

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144765.5 - phase: 0 /pseudo
(443 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
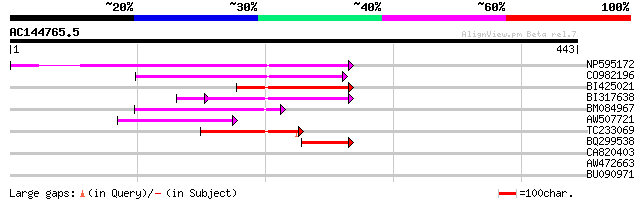
Score E
Sequences producing significant alignments: (bits) Value
NP595172 polyprotein [Glycine max] 169 3e-42
CO982196 125 5e-29
BI425021 87 2e-17
BI317638 weakly similar to GP|9294238|dbj| contains similarity t... 84 2e-17
BM084967 85 6e-17
AW507721 weakly similar to GP|27764548|gb polyprotein {Glycine m... 72 7e-13
TC233069 69 6e-12
BQ299538 50 2e-06
CA820403 weakly similar to GP|13273463|gb| pol protein integrase... 32 0.75
AW472663 31 0.98
BU090971 28 8.3
>NP595172 polyprotein [Glycine max]
Length = 4659
Score = 169 bits (427), Expect = 3e-42
Identities = 98/268 (36%), Positives = 139/268 (51%)
Frame = +1
Query: 1 NFGLNYHPDKAKVVADALSRKTLHMSALMVKEFELLEQFRDLSLVCELSSQSVQLGMLKI 60
+F + Y P K ADALSR + L +
Sbjct: 2959 DFKIEYKPGKDNQAADALSRMFM-------------------------------LAWSEP 3045
Query: 61 NSDFLGSIREAQQVDVKFVDLMVVSNQAEESDFKVDEQGVLRFRGRICIPDNEELKKLIL 120
+S FL +R D LM Q ++ +G+L ++ R+ IP EE+ IL
Sbjct: 3046 HSIFLEELRARLISDPHLKQLMETYKQGADASHYTVREGLLYWKDRVVIPAEEEIVNKIL 3225
Query: 121 EEGHKSNLSIHLGATKMYQDLKKLFWWSGLKKDVARFVYACLTCQKSKVEHQRPAGLLTP 180
+E H S + H G T+ LK F+W +++DV ++ CL CQ++K + PAGLL P
Sbjct: 3226 QEYHSSPIGGHAGITRTLARLKAQFYWPKMQEDVKAYIQKCLICQQAKSNNTLPAGLLQP 3405
Query: 181 LDVPEWKWDSISMDFVSSLPNTSRGHDSIWVVVDRLTKSAHFIPINISYPVAQLAEIYIQ 240
L +P+ W+ ++MDF++ LPN S G I VV+DRLTK AHFIP+ Y +AE ++
Sbjct: 3406 LPIPQQVWEDVAMDFITGLPN-SFGLSVIMVVIDRLTKYAHFIPLKADYNSKVVAEAFMS 3582
Query: 241 NIVKLHGVPSSIVSDRDPRFTSRFWRSL 268
+IVKLHG+P SIVSDRD FTS FW+ L
Sbjct: 3583 HIVKLHGIPRSIVSDRDRVFTSTFWQHL 3666
>CO982196
Length = 812
Score = 125 bits (313), Expect = 5e-29
Identities = 67/166 (40%), Positives = 98/166 (58%)
Frame = +1
Query: 99 GVLRFRGRICIPDNEELKKLILEEGHKSNLSIHLGATKMYQDLKKLFWWSGLKKDVARFV 158
G L F+ R+ + N L+L+E S L H G + ++ + + +W G+KK +V
Sbjct: 316 GKLYFKDRLVLSKNSTKIPLLLKELQDSPLGGHSGFFRTFKRVANVVFWQGMKKTTRDYV 495
Query: 159 YACLTCQKSKVEHQRPAGLLTPLDVPEWKWDSISMDFVSSLPNTSRGHDSIWVVVDRLTK 218
AC C+++K PAGLL L +P W ISMDF+ LP ++G D+I VVVDRLTK
Sbjct: 496 AACEICRRNKTSTLSPAGLL*LLPIPTKVWTDISMDFIGGLPK-AQGKDNILVVVDRLTK 672
Query: 219 SAHFIPINISYPVAQLAEIYIQNIVKLHGVPSSIVSDRDPRFTSRF 264
AHF ++ Y ++AE++I+ +V+LHG P+SIVSD F S F
Sbjct: 673 YAHFFALSHPYTAKEVAELFIKELVRLHGFPASIVSDXXRLFMSLF 810
>BI425021
Length = 426
Score = 87.0 bits (214), Expect = 2e-17
Identities = 41/91 (45%), Positives = 58/91 (63%)
Frame = -1
Query: 178 LTPLDVPEWKWDSISMDFVSSLPNTSRGHDSIWVVVDRLTKSAHFIPINISYPVAQLAEI 237
L PL VP+ W+ +SMDF+ LP GH +I+VVV+R +K H + S+ +A +
Sbjct: 426 LCPLPVPQRPWEDLSMDFIVGLP-PYHGHTTIFVVVNRFSKGIHLGTLPTSHTAHMVASL 250
Query: 238 YIQNIVKLHGVPSSIVSDRDPRFTSRFWRSL 268
++ ++KLHG P SIVSDRDP F S FW+ L
Sbjct: 249 FLNIVIKLHGFPRSIVSDRDPLFISHFWQDL 157
>BI317638 weakly similar to GP|9294238|dbj| contains similarity to reverse
transcriptase~gene_id:K11J14.5 {Arabidopsis thaliana},
partial (5%)
Length = 420
Score = 84.3 bits (207), Expect(2) = 2e-17
Identities = 45/116 (38%), Positives = 67/116 (56%)
Frame = -2
Query: 153 DVARFVYACLTCQKSKVEHQRPAGLLTPLDVPEWKWDSISMDFVSSLPNTSRGHDSIWVV 212
+ A+ + CL CQ +K E +R LL PL VP W+ +S+DF++ L H +I VV
Sbjct: 350 ECAQMLPNCLDCQHTKYETKRIVDLLCPLLVPHRPWEDLSLDFITGLL-PYHVHTAILVV 174
Query: 213 VDRLTKSAHFIPINISYPVAQLAEIYIQNIVKLHGVPSSIVSDRDPRFTSRFWRSL 268
VD +K H + S+ +A ++I ++ KLHG+P S+VSD D F S FW+ L
Sbjct: 173 VDHFSKGIHLGMLPSSHTAHTVACLFIDSVAKLHGLPRSLVSDCDLLFVSHFWQEL 6
Score = 22.7 bits (47), Expect(2) = 2e-17
Identities = 9/26 (34%), Positives = 13/26 (49%)
Frame = -1
Query: 131 HLGATKMYQDLKKLFWWSGLKKDVAR 156
H G K L K +W G++ DV +
Sbjct: 405 HTGIAKTLA*LSKNIYWFGMRTDVTQ 328
>BM084967
Length = 426
Score = 85.1 bits (209), Expect = 6e-17
Identities = 48/118 (40%), Positives = 66/118 (55%)
Frame = -2
Query: 98 QGVLRFRGRICIPDNEELKKLILEEGHKSNLSIHLGATKMYQDLKKLFWWSGLKKDVARF 157
Q ++ G I +P +L E H S H+G TK L + F W G++KDV +F
Sbjct: 356 QDLILKNGCIWLPSGFSFIPTLLLEYHSSPTDAHIGVTKTMARLSENFTWIGIRKDVEQF 177
Query: 158 VYACLTCQKSKVEHQRPAGLLTPLDVPEWKWDSISMDFVSSLPNTSRGHDSIWVVVDR 215
V ACL CQ +K E Q+ AGLL PL VP W+ +S +F+ L + RG+ +I VVV R
Sbjct: 176 VAACLDCQYTKYEAQKMAGLLCPLPVPCRPWEDLSFNFIIGL-SEFRGYTAILVVVGR 6
>AW507721 weakly similar to GP|27764548|gb polyprotein {Glycine max}, partial
(3%)
Length = 464
Score = 71.6 bits (174), Expect = 7e-13
Identities = 35/95 (36%), Positives = 57/95 (59%), Gaps = 1/95 (1%)
Frame = +1
Query: 85 SNQAEESDFKVDEQGVLRFRGRICIPDNEELKKLILEEGHKSNLSIHLGATKMYQDLKKL 144
SN A +S V V+ ++GRI +P++ +L K+I+ E H S + H G T+ +
Sbjct: 178 SNNAGKSGDYVLHHDVIIWKGRIMLPNDSQLLKMIMTESHASKVGGHAGTTRTIVRINAQ 357
Query: 145 FWWSGLKKDVARFVYACLTCQKSKVEHQ-RPAGLL 178
F+W +++D+ +FV C+ CQ++KV H PAGLL
Sbjct: 358 FYWPKMREDIMKFVQECVICQQAKVTHSLLPAGLL 462
>TC233069
Length = 881
Score = 68.6 bits (166), Expect = 6e-12
Identities = 37/82 (45%), Positives = 50/82 (60%), Gaps = 2/82 (2%)
Frame = +2
Query: 150 LKKDVARFVYACLTCQKSKVEHQRPAGLLTPLDVPEWKWDSISMDFVSSLPNTSRGHDSI 209
+ +DV + AC CQ++K Q+ GLL PL +P W ISMDFV+ LP S G I
Sbjct: 224 MARDVCDHICACTNCQQNKYSTQK*FGLLQPLPIP*QVWKDISMDFVTHLP-PS*GKKMI 400
Query: 210 WVVVDRLTKSAHF--IPINISY 229
WV+VD TK +HF +P ++SY
Sbjct: 401 WVIVDCWTKYSHFLSLPAHLSY 466
>BQ299538
Length = 426
Score = 50.4 bits (119), Expect = 2e-06
Identities = 21/40 (52%), Positives = 28/40 (69%)
Frame = +1
Query: 229 YPVAQLAEIYIQNIVKLHGVPSSIVSDRDPRFTSRFWRSL 268
Y LAEI+ + +V LHGVP+S++SD DP F S FW+ L
Sbjct: 13 YSARVLAEIFTKEVVHLHGVPASVLSDEDPIFVSSFWKEL 132
>CA820403 weakly similar to GP|13273463|gb| pol protein integrase region
{Ginkgo biloba}, partial (52%)
Length = 421
Score = 31.6 bits (70), Expect = 0.75
Identities = 18/32 (56%), Positives = 20/32 (62%), Gaps = 1/32 (3%)
Frame = -3
Query: 238 YIQNIVKLHGVPSSIVSDRDPRF-TSRFWRSL 268
+I+ VKLHG SSIVSD D F S FW L
Sbjct: 335 FIKEAVKLHGCSSSIVSDWDRLFLIS*FWTEL 240
>AW472663
Length = 190
Score = 31.2 bits (69), Expect = 0.98
Identities = 15/41 (36%), Positives = 22/41 (53%)
Frame = -2
Query: 206 HDSIWVVVDRLTKSAHFIPINISYPVAQLAEIYIQNIVKLH 246
HD VV R++K A+F + SY + L I ++KLH
Sbjct: 168 HDMAIVVTFRVSKQAYFYSFSFSYSILGLNMILYLELIKLH 46
>BU090971
Length = 447
Score = 28.1 bits (61), Expect = 8.3
Identities = 11/21 (52%), Positives = 15/21 (71%)
Frame = -3
Query: 1 NFGLNYHPDKAKVVADALSRK 21
NF + Y PDK+ +V DALS +
Sbjct: 361 NFDILYKPDKSNIVVDALSHQ 299
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.341 0.148 0.472
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,789,759
Number of Sequences: 63676
Number of extensions: 246768
Number of successful extensions: 1883
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 1865
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1878
length of query: 443
length of database: 12,639,632
effective HSP length: 100
effective length of query: 343
effective length of database: 6,272,032
effective search space: 2151306976
effective search space used: 2151306976
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC144765.5


