

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144726.5 - phase: 0 /pseudo
(293 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
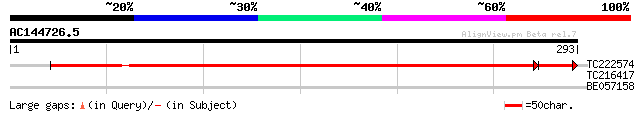
TC222574 similar to UP|Q8H1H1 (Q8H1H1) Translin-associated facto... 388 e-110
TC216417 similar to UP|Q7XJ64 (Q7XJ64) At1g15860, partial (87%) 28 3.7
BE057158 28 6.4
>TC222574 similar to UP|Q8H1H1 (Q8H1H1) Translin-associated factor X
(Fragment), complete
Length = 1036
Score = 388 bits (996), Expect(2) = e-110
Identities = 199/252 (78%), Positives = 220/252 (86%)
Frame = +2
Query: 22 QRLHQSILLFFDFGFSTVTGTNFQNTSKRPKTMSIATDTATVTDSAMKEPFTKYTEYLNN 81
Q LH F + GTN Q++ KR +TM+ ++ T + A+KE F++YT+ LN+
Sbjct: 2 QMLHSLRFSLFMASKHRIAGTNIQSSPKRARTMATSS---TAIEPALKEAFSRYTQCLND 172
Query: 82 LNDKRERVVKASRDITMNSKKVIFQVHRMSKYNKDEVLEKAEKDLAAVTNQHVSRLVKEL 141
LNDKRERVVKASRD+TMNSKKVIFQVHRMSKYNK E+LEKAEKDLAAVT+Q++SRLVKEL
Sbjct: 173 LNDKRERVVKASRDVTMNSKKVIFQVHRMSKYNKVEILEKAEKDLAAVTDQYMSRLVKEL 352
Query: 142 QGTDFWKLRRAYSPGIQEYVEAATFCSFCKNGTLLKLDEINKTLLPLSDPSLQPLQINIL 201
QGTDFWKLRRAYSPGIQEYVEAATF FCK+GTLLKLDEINKTLLPLSDPSL PLQINIL
Sbjct: 353 QGTDFWKLRRAYSPGIQEYVEAATFYGFCKSGTLLKLDEINKTLLPLSDPSLDPLQINIL 532
Query: 202 DYILGLADLTGELMRLAIGRISDGELEFAEKICSFARDIYRELTLVVPHMDDSSDMKTKM 261
DYILG+ADLTGELMRLAIGRISDGELEFAEKIC FARDIYRELTLVVPHMDDSSDMKTKM
Sbjct: 533 DYILGVADLTGELMRLAIGRISDGELEFAEKICRFARDIYRELTLVVPHMDDSSDMKTKM 712
Query: 262 ETMLQSVMKIEN 273
+ MLQSVMKIEN
Sbjct: 713 DVMLQSVMKIEN 748
Score = 29.3 bits (64), Expect(2) = e-110
Identities = 14/20 (70%), Positives = 15/20 (75%)
Frame = +3
Query: 274 VFMSEDQSIYHFWDQMIQVL 293
VFM E QSI+HF D IQVL
Sbjct: 759 VFM*EGQSIFHFLDPTIQVL 818
>TC216417 similar to UP|Q7XJ64 (Q7XJ64) At1g15860, partial (87%)
Length = 826
Score = 28.5 bits (62), Expect = 3.7
Identities = 26/98 (26%), Positives = 46/98 (46%)
Frame = +1
Query: 103 VIFQVHRMSKYNKDEVLEKAEKDLAAVTNQHVSRLVKELQGTDFWKLRRAYSPGIQEYVE 162
++F VH+ +Y+K ++ ++ + L A+ VS+L L D K R S Y
Sbjct: 205 IMFIVHKQQRYSKQQI*DEWRRGLKALRADTVSKLKNLL--PDLEKEVRRPSNFADFYSY 378
Query: 163 AATFCSFCKNGTLLKLDEINKTLLPLSDPSLQPLQINI 200
A +C + + ++ I + LL L S P Q+N+
Sbjct: 379 AFQYCLTEEKQKSIDIESICE-LLTLVLGSTFPAQVNL 489
>BE057158
Length = 409
Score = 27.7 bits (60), Expect = 6.4
Identities = 13/37 (35%), Positives = 21/37 (56%)
Frame = -1
Query: 4 SFSSSFHRLSLFMASEKTQRLHQSILLFFDFGFSTVT 40
S S + R+ + ++ LH +I LF+ F FST+T
Sbjct: 406 SHSHVYMRMKVIFYNKYIFLLHSTIFLFYLFNFSTIT 296
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.320 0.133 0.371
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,461,443
Number of Sequences: 63676
Number of extensions: 113411
Number of successful extensions: 581
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 580
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 581
length of query: 293
length of database: 12,639,632
effective HSP length: 97
effective length of query: 196
effective length of database: 6,463,060
effective search space: 1266759760
effective search space used: 1266759760
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC144726.5


