

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144766.6 - phase: 0
(381 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
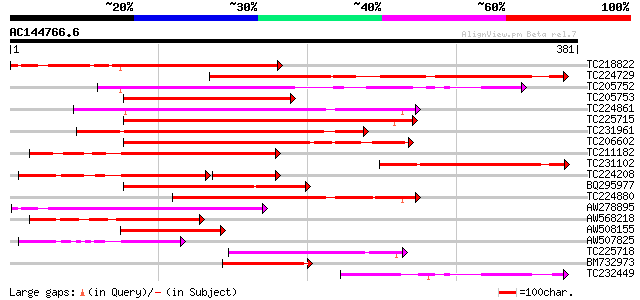
Score E
Sequences producing significant alignments: (bits) Value
TC218822 295 2e-80
TC224729 249 2e-66
TC205752 247 7e-66
TC205753 211 6e-55
TC224861 similar to UP|Q9LUZ4 (Q9LUZ4) Zinc finger transcription... 203 1e-52
TC225715 similar to GB|AAM16218.1|20334714|AY093957 At2g41900/T6... 201 5e-52
TC231961 homologue to GB|AAM16218.1|20334714|AY093957 At2g41900/... 200 1e-51
TC206602 similar to UP|Q9LUZ4 (Q9LUZ4) Zinc finger transcription... 200 1e-51
TC211182 similar to GB|AAO63406.1|28950965|BT005342 At1g03790 {A... 191 5e-49
TC231102 189 2e-48
TC224208 similar to GB|AAO63406.1|28950965|BT005342 At1g03790 {A... 127 2e-48
BQ295977 similar to GP|8843784|dbj zinc finger transcription fac... 181 6e-46
TC224880 similar to UP|Q9LUZ4 (Q9LUZ4) Zinc finger transcription... 149 2e-36
AW278895 123 2e-28
AW568218 111 5e-25
AW508155 102 3e-22
AW507825 96 3e-20
TC225718 similar to GB|AAM16218.1|20334714|AY093957 At2g41900/T6... 85 5e-17
BM732973 similar to GP|14335106|gb| At2g41900/T6D20.20 {Arabidop... 80 1e-15
TC232449 68 8e-12
>TC218822
Length = 554
Score = 295 bits (756), Expect = 2e-80
Identities = 140/186 (75%), Positives = 156/186 (83%), Gaps = 3/186 (1%)
Frame = +3
Query: 1 MMLGEPPHRTNPTVHVPPWPTLNNPTAEIFSPLTSNDDYSQFYMQEALSAFQHYVNENND 60
MMLGE HR NPTVHVPPW EIFSP T N DYS + MQEALSA QHY E+ D
Sbjct: 27 MMLGET-HRPNPTVHVPPW------APEIFSPYTGNADYSPYSMQEALSALQHY--ESTD 179
Query: 61 SDSDSEIFPTHES---VDSYSNDHFRMFEFKIRRCARGRSHDWTECPFSHPGEKARRRDP 117
++SDSE+ P+ E VD+YS DHFRMFEFK+RRCAR RSHDWT+CP++HPGEKARRRDP
Sbjct: 180 AESDSEV-PSREPEVPVDAYSCDHFRMFEFKVRRCARCRSHDWTDCPYAHPGEKARRRDP 356
Query: 118 RKYNYSGTSCPDFRKGSCKKGDSCEFAHGVFECWLHPSRYRTQPCKDGTSCRRPVCFFAH 177
RKY+YSGT+CPDFRKGSCKKGD+CE+AHGVFECWLHP+RYRTQPCKDGTSCRR VCFFAH
Sbjct: 357 RKYHYSGTACPDFRKGSCKKGDACEYAHGVFECWLHPARYRTQPCKDGTSCRRCVCFFAH 536
Query: 178 TTEQLR 183
T EQLR
Sbjct: 537 TPEQLR 554
>TC224729
Length = 1018
Score = 249 bits (635), Expect = 2e-66
Identities = 142/242 (58%), Positives = 155/242 (63%), Gaps = 1/242 (0%)
Frame = +2
Query: 135 CKKGDSCEFAHGVFECWLHPSRYRTQPCKDGTSCRRPVCFFAHTTEQLRAPTQQSPRSVP 194
C+ G SC F H F CWLH RYRT CKD TSCRR VCFFAHT EQLR QSPRSV
Sbjct: 11 CRIGYSCYFFHCFFYCWLHSVRYRTHSCKDCTSCRRRVCFFAHTPEQLRVLPMQSPRSVA 190
Query: 195 -SVDSYDGSPLRLAFESSCVKTLQFMSSPGSVSPPVESPPMSPMTRSLGRSVGSSSVNEM 253
S +SYDGSP+R SS FMSSP + P ESPP SVNEM
Sbjct: 191 NSSESYDGSPMRQVSLSSAAAAA-FMSSPAASLSPPESPP---------------SVNEM 322
Query: 254 VASLRNLQLGTMKSLPSSWNVQMGSPRFGSPRGPVIRPGFCSLPSTPTQVPSRGRVNHFD 313
VASLRNLQLG MKS+P + NV +GSPR G V+RPGF SLP+TPTQ P R V FD
Sbjct: 323 VASLRNLQLGKMKSMPHNRNVSVGSPR-----GSVLRPGFLSLPTTPTQQPVRSGVKCFD 487
Query: 314 LWDQSCEEEPVMERVESGRDIRVKMFEKLSKENSFNGSGMGSGSGLGEVVEDPDVGWVSE 373
+WD+S EEEPVMERVESGR IR KMFEKLSKENS + S PD+GWVSE
Sbjct: 488 VWDESFEEEPVMERVESGRGIRAKMFEKLSKENSLDAS-----------ASPPDLGWVSE 634
Query: 374 LV 375
LV
Sbjct: 635 LV 640
>TC205752
Length = 1020
Score = 247 bits (630), Expect = 7e-66
Identities = 138/293 (47%), Positives = 166/293 (56%), Gaps = 5/293 (1%)
Frame = +1
Query: 60 DSDSDSEIFPTHES----VDSYSNDHFRMFEFKIRRCARGRSHDWTECPFSHPGEKARRR 115
D+D + P+ S +S+DHFRMF+FK+R C RGRSHDWTECP++HP EKARRR
Sbjct: 31 DADGGGALSPSISSNADTCSLFSSDHFRMFQFKVRNCPRGRSHDWTECPYAHPAEKARRR 210
Query: 116 DPRKYNYSGTSCPDFRKGSCKKGDSCEFAHGVFECWLHPSRYRTQPCKDGTSCRRPVCFF 175
DPRKY+YSGTSCPD+RKG+CK+GD+C+FAHGVFECWLHPSRYRTQ CKDGT+CRR VCFF
Sbjct: 211 DPRKYHYSGTSCPDYRKGNCKRGDTCQFAHGVFECWLHPSRYRTQLCKDGTNCRRRVCFF 390
Query: 176 AHTTEQLRAPTQQSPRSVPSVDSYDGSPLRLAFESSCVKTLQFMSSPGSVSPPVESPPMS 235
AHT++QLR T S S F+SSP SV
Sbjct: 391 AHTSDQLRVVTDSSSSS-------------------------FVSSPTSV---------- 465
Query: 236 PMTRSLGRSVGSSSVNEMVASLRNL-QLGTMKSLPSSWNVQMGSPRFGSPRGPVIRPGFC 294
+ S SS V + L + G+ L FGSPRG GF
Sbjct: 466 ----LINSSFSDSSPYSFVRGIVKLDEEGSYSGLTRC--------VFGSPRG-----GFM 594
Query: 295 SLPSTPTQVPSRGRVNHFDLWDQSCEEEPVMERVESGRDIRVKMFEKLSKENS 347
S+ CEEEP MERVESGRDIR +++ KLS+ENS
Sbjct: 595 SV--------------------NGCEEEPAMERVESGRDIRARIYAKLSRENS 693
>TC205753
Length = 457
Score = 211 bits (536), Expect = 6e-55
Identities = 87/116 (75%), Positives = 103/116 (88%)
Frame = +3
Query: 77 YSNDHFRMFEFKIRRCARGRSHDWTECPFSHPGEKARRRDPRKYNYSGTSCPDFRKGSCK 136
+S+DHFRMF+FK+R C RGRSHDWTECP++HP EKARRRDPRKY+YSGT+CPD++KG+CK
Sbjct: 108 FSSDHFRMFQFKVRICPRGRSHDWTECPYAHPAEKARRRDPRKYHYSGTACPDYQKGNCK 287
Query: 137 KGDSCEFAHGVFECWLHPSRYRTQPCKDGTSCRRPVCFFAHTTEQLRAPTQQSPRS 192
+GD+C+F+HGVFECWLHPSRYRT CKDGT+CRR VCFFAHTTEQLR S S
Sbjct: 288 RGDTCQFSHGVFECWLHPSRYRTHLCKDGTTCRRRVCFFAHTTEQLRLVVTDSSSS 455
>TC224861 similar to UP|Q9LUZ4 (Q9LUZ4) Zinc finger transcription factor-like
protein (AT5g58620/mzn1_70), partial (42%)
Length = 2560
Score = 203 bits (516), Expect = 1e-52
Identities = 104/240 (43%), Positives = 141/240 (58%), Gaps = 7/240 (2%)
Frame = +1
Query: 44 MQEALSAFQHYVNENNDSDSDSEIFPTHESVDS-----YSNDHFRMFEFKIRRCARGRSH 98
M + + V E + + + +P S+ Y D FRM+ FK++ C+R SH
Sbjct: 664 MSQEMELQMFSVPEKKEGSDNKKEYPVDISLPDINNGVYGTDEFRMYNFKVKPCSRAYSH 843
Query: 99 DWTECPFSHPGEKARRRDPRKYNYSGTSCPDFRKGSCKKGDSCEFAHGVFECWLHPSRYR 158
DWTECPF HPGE ARRRDPRKY YS CP+FRKG+C+KGDSCE+AHGVFE WLHP++YR
Sbjct: 844 DWTECPFVHPGENARRRDPRKYPYSCVPCPEFRKGTCQKGDSCEYAHGVFESWLHPAQYR 1023
Query: 159 TQPCKDGTSCRRPVCFFAHTTEQLRAPTQQSPRSVPSVDSYDGSPLRLAFESSCVKTLQF 218
T+ CKD T C R VCFFAH E+LR + ++PS SY S L + S
Sbjct: 1024TRLCKDETGCARKVCFFAHKPEELRPVYASTGSAMPSPKSYSASGLDMTAMSPLA----- 1188
Query: 219 MSSPGSVSPPVESPPMSPMTRSLGRSVGSSSVNEMVASLRNLQL--GTMKSLPSSWNVQM 276
+SS P V +PPMSP+ + GS N++ + +LQL +K+ S+ +++M
Sbjct: 1189LSSTSLPMPTVSTPPMSPLAAASSPKSGSMWQNKINLTPPSLQLPGSRLKAALSARDLEM 1368
>TC225715 similar to GB|AAM16218.1|20334714|AY093957 At2g41900/T6D20.20
{Arabidopsis thaliana;} , partial (42%)
Length = 2142
Score = 201 bits (511), Expect = 5e-52
Identities = 99/200 (49%), Positives = 125/200 (62%), Gaps = 2/200 (1%)
Frame = +3
Query: 77 YSNDHFRMFEFKIRRCARGRSHDWTECPFSHPGEKARRRDPRKYNYSGTSCPDFRKGSCK 136
Y+ D FRMF FK+R C+R SHDWTECPF HPGE ARRRDPRK++YS CPDFRKG+C+
Sbjct: 66 YATDEFRMFSFKVRPCSRAYSHDWTECPFVHPGENARRRDPRKFHYSCVPCPDFRKGACR 245
Query: 137 KGDSCEFAHGVFECWLHPSRYRTQPCKDGTSCRRPVCFFAHTTEQLRAPTQQSPRSVPSV 196
+GD CE+AHGVFECWLHP++YRT+ CKDGTSC R VCFFAHT E+LR + + PS
Sbjct: 246 RGDMCEYAHGVFECWLHPAQYRTRLCKDGTSCNRRVCFFAHTAEELRPLYVSTGSAAPSP 425
Query: 197 DSYDGSPLRLAFESSCVKTLQFMSSPGSVSPPVESPPMSPMTRSLGRSVGSSSVNEMVAS 256
S P + ++ SS S+SP PMSP + S + N
Sbjct: 426 RSSASGPNVMDMAAAMSLFPGSPSSGSSMSPSHFGQPMSPSANGMPLSSAWAQPNVPALH 605
Query: 257 L--RNLQLGTMKSLPSSWNV 274
L NLQ ++S S+ ++
Sbjct: 606 LPGSNLQSSRLRSSLSARDI 665
>TC231961 homologue to GB|AAM16218.1|20334714|AY093957 At2g41900/T6D20.20
{Arabidopsis thaliana;} , partial (25%)
Length = 893
Score = 200 bits (508), Expect = 1e-51
Identities = 98/196 (50%), Positives = 126/196 (64%)
Frame = +2
Query: 46 EALSAFQHYVNENNDSDSDSEIFPTHESVDSYSNDHFRMFEFKIRRCARGRSHDWTECPF 105
E L + V+E + D + S+ YS+D FRM+ FK+R C+R SHDWTECPF
Sbjct: 86 EDLKMKTNEVSEKKEYPVDLSLPDIKNSI--YSSDEFRMYSFKVRPCSRAYSHDWTECPF 259
Query: 106 SHPGEKARRRDPRKYNYSGTSCPDFRKGSCKKGDSCEFAHGVFECWLHPSRYRTQPCKDG 165
HPGE ARRRDPRK++YS CP+FRKG+C++GD CE+AHGVFECWLHP++YRT+ CKDG
Sbjct: 260 VHPGENARRRDPRKFHYSCVPCPEFRKGACRRGDMCEYAHGVFECWLHPAQYRTRLCKDG 439
Query: 166 TSCRRPVCFFAHTTEQLRAPTQQSPRSVPSVDSYDGSPLRLAFESSCVKTLQFMSSPGSV 225
T+C R VCFFAHT E+LR + +VPS S S + S SS +
Sbjct: 440 TNCARRVCFFAHTNEELRPLYVSTGSAVPSPRSSASSAMDFVAAIS-------PSSMSVM 598
Query: 226 SPPVESPPMSPMTRSL 241
SP +PPMSP + S+
Sbjct: 599 SPSPFTPPMSPSSASI 646
>TC206602 similar to UP|Q9LUZ4 (Q9LUZ4) Zinc finger transcription factor-like
protein (AT5g58620/mzn1_70), partial (31%)
Length = 1855
Score = 200 bits (508), Expect = 1e-51
Identities = 97/195 (49%), Positives = 125/195 (63%)
Frame = +1
Query: 77 YSNDHFRMFEFKIRRCARGRSHDWTECPFSHPGEKARRRDPRKYNYSGTSCPDFRKGSCK 136
Y D FRM+ FK++ C+R SHDWTECPF HPGE ARRRDPRKY+YS CP+FRKGSC
Sbjct: 67 YGTDEFRMYTFKVKPCSRAYSHDWTECPFVHPGENARRRDPRKYHYSCVPCPEFRKGSCS 246
Query: 137 KGDSCEFAHGVFECWLHPSRYRTQPCKDGTSCRRPVCFFAHTTEQLRAPTQQSPRSVPSV 196
KGD+CE+AHG+FECWLHP++YRT+ CKD C R VCFFAH E+LR + ++PS
Sbjct: 247 KGDACEYAHGIFECWLHPAQYRTRLCKDEGGCTRRVCFFAHKLEELRPLYASTGSAIPSP 426
Query: 197 DSYDGSPLRLAFESSCVKTLQFMSSPGSVSPPVESPPMSPMTRSLGRSVGSSSVNEMVAS 256
SY S A E V + + SP + PP +PP++P S SS + + S
Sbjct: 427 RSYSAS--ASALEMGSVNPIA-LGSPSVLMPPTSTPPLTP-------SGASSPIAGSMWS 576
Query: 257 LRNLQLGTMKSLPSS 271
N+ + T++ LP S
Sbjct: 577 QSNVSVPTLQ-LPKS 618
>TC211182 similar to GB|AAO63406.1|28950965|BT005342 At1g03790 {Arabidopsis
thaliana;} , partial (27%)
Length = 700
Score = 191 bits (485), Expect = 5e-49
Identities = 88/169 (52%), Positives = 112/169 (66%)
Frame = +2
Query: 14 VHVPPWPTLNNPTAEIFSPLTSNDDYSQFYMQEALSAFQHYVNENNDSDSDSEIFPTHES 73
+ VPP L +A + +D Y+ + S + + D DSD
Sbjct: 143 IPVPPRKLLTRRSAAV------HDGSGDTYLPHSGST-----DSSTDDDSDG-------- 265
Query: 74 VDSYSNDHFRMFEFKIRRCARGRSHDWTECPFSHPGEKARRRDPRKYNYSGTSCPDFRKG 133
D Y++D FRMFEFK+RRC+R RSHDWT+CPF HPGEKARRRDPR+++YS T CP+FR+G
Sbjct: 266 -DPYASDQFRMFEFKVRRCSRSRSHDWTDCPFVHPGEKARRRDPRRFHYSATVCPEFRRG 442
Query: 134 SCKKGDSCEFAHGVFECWLHPSRYRTQPCKDGTSCRRPVCFFAHTTEQL 182
C +GD+CEF+HGVFECWLHPSRYRT+ CKDG +C+R VCFF QL
Sbjct: 443 QCDRGDACEFSHGVFECWLHPSRYRTEACKDGKNCKRKVCFFXTPLGQL 589
>TC231102
Length = 761
Score = 189 bits (479), Expect = 2e-48
Identities = 96/128 (75%), Positives = 105/128 (82%)
Frame = +3
Query: 249 SVNEMVASLRNLQLGTMKSLPSSWNVQMGSPRFGSPRGPVIRPGFCSLPSTPTQVPSRGR 308
SVNEMVASLRNLQLG +KSLPSSWNV MG FGSPRGP+IRPGF SLP+TP Q P+RG
Sbjct: 12 SVNEMVASLRNLQLGKVKSLPSSWNV-MGPSGFGSPRGPMIRPGFFSLPTTPAQAPTRGG 188
Query: 309 VNHFDLWDQSCEEEPVMERVESGRDIRVKMFEKLSKENSFNGSGMGSGSGLGEVVEDPDV 368
VN+FD WDQSCEEEPVMERVESGR+IR +MF KLSKEN +GS GSG + DPDV
Sbjct: 189 VNYFDKWDQSCEEEPVMERVESGRNIRARMFAKLSKENHLDGSDSGSGQ-----IGDPDV 353
Query: 369 GWVSELVS 376
GWVSELVS
Sbjct: 354 GWVSELVS 377
>TC224208 similar to GB|AAO63406.1|28950965|BT005342 At1g03790 {Arabidopsis
thaliana;} , partial (28%)
Length = 884
Score = 127 bits (318), Expect(2) = 2e-48
Identities = 62/129 (48%), Positives = 79/129 (61%)
Frame = +2
Query: 7 PHRTNPTVHVPPWPTLNNPTAEIFSPLTSNDDYSQFYMQEALSAFQHYVNENNDSDSDSE 66
P +T + +PP L A+ D YS+ MQ+ +NDSD D
Sbjct: 131 PKKTLREIDIPPRKLLTRRAAKCVG---GTDVYSEETMQQKFLP-------SNDSDED-- 274
Query: 67 IFPTHESVDSYSNDHFRMFEFKIRRCARGRSHDWTECPFSHPGEKARRRDPRKYNYSGTS 126
D YS+DHFRMFEFK+R+C R RSHDWT+CPF+HPGEKARRRDPR+Y+YSGT
Sbjct: 275 --------DPYSSDHFRMFEFKVRQCTRSRSHDWTDCPFAHPGEKARRRDPRRYHYSGTV 430
Query: 127 CPDFRKGSC 135
CP++R+G C
Sbjct: 431 CPEYRRGGC 457
Score = 83.6 bits (205), Expect(2) = 2e-48
Identities = 32/46 (69%), Positives = 39/46 (84%)
Frame = +3
Query: 137 KGDSCEFAHGVFECWLHPSRYRTQPCKDGTSCRRPVCFFAHTTEQL 182
+ D+CE+AHGVFECWLHP RYRT+ CKDG +C+R VCFFAHT Q+
Sbjct: 462 RDDACEYAHGVFECWLHPFRYRTEACKDGRNCKRKVCFFAHTPXQV 599
>BQ295977 similar to GP|8843784|dbj zinc finger transcription factor-like
protein {Arabidopsis thaliana}, partial (22%)
Length = 422
Score = 181 bits (458), Expect = 6e-46
Identities = 77/126 (61%), Positives = 95/126 (75%)
Frame = +3
Query: 77 YSNDHFRMFEFKIRRCARGRSHDWTECPFSHPGEKARRRDPRKYNYSGTSCPDFRKGSCK 136
YS+D FRM+ FK+R C+R SHDWTECPF HPGE ARRRDPR+Y YS CP+FRKGSC
Sbjct: 42 YSSDEFRMYTFKVRPCSRAYSHDWTECPFVHPGENARRRDPRRYQYSCVPCPEFRKGSCS 221
Query: 137 KGDSCEFAHGVFECWLHPSRYRTQPCKDGTSCRRPVCFFAHTTEQLRAPTQQSPRSVPSV 196
KGD+C++AHG+FECWLHP++Y+T+ CK+ T C R VCFFAH E LR + ++PS
Sbjct: 222 KGDACDYAHGIFECWLHPAQYKTRLCKE-TGCTRRVCFFAHNVEDLRPVYASTGSAMPSP 398
Query: 197 DSYDGS 202
SY S
Sbjct: 399 RSYSVS 416
>TC224880 similar to UP|Q9LUZ4 (Q9LUZ4) Zinc finger transcription factor-like
protein (AT5g58620/mzn1_70), partial (21%)
Length = 1723
Score = 149 bits (377), Expect = 2e-36
Identities = 81/169 (47%), Positives = 107/169 (62%), Gaps = 2/169 (1%)
Frame = +3
Query: 110 EKARRRDPRKYNYSGTSCPDFRKGSCKKGDSCEFAHGVFECWLHPSRYRTQPCKDGTSCR 169
E ARRRDPRKY YS CP+FRKG+C+KGDSCE+AHGVFE WLHP++YRT+ CKD T C
Sbjct: 9 ENARRRDPRKYPYSCVPCPEFRKGTCQKGDSCEYAHGVFESWLHPAQYRTRLCKDETGCA 188
Query: 170 RPVCFFAHTTEQLRAPTQQSPRSVPSVDSYDGSPLRLAFESSCVKTLQFMSSPGSVSPPV 229
R VCFFAH E+LR + ++PS SY S L + S +SS P V
Sbjct: 189 RKVCFFAHKPEELRPVYASTGSAMPSPKSYSASGLDMTAMSPLA-----LSSTSLPMPTV 353
Query: 230 ESPPMSPMTRSLGRSVGSSSVNEMVASLRNLQL--GTMKSLPSSWNVQM 276
+PPMSP+T S +S GS N++ + +LQL +K+ S+ +++M
Sbjct: 354 STPPMSPLTASSPKS-GSLWQNKINLTPPSLQLPGSRLKAALSARDLEM 497
>AW278895
Length = 496
Score = 123 bits (308), Expect = 2e-28
Identities = 69/172 (40%), Positives = 97/172 (56%)
Frame = +2
Query: 2 MLGEPPHRTNPTVHVPPWPTLNNPTAEIFSPLTSNDDYSQFYMQEALSAFQHYVNENNDS 61
MLGE HR N TVH+P W EIFSP T++ DYS + +++ LSA + + + S
Sbjct: 2 MLGET-HRPNATVHMPAW------APEIFSPYTNDADYSPYSVRKTLSALEDCESTDAKS 160
Query: 62 DSDSEIFPTHESVDSYSNDHFRMFEFKIRRCARGRSHDWTECPFSHPGEKARRRDPRKYN 121
DS+ VD+YS DH RMFE K+ RC RSH CP++ P +KAR +DPR Y+
Sbjct: 161 DSEVPCSKAEGPVDTYSYDHCRMFECKVSRCTXYRSHK*DRCPYARPHDKARGQDPRDYH 340
Query: 122 YSGTSCPDFRKGSCKKGDSCEFAHGVFECWLHPSRYRTQPCKDGTSCRRPVC 173
YSG P + + S K+ ++CE + E + P RT+ C+ T + VC
Sbjct: 341 YSGRXEPGWPEESGKRSNACE*SAQSVE*LVDPRHGRTEACEK*TEGGQHVC 496
>AW568218
Length = 427
Score = 111 bits (278), Expect = 5e-25
Identities = 55/118 (46%), Positives = 73/118 (61%)
Frame = +3
Query: 14 VHVPPWPTLNNPTAEIFSPLTSNDDYSQFYMQEALSAFQHYVNENNDSDSDSEIFPTHES 73
+ VPP L +A + S D +S ++ Y+ N +DS + +S
Sbjct: 111 IPVPPRKLLTRRSAAVHD--VSGDMFSDDTLRHK------YLPHNGSTDSSDD-----DS 251
Query: 74 VDSYSNDHFRMFEFKIRRCARGRSHDWTECPFSHPGEKARRRDPRKYNYSGTSCPDFR 131
D Y++D FRMFEFK+RRC R RSHDWT+CPF HPGEKARRRDPR+++YS T CP+FR
Sbjct: 252 GDPYASDQFRMFEFKVRRCTRSRSHDWTDCPFVHPGEKARRRDPRRFHYSATVCPEFR 425
>AW508155
Length = 220
Score = 102 bits (254), Expect = 3e-22
Identities = 41/71 (57%), Positives = 52/71 (72%)
Frame = +2
Query: 75 DSYSNDHFRMFEFKIRRCARGRSHDWTECPFSHPGEKARRRDPRKYNYSGTSCPDFRKGS 134
D Y++D FRMFEFK+RR R RSHDWT+ PF H GE ARRRDPR+ Y T CP+F +
Sbjct: 8 DPYASDQFRMFEFKVRRLTRSRSHDWTD*PFVHTGENARRRDPRRIQYYATGCPEFLRSQ 187
Query: 135 CKKGDSCEFAH 145
C +G++CE +H
Sbjct: 188 CDRGNACEISH 220
>AW507825
Length = 375
Score = 95.9 bits (237), Expect = 3e-20
Identities = 53/112 (47%), Positives = 65/112 (57%)
Frame = +1
Query: 7 PHRTNPTVHVPPWPTLNNPTAEIFSPLTSNDDYSQFYMQEALSAFQHYVNENNDSDSDSE 66
P +T + +PP L A D YS +E + Q ++ +NDSD D
Sbjct: 97 PKKTLREIDIPPRKLLTRRAAACVG---GTDVYSS---EETM--LQKFL-PSNDSDED-- 243
Query: 67 IFPTHESVDSYSNDHFRMFEFKIRRCARGRSHDWTECPFSHPGEKARRRDPR 118
D YS+DHFRMFEFK+RRC R RSHDWT+CPF+HPGEKARRRDPR
Sbjct: 244 --------DPYSSDHFRMFEFKVRRCTRSRSHDWTDCPFAHPGEKARRRDPR 375
>TC225718 similar to GB|AAM16218.1|20334714|AY093957 At2g41900/T6D20.20
{Arabidopsis thaliana;} , partial (38%)
Length = 1643
Score = 85.1 bits (209), Expect = 5e-17
Identities = 51/124 (41%), Positives = 68/124 (54%), Gaps = 4/124 (3%)
Frame = +2
Query: 148 FECWLHPSRYRTQPCKDGTSCRRPVCFFAHTTEQLRAPTQQSPRSVPSVDSYDGSPLRLA 207
FECWLHP++YRT+ CKDGTSC R VCFFAHT E+LR + +VPS S +P +
Sbjct: 2 FECWLHPAQYRTRLCKDGTSCNRRVCFFAHTAEELRPLYVSTGSAVPSPRSSASAPNVMD 181
Query: 208 FESSCVKTLQFMSSPGSVSPPVESPPMSPMTRSLGRSVGSSSVNEMVASLR----NLQLG 263
++ SS S+SP PMSP G S+ S+ V++L NLQ
Sbjct: 182 MAAAMSLLPGSPSSVSSMSPSHFGQPMSPSAN--GMSLSSAWAQPNVSALHLPGSNLQSS 355
Query: 264 TMKS 267
++S
Sbjct: 356 RLRS 367
>BM732973 similar to GP|14335106|gb| At2g41900/T6D20.20 {Arabidopsis
thaliana}, partial (12%)
Length = 421
Score = 80.5 bits (197), Expect = 1e-15
Identities = 37/60 (61%), Positives = 45/60 (74%)
Frame = +2
Query: 144 AHGVFECWLHPSRYRTQPCKDGTSCRRPVCFFAHTTEQLRAPTQQSPRSVPSVDSYDGSP 203
AHGVFECWLHP++YRT+ CKDGT+C R VCFFAHT E+LR + +VPS S G+P
Sbjct: 2 AHGVFECWLHPAQYRTRLCKDGTNCARRVCFFAHTNEELRPLYVSTGSAVPSPRS--GAP 175
>TC232449
Length = 524
Score = 67.8 bits (164), Expect = 8e-12
Identities = 56/155 (36%), Positives = 69/155 (44%), Gaps = 2/155 (1%)
Frame = +1
Query: 223 GSVSPPVESPPMSPMTRSLGRSVGSSSVNEMVASLRNLQLGTMKSLPSSWNVQMGSPR-- 280
G+ S V SP S S S V E+V S RN+++ G R
Sbjct: 7 GTRSXFVSSPTSVLNYSSFSDSSPYSFVREIVNSTRNVKIE-----------DSGLTRCV 153
Query: 281 FGSPRGPVIRPGFCSLPSTPTQVPSRGRVNHFDLWDQSCEEEPVMERVESGRDIRVKMFE 340
FGSPRG GF S+ CEEEP MERVESG+DIR +++
Sbjct: 154 FGSPRG-----GFLSV--------------------NGCEEEPAMERVESGKDIRARIYA 258
Query: 341 KLSKENSFNGSGMGSGSGLGEVVEDPDVGWVSELV 375
KLS+EN+ S DPD+GWVSELV
Sbjct: 259 KLSRENTIGAS-------------DPDIGWVSELV 324
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.316 0.133 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,750,758
Number of Sequences: 63676
Number of extensions: 368121
Number of successful extensions: 3008
Number of sequences better than 10.0: 109
Number of HSP's better than 10.0 without gapping: 2706
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2943
length of query: 381
length of database: 12,639,632
effective HSP length: 99
effective length of query: 282
effective length of database: 6,335,708
effective search space: 1786669656
effective search space used: 1786669656
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 60 (27.7 bits)
Medicago: description of AC144766.6


