

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144765.1 + phase: 0 /pseudo
(52 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
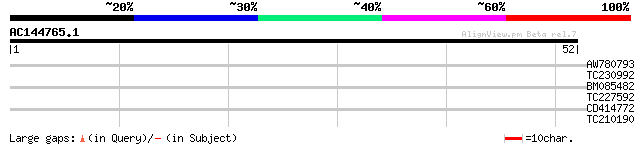
Score E
Sequences producing significant alignments: (bits) Value
AW780793 29 0.59
TC230992 similar to UP|Q944A6 (Q944A6) At1g09020/F7G19_11, parti... 28 0.77
BM085482 28 0.77
TC227592 homologue to GB|BAC42746.1|26451290|AK118120 protein tr... 27 2.9
CD414772 homologue to SP|P14226|PSBO_ Oxygen-evolving enhancer p... 26 3.8
TC210190 similar to UP|Q93Z68 (Q93Z68) AT3g59470/T16L24_20, part... 25 8.5
>AW780793
Length = 419
Score = 28.9 bits (63), Expect = 0.59
Identities = 11/20 (55%), Positives = 16/20 (80%)
Frame = +3
Query: 2 EEANKFWLAYSFRVGFGVRV 21
EEA +F++AY R+GF VR+
Sbjct: 162 EEAREFYIAYGRRIGFTVRI 221
>TC230992 similar to UP|Q944A6 (Q944A6) At1g09020/F7G19_11, partial (24%)
Length = 826
Score = 28.5 bits (62), Expect = 0.77
Identities = 13/38 (34%), Positives = 21/38 (55%)
Frame = -1
Query: 9 LAYSFRVGFGVRVRFANKREDGSVSSCRMLYEGSVNTW 46
LA ++GF +R N++E +S C YE + +TW
Sbjct: 121 LAKKKKLGF*CLIRRENQKEKVPISHCNRTYEETNSTW 8
>BM085482
Length = 415
Score = 28.5 bits (62), Expect = 0.77
Identities = 11/37 (29%), Positives = 20/37 (53%)
Frame = -3
Query: 6 KFWLAYSFRVGFGVRVRFANKREDGSVSSCRMLYEGS 42
K+W +S + FG+ ++F GS+ + YEG+
Sbjct: 353 KYWFLFSIKGTFGIFIKFGESFSLGSLLNQSQTYEGN 243
>TC227592 homologue to GB|BAC42746.1|26451290|AK118120 protein transport
protein subunit - like {Arabidopsis thaliana;} , partial
(80%)
Length = 574
Score = 26.6 bits (57), Expect = 2.9
Identities = 9/25 (36%), Positives = 15/25 (60%)
Frame = -2
Query: 19 VRVRFANKREDGSVSSCRMLYEGSV 43
+R+R N +ED + CR ++ G V
Sbjct: 429 MRIRTTNPKEDRKIKRCRQIWSGRV 355
>CD414772 homologue to SP|P14226|PSBO_ Oxygen-evolving enhancer protein 1
chloroplast precursor (OEE1), partial (28%)
Length = 580
Score = 26.2 bits (56), Expect = 3.8
Identities = 9/20 (45%), Positives = 15/20 (75%)
Frame = -3
Query: 31 SVSSCRMLYEGSVNTWYVKY 50
++SSCR+ +EGS N W ++
Sbjct: 299 AISSCRIRHEGSWNFWTCRF 240
>TC210190 similar to UP|Q93Z68 (Q93Z68) AT3g59470/T16L24_20, partial (65%)
Length = 866
Score = 25.0 bits (53), Expect = 8.5
Identities = 13/29 (44%), Positives = 19/29 (64%), Gaps = 1/29 (3%)
Frame = +1
Query: 4 ANKFWLAYSFRVGFGVRV-RFANKREDGS 31
A+ F+ AY+ VGF VRV + + R DG+
Sbjct: 25 AHAFYNAYATEVGFIVRVSKLSRSRRDGT 111
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.322 0.134 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,547,236
Number of Sequences: 63676
Number of extensions: 22390
Number of successful extensions: 101
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 101
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 101
length of query: 52
length of database: 12,639,632
effective HSP length: 28
effective length of query: 24
effective length of database: 10,856,704
effective search space: 260560896
effective search space used: 260560896
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 52 (24.6 bits)
Medicago: description of AC144765.1


