

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144731.10 + phase: 0 /pseudo
(222 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
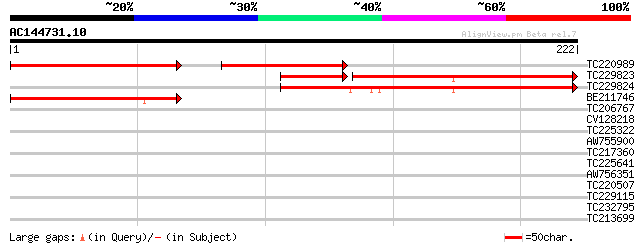
Score E
Sequences producing significant alignments: (bits) Value
TC220989 119 1e-44
TC229823 142 4e-43
TC229824 similar to UP|Q7PL26 (Q7PL26) CG40196-PA.3 (LD17963p), ... 150 4e-37
BE211746 106 9e-24
TC206767 similar to UP|Q9FVN7 (Q9FVN7) NADPH-dependent mannose 6... 28 3.3
CV128218 28 4.3
TC225322 similar to UP|TI41_ARATH (O82316) Probable aquaporin TI... 27 5.6
AW755900 similar to GP|7839365|gb|A S-adenosyl-L-methionine Mg-p... 27 7.3
TC217360 similar to GB|AAL31246.1|16974485|AY061919 AT4g32480/F8... 27 7.3
TC225641 similar to GB|AAA18379.1|475719|ATU08467 RNA-binding pr... 27 9.5
AW756351 27 9.5
TC220507 similar to UP|Q7Q3J9 (Q7Q3J9) AgCP11343 (Fragment), par... 27 9.5
TC229115 similar to UP|RK3B_ARATH (Q9LRN8) 50S ribosomal protein... 27 9.5
TC232795 similar to UP|Q9XI26 (Q9XI26) F9L1.38, partial (49%) 27 9.5
TC213699 similar to UP|Q9XI26 (Q9XI26) F9L1.38, partial (11%) 27 9.5
>TC220989
Length = 568
Score = 119 bits (298), Expect(2) = 1e-44
Identities = 56/67 (83%), Positives = 61/67 (90%)
Frame = +2
Query: 1 MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNEILDYLGKSS 60
MKFLEC+ LDRL+DFL NLNLGERTIKGC+EAYSCKH+G+DKKLSISL EILDYLGKSS
Sbjct: 158 MKFLECSALDRLNDFLGNLNLGERTIKGCLEAYSCKHAGSDKKLSISLETEILDYLGKSS 337
Query: 61 DNDSPSP 67
D DS SP
Sbjct: 338 DTDSSSP 358
Score = 77.8 bits (190), Expect(2) = 1e-44
Identities = 37/49 (75%), Positives = 39/49 (79%)
Frame = +3
Query: 84 PAGRHWFT*FLPSITCIQIMISAQLKLTSFSQRNHGTVLSRFLKPTCLK 132
PA RHWFT F P ITCI IM S QL+LTS SQRNHGT LSRFL PTC+K
Sbjct: 381 PAERHWFTWF*PCITCIPIMTSVQLRLTSCSQRNHGTALSRFLIPTCMK 527
>TC229823
Length = 870
Score = 142 bits (358), Expect(2) = 4e-43
Identities = 70/89 (78%), Positives = 77/89 (85%), Gaps = 1/89 (1%)
Frame = +2
Query: 135 SLLDALFKAVDEVVSLDDCEIYGYLPDSEADPLPERGA-IWSFNFLFYNRKLKRIVTFRL 193
SLLD LFKA+DEVV L DC+IYGY+PD EADPL E G IWSFNFLFYNRKLKRIV+FR
Sbjct: 284 SLLDTLFKALDEVVKLVDCDIYGYVPDFEADPLLESGGTIWSFNFLFYNRKLKRIVSFRF 463
Query: 194 SCFSNLIADGFSLDEICDEYDEEIFADMD 222
SCFSNLIADG +D+I +EYDEEIFADMD
Sbjct: 464 SCFSNLIADGLLIDDIHNEYDEEIFADMD 550
Score = 49.7 bits (117), Expect(2) = 4e-43
Identities = 22/26 (84%), Positives = 24/26 (91%)
Frame = +3
Query: 107 QLKLTSFSQRNHGTVLSRFLKPTCLK 132
QL+LTSFSQRNHGT LSRFL PTC+K
Sbjct: 171 QLRLTSFSQRNHGTALSRFLIPTCMK 248
>TC229824 similar to UP|Q7PL26 (Q7PL26) CG40196-PA.3 (LD17963p), partial (8%)
Length = 779
Score = 150 bits (379), Expect = 4e-37
Identities = 87/151 (57%), Positives = 99/151 (64%), Gaps = 35/151 (23%)
Frame = +1
Query: 107 QLKLTSFSQRNHGTVLSRFLKPTCLK-PQSLLDAL------------------FKA---- 143
QL+LTSFSQRNHGT LSRFL PTC+K P++ L L FK
Sbjct: 100 QLRLTSFSQRNHGTALSRFLIPTCMKLPRNGLKLLGVLLCWTPCSRPLMRYGFFKM*IIF 279
Query: 144 -----------VDEVVSLDDCEIYGYLPDSEADPLPERGA-IWSFNFLFYNRKLKRIVTF 191
V +VV L DC+IYGY+PD EADPL E G IWSFNFLFYNRKLKRIV+F
Sbjct: 280 EIYQDALL*YLVVQVVKLVDCDIYGYVPDFEADPLLESGGTIWSFNFLFYNRKLKRIVSF 459
Query: 192 RLSCFSNLIADGFSLDEICDEYDEEIFADMD 222
R SCFSNLIA+G +D+I +EYDEEIFADMD
Sbjct: 460 RFSCFSNLIAEGLLIDDIHNEYDEEIFADMD 552
>BE211746
Length = 476
Score = 106 bits (264), Expect = 9e-24
Identities = 54/80 (67%), Positives = 60/80 (74%), Gaps = 13/80 (16%)
Frame = +1
Query: 1 MKFLECAPLDRLSDFLSNLNLGERTIKGCIEAYSCKHSGADKKLSISLSNE--------- 51
MKFLEC+ L RL+DFL NLNLG+RTIKGC+EAYSCKH+G+DKKLSISL E
Sbjct: 232 MKFLECSALHRLNDFLGNLNLGQRTIKGCLEAYSCKHAGSDKKLSISLETEVD*CNVYCV 411
Query: 52 ----ILDYLGKSSDNDSPSP 67
ILDYLGKSSD DS SP
Sbjct: 412 YHFTILDYLGKSSDTDSSSP 471
>TC206767 similar to UP|Q9FVN7 (Q9FVN7) NADPH-dependent mannose 6-phosphate
reductase, complete
Length = 1323
Score = 28.1 bits (61), Expect = 3.3
Identities = 15/42 (35%), Positives = 22/42 (51%), Gaps = 9/42 (21%)
Frame = -3
Query: 167 LPERGAIWSFNFLFY---------NRKLKRIVTFRLSCFSNL 199
+P+R IW+ FL++ NRKL + F LS F +L
Sbjct: 946 IPQRRLIWAAVFLYFSARPLEITQNRKLSKYHLFNLSSFDDL 821
>CV128218
Length = 826
Score = 27.7 bits (60), Expect = 4.3
Identities = 11/31 (35%), Positives = 21/31 (67%)
Frame = -3
Query: 30 IEAYSCKHSGADKKLSISLSNEILDYLGKSS 60
++ +SC H+ + SLS+E++DYL +S+
Sbjct: 368 VDLHSCYHAKQKTLIRNSLSSEVMDYLCESA 276
>TC225322 similar to UP|TI41_ARATH (O82316) Probable aquaporin TIP4.1
(Tonoplast intrinsic protein 4.1) (Epsilon-tonoplast
intrinsic protein) (Epsilon-TIP), partial (98%)
Length = 1378
Score = 27.3 bits (59), Expect = 5.6
Identities = 11/30 (36%), Positives = 20/30 (66%)
Frame = -1
Query: 52 ILDYLGKSSDNDSPSPVDSLSSPVDSLSAR 81
++D K++ N+ P+PV+ + SPV S +R
Sbjct: 1231 LIDETSKATTNERPNPVNPMISPVASNQSR 1142
>AW755900 similar to GP|7839365|gb|A S-adenosyl-L-methionine
Mg-protoporphyrin IX methyltranserase {Nicotiana
tabacum}, partial (28%)
Length = 439
Score = 26.9 bits (58), Expect = 7.3
Identities = 12/28 (42%), Positives = 19/28 (67%)
Frame = +2
Query: 60 SDNDSPSPVDSLSSPVDSLSARTSPAGR 87
+ + P P+DSL++ +SLS R SP+ R
Sbjct: 77 TQTEPPFPLDSLTNRPNSLSRRRSPSRR 160
>TC217360 similar to GB|AAL31246.1|16974485|AY061919 AT4g32480/F8B4_180
{Arabidopsis thaliana;} , partial (31%)
Length = 1173
Score = 26.9 bits (58), Expect = 7.3
Identities = 16/40 (40%), Positives = 25/40 (62%), Gaps = 1/40 (2%)
Frame = -2
Query: 43 KLSISLSNEIL-DYLGKSSDNDSPSPVDSLSSPVDSLSAR 81
+LS L+N+I+ ++S++ SP+P S S P DSL R
Sbjct: 209 ELSSLLTNDIIISPSSRNSNSTSPTPSCSFSHPNDSLRGR 90
>TC225641 similar to GB|AAA18379.1|475719|ATU08467 RNA-binding protein 2
{Arabidopsis thaliana;} , partial (59%)
Length = 1223
Score = 26.6 bits (57), Expect = 9.5
Identities = 20/60 (33%), Positives = 28/60 (46%), Gaps = 2/60 (3%)
Frame = -2
Query: 34 SCKHSGADKKLSISLSNEILDYLGKSSDNDS--PSPVDSLSSPVDSLSARTSPAGRHWFT 91
+C S A+ LS S L S+ + + PSP S+SSP S+ SP WF+
Sbjct: 496 ACSKSEANFSLSKSKGRFPTKILTSSASSSAKPPSPPSSISSPPSSVPHAVSP----WFS 329
>AW756351
Length = 426
Score = 26.6 bits (57), Expect = 9.5
Identities = 13/33 (39%), Positives = 20/33 (60%), Gaps = 1/33 (3%)
Frame = -3
Query: 87 RHWFT*FLPSITCIQIMISAQL-KLTSFSQRNH 118
RHW +PSI+ Q++I + + SF+ RNH
Sbjct: 244 RHWVHNPVPSISSAQLIIDSSITNYLSFNLRNH 146
>TC220507 similar to UP|Q7Q3J9 (Q7Q3J9) AgCP11343 (Fragment), partial (3%)
Length = 829
Score = 26.6 bits (57), Expect = 9.5
Identities = 13/29 (44%), Positives = 19/29 (64%)
Frame = +3
Query: 49 SNEILDYLGKSSDNDSPSPVDSLSSPVDS 77
+ E +D G SSD+ S S +S SSP+D+
Sbjct: 165 AEEEIDGFGSSSDSSSVSWFESDSSPIDA 251
>TC229115 similar to UP|RK3B_ARATH (Q9LRN8) 50S ribosomal protein L3-2,
chloroplast precursor, partial (81%)
Length = 1211
Score = 26.6 bits (57), Expect = 9.5
Identities = 17/64 (26%), Positives = 29/64 (44%), Gaps = 7/64 (10%)
Frame = +1
Query: 25 TIKGCIEAYSC-------KHSGADKKLSISLSNEILDYLGKSSDNDSPSPVDSLSSPVDS 77
+IK C C +H ++LS ++ L + + + SPSP+ SLSS +
Sbjct: 16 SIKSCERTRFCDLPLHHRRHLHLSGSAMLALSRGLVSRLQRFAVDSSPSPLRSLSSDAAA 195
Query: 78 LSAR 81
+ R
Sbjct: 196IQTR 207
>TC232795 similar to UP|Q9XI26 (Q9XI26) F9L1.38, partial (49%)
Length = 1027
Score = 26.6 bits (57), Expect = 9.5
Identities = 16/53 (30%), Positives = 27/53 (50%)
Frame = -2
Query: 28 GCIEAYSCKHSGADKKLSISLSNEILDYLGKSSDNDSPSPVDSLSSPVDSLSA 80
GC + S +D +SLS+ I + SS + S +P+ S+S ++S A
Sbjct: 738 GCCSSGSVSMVSSDSSSLLSLSS*ITGTISPSSSSSSIAPM*SVSKQLESWMA 580
>TC213699 similar to UP|Q9XI26 (Q9XI26) F9L1.38, partial (11%)
Length = 822
Score = 26.6 bits (57), Expect = 9.5
Identities = 16/53 (30%), Positives = 27/53 (50%)
Frame = -3
Query: 28 GCIEAYSCKHSGADKKLSISLSNEILDYLGKSSDNDSPSPVDSLSSPVDSLSA 80
GC + S +D +SLS+ I + SS + S +P+ S+S ++S A
Sbjct: 760 GCCSSGSVSMVSSDSSSLLSLSS*ITGTISPSSSSSSIAPM*SVSKQLESWMA 602
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.323 0.139 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,373,757
Number of Sequences: 63676
Number of extensions: 149135
Number of successful extensions: 761
Number of sequences better than 10.0: 30
Number of HSP's better than 10.0 without gapping: 753
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 758
length of query: 222
length of database: 12,639,632
effective HSP length: 94
effective length of query: 128
effective length of database: 6,654,088
effective search space: 851723264
effective search space used: 851723264
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 57 (26.6 bits)
Medicago: description of AC144731.10


