWCW 107
>TC210857 similar to UP|Q9M7I2 (Q9M7I2) WD-repeat cell cycle regulatory
protein, partial (5%)
Length = 462
Score = 24.3 bits (51), Expect = 9.4
Identities = 11/28 (39%), Positives = 15/28 (53%)
Frame = -1
Query: 29 FAILGARSYNQQKIIEALEAEKDSLTKT 56
+AIL + YN ++I E K LT T
Sbjct: 372 YAILNSPKYNARRITEKNNGRKPKLTST 289
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.316 0.131 0.369
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,140,253
Number of Sequences: 63676
Number of extensions: 39137
Number of successful extensions: 215
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 214
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 215
length of query: 112
length of database: 12,639,632
effective HSP length: 88
effective length of query: 24
effective length of database: 7,036,144
effective search space: 168867456
effective search space used: 168867456
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 51 (24.3 bits)
Medicago: description of AC144807.5


