

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
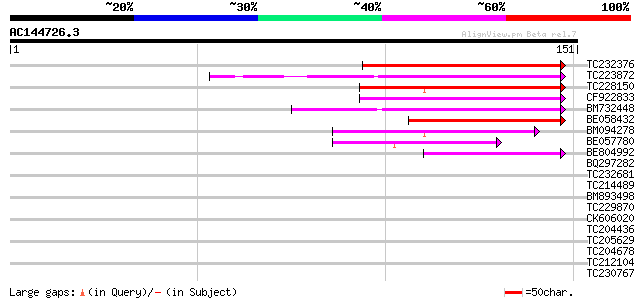
Query= AC144726.3 - phase: 0
(151 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC232376 86 5e-18
TC223872 similar to UP|Q9MF81 (Q9MF81) Orf102a protein, partial ... 71 2e-13
TC228150 homologue to UP|Q9S849 (Q9S849) Squamosa promoter bindi... 53 5e-08
CF922833 51 2e-07
BM732448 similar to SP|O04003|LG1_M LIGULELESS1 protein. [Maize]... 49 7e-07
BE058432 similar to SP|Q38741|SBP1_ Squamosa-promoter binding pr... 48 2e-06
BM094278 47 3e-06
BE057780 similar to GP|5931778|emb| SBP-domain protein 1 {Zea ma... 41 2e-04
BE804992 similar to PIR|T52297|T522 squamosa promoter binding pr... 41 2e-04
BQ297282 similar to PIR|T52607|T526 squamosa promoter binding pr... 35 0.018
TC232681 weakly similar to GB|AAP21377.1|30102918|BT006569 At1g4... 31 0.26
TC214489 similar to UP|O65450 (O65450) Glycine-rich protein, par... 30 0.44
BM893498 similar to GP|18650611|gb| At1g27370/F17L21_16 {Arabido... 30 0.57
TC229870 similar to UP|Q9FVL1 (Q9FVL1) Phosphatidic acid phospha... 29 0.75
CK606020 29 0.98
TC204436 28 1.3
TC205629 similar to UP|Q9LEX1 (Q9LEX1) CaLB protein (At3g61050),... 28 1.7
TC204678 similar to UP|Q94EJ6 (Q94EJ6) AT4g18030/T6K21_210, part... 28 1.7
TC212104 UP|Q6QI42 (Q6QI42) LRRGT00166, partial (22%) 28 1.7
TC230767 similar to UP|O49361 (O49361) S-receptor kinase-like pr... 28 2.2
>TC232376
Length = 1033
Score = 86.3 bits (212), Expect = 5e-18
Identities = 38/54 (70%), Positives = 46/54 (84%)
Frame = +2
Query: 95 KARAINNGTQIVSSLADGCQADLSNYRGYHMRHKVCKLHSKTSQVTLGGHKQRF 148
+ARA++NGTQ VS L DGC ADLSN R YH RHKVC++HSKT+QV++GG KQRF
Sbjct: 2 RARALSNGTQTVSCLVDGCHADLSNCRDYHRRHKVCEVHSKTAQVSIGGQKQRF 163
>TC223872 similar to UP|Q9MF81 (Q9MF81) Orf102a protein, partial (44%)
Length = 411
Score = 70.9 bits (172), Expect = 2e-13
Identities = 42/95 (44%), Positives = 57/95 (59%)
Frame = +1
Query: 54 VDLKLGHVGNSSAESGLTKSKDVGGFTKITSSSSSTSGSSKKARAINNGTQIVSSLADGC 113
VDL+LG +A + K TK + + SS SGSSK++R + NG Q + DGC
Sbjct: 142 VDLRLGE--EKTAPDVVAKD------TKDSKTVSSPSGSSKRSR-LQNGLQNMCCSVDGC 294
Query: 114 QADLSNYRGYHMRHKVCKLHSKTSQVTLGGHKQRF 148
+DLS+ R YH RH+VC+ HSKT V +GG + RF
Sbjct: 295 NSDLSDCREYHRRHRVCEKHSKTPVVMVGGKQXRF 399
>TC228150 homologue to UP|Q9S849 (Q9S849) Squamosa promoter binding
protein-like 8, partial (23%)
Length = 685
Score = 53.1 bits (126), Expect = 5e-08
Identities = 26/59 (44%), Positives = 36/59 (60%), Gaps = 4/59 (6%)
Frame = +1
Query: 94 KKARAINNGTQIVSSL----ADGCQADLSNYRGYHMRHKVCKLHSKTSQVTLGGHKQRF 148
+++R + G+ I S+ A+GC ADLS + YH RHKVC+ HSK + V G QRF
Sbjct: 13 RRSRPVEPGSTISSNSPRCQAEGCNADLSQAKHYHRRHKVCEFHSKAATVIAAGLTQRF 189
>CF922833
Length = 664
Score = 50.8 bits (120), Expect = 2e-07
Identities = 23/55 (41%), Positives = 32/55 (57%)
Frame = +2
Query: 94 KKARAINNGTQIVSSLADGCQADLSNYRGYHMRHKVCKLHSKTSQVTLGGHKQRF 148
K++ + G+ S D C ADLS + YH RHKVC+ H+K V + G +QRF
Sbjct: 209 KRSGSKGGGSMPPSCQVDNCDADLSEAKQYHRRHKVCEYHAKAPSVHMAGLQQRF 373
>BM732448 similar to SP|O04003|LG1_M LIGULELESS1 protein. [Maize] {Zea mays},
partial (12%)
Length = 356
Score = 49.3 bits (116), Expect = 7e-07
Identities = 28/73 (38%), Positives = 37/73 (50%)
Frame = +1
Query: 76 VGGFTKITSSSSSTSGSSKKARAINNGTQIVSSLADGCQADLSNYRGYHMRHKVCKLHSK 135
VGG + G R ++G S A+ C ADL+ + YH RHKVC+LHSK
Sbjct: 115 VGGEGGLAEDEKKKGGGGSGRRG-SSGFFPPSCQAEMCGADLTVAKRYHRRHKVCELHSK 291
Query: 136 TSQVTLGGHKQRF 148
V + G +QRF
Sbjct: 292 APSVMVAGLRQRF 330
>BE058432 similar to SP|Q38741|SBP1_ Squamosa-promoter binding protein 1.
[Garden snapdragon] {Antirrhinum majus}, partial (58%)
Length = 433
Score = 47.8 bits (112), Expect = 2e-06
Identities = 21/42 (50%), Positives = 28/42 (66%)
Frame = +1
Query: 107 SSLADGCQADLSNYRGYHMRHKVCKLHSKTSQVTLGGHKQRF 148
S A+ C ADL++ + YH RHKVC+ HSK V + G +QRF
Sbjct: 19 SCQAERCGADLTDAKRYHRRHKVCEFHSKAPVVVVAGLRQRF 144
>BM094278
Length = 428
Score = 47.0 bits (110), Expect = 3e-06
Identities = 23/56 (41%), Positives = 33/56 (58%), Gaps = 1/56 (1%)
Frame = +2
Query: 87 SSTSGSSKKARAINNGTQIVSSL-ADGCQADLSNYRGYHMRHKVCKLHSKTSQVTL 141
++ S S+K+ R+ + GT D C+ DLS + YH RHKVC+ HSK S+ L
Sbjct: 251 TNNSNSNKRVRSGSPGTSSYPMCQVDNCREDLSKAKDYHRRHKVCEAHSKASKALL 418
>BE057780 similar to GP|5931778|emb| SBP-domain protein 1 {Zea mays}, partial
(6%)
Length = 286
Score = 41.2 bits (95), Expect = 2e-04
Identities = 21/52 (40%), Positives = 27/52 (51%), Gaps = 7/52 (13%)
Frame = +3
Query: 87 SSTSGSSKKARAINN-------GTQIVSSLADGCQADLSNYRGYHMRHKVCK 131
S KK R ++N G++ S DGC ADLS + YH RHKVC+
Sbjct: 126 SEYGDDGKKKRVVSNKRGSKAGGSEPPSCQVDGCNADLSEAKPYHRRHKVCE 281
>BE804992 similar to PIR|T52297|T522 squamosa promoter binding
protein-homolog 5 [imported] - garden snapdragon
(fragment), partial (22%)
Length = 418
Score = 40.8 bits (94), Expect = 2e-04
Identities = 17/38 (44%), Positives = 23/38 (59%)
Frame = +2
Query: 111 DGCQADLSNYRGYHMRHKVCKLHSKTSQVTLGGHKQRF 148
+GC L N + YH RH+VC HSK + + G +QRF
Sbjct: 5 EGCHVALVNAKEYHRRHRVCDKHSKAPKAVVLGLEQRF 118
>BQ297282 similar to PIR|T52607|T526 squamosa promoter binding protein 5
[imported] - Arabidopsis thaliana, partial (42%)
Length = 427
Score = 34.7 bits (78), Expect = 0.018
Identities = 15/34 (44%), Positives = 20/34 (58%)
Frame = +2
Query: 115 ADLSNYRGYHMRHKVCKLHSKTSQVTLGGHKQRF 148
ADL + YH RH+VC+ H K V + +QRF
Sbjct: 170 ADLHEAKQYHRRHRVCEYHVKAQVVLVDEVRQRF 271
>TC232681 weakly similar to GB|AAP21377.1|30102918|BT006569 At1g47970
{Arabidopsis thaliana;} , partial (39%)
Length = 867
Score = 30.8 bits (68), Expect = 0.26
Identities = 25/85 (29%), Positives = 42/85 (49%)
Frame = -2
Query: 63 NSSAESGLTKSKDVGGFTKITSSSSSTSGSSKKARAINNGTQIVSSLADGCQADLSNYRG 122
+SS+ S + S +T ++SSSSS+S SS + A ++ + SSL+ D + R
Sbjct: 365 SSSSSSLSSSSGSPPRWTSLSSSSSSSSSSSPPSGASSSSSSSSSSLSSSSSKD--SRRR 192
Query: 123 YHMRHKVCKLHSKTSQVTLGGHKQR 147
H + LH +T + H+ R
Sbjct: 191 LHRPLRRLLLHPRTRILRPPPHQPR 117
Score = 26.2 bits (56), Expect = 6.3
Identities = 15/31 (48%), Positives = 20/31 (64%)
Frame = -2
Query: 63 NSSAESGLTKSKDVGGFTKITSSSSSTSGSS 93
+SS+ES L+ GG + + SSSSTS SS
Sbjct: 590 SSSSESSLSDLFLFGGASDFSPSSSSTSSSS 498
>TC214489 similar to UP|O65450 (O65450) Glycine-rich protein, partial (9%)
Length = 443
Score = 30.0 bits (66), Expect = 0.44
Identities = 28/90 (31%), Positives = 41/90 (45%), Gaps = 6/90 (6%)
Frame = -1
Query: 14 ILYIVLFLNNDEVFWRPWHDSIWNILYGFVCFSMCANEFFVDLKLGH--VGNSSAESGL- 70
ILY + F NN+ + + + +I F FSM FF+ S+ S L
Sbjct: 266 ILYAIAFFNNNSI--SSFATFVSSIFPTFSSFSMMIVSFFITSNSSSFIAATSTVASALA 93
Query: 71 -TKSKDVGGFTKI--TSSSSSTSGSSKKAR 97
T++ T I TSSSSS+S +S + R
Sbjct: 92 TTRTSSFTTTTSISSTSSSSSSSSTSSRTR 3
>BM893498 similar to GP|18650611|gb| At1g27370/F17L21_16 {Arabidopsis
thaliana}, partial (14%)
Length = 421
Score = 29.6 bits (65), Expect = 0.57
Identities = 10/23 (43%), Positives = 18/23 (77%)
Frame = +1
Query: 126 RHKVCKLHSKTSQVTLGGHKQRF 148
+H+VC+ HSK+ +V + G ++RF
Sbjct: 4 KHRVCEAHSKSPKVVIAGLERRF 72
>TC229870 similar to UP|Q9FVL1 (Q9FVL1) Phosphatidic acid phosphatase beta ,
partial (73%)
Length = 935
Score = 29.3 bits (64), Expect = 0.75
Identities = 13/51 (25%), Positives = 24/51 (46%)
Frame = +1
Query: 3 IPMLCLFNVCSILYIVLFLNNDEVFWRPWHDSIWNILYGFVCFSMCANEFF 53
+ LCL + ++ ++ ++ + +W W D L G V S C +FF
Sbjct: 319 VAKLCLVFLPFLVAAMIAVSRVDDYWHHWQDVFAGALIGMVIASFCYLQFF 471
>CK606020
Length = 340
Score = 28.9 bits (63), Expect = 0.98
Identities = 16/50 (32%), Positives = 28/50 (56%)
Frame = -3
Query: 63 NSSAESGLTKSKDVGGFTKITSSSSSTSGSSKKARAINNGTQIVSSLADG 112
++S+ + + S + +SSSSS+S SS + + ++ I SSLA G
Sbjct: 308 STSSSTSSSSSSSSSSISSSSSSSSSSSSSSSSSSSSSSDGSISSSLATG 159
>TC204436
Length = 2771
Score = 28.5 bits (62), Expect = 1.3
Identities = 13/36 (36%), Positives = 22/36 (61%), Gaps = 1/36 (2%)
Frame = -2
Query: 3 IPMLCLFNVCSILYIVLFLNNDEVF-WRPWHDSIWN 37
+P LCL+ + ++ LFL + VF + P + SIW+
Sbjct: 1783 VPFLCLYILYTLFIFFLFLWSTRVFPFAPTYSSIWD 1676
>TC205629 similar to UP|Q9LEX1 (Q9LEX1) CaLB protein (At3g61050), partial (92%)
Length = 1766
Score = 28.1 bits (61), Expect = 1.7
Identities = 16/55 (29%), Positives = 25/55 (45%)
Frame = -3
Query: 70 LTKSKDVGGFTKITSSSSSTSGSSKKARAINNGTQIVSSLADGCQADLSNYRGYH 124
L +S G T +++SST +S K + N T + + + SN R YH
Sbjct: 1368 LAQSTTNGSKTTPNNTNSSTDAASNKPGSSTNTTSHKTGTGNNTSTNQSNTRAYH 1204
>TC204678 similar to UP|Q94EJ6 (Q94EJ6) AT4g18030/T6K21_210, partial (91%)
Length = 2339
Score = 28.1 bits (61), Expect = 1.7
Identities = 19/43 (44%), Positives = 23/43 (53%), Gaps = 4/43 (9%)
Frame = +3
Query: 27 FWRPWHDS----IWNILYGFVCFSMCANEFFVDLKLGHVGNSS 65
+W PWH I ++ YG V FSM +DLK GHV N S
Sbjct: 864 YWCPWHHPPSIPIKSLRYGPV-FSMSNT---MDLK*GHVSNGS 980
>TC212104 UP|Q6QI42 (Q6QI42) LRRGT00166, partial (22%)
Length = 655
Score = 28.1 bits (61), Expect = 1.7
Identities = 14/27 (51%), Positives = 20/27 (73%)
Frame = +2
Query: 64 SSAESGLTKSKDVGGFTKITSSSSSTS 90
SSAE+G KS++ +K +SSSSS+S
Sbjct: 200 SSAETGEAKSRETNANSKSSSSSSSSS 280
>TC230767 similar to UP|O49361 (O49361) S-receptor kinase-like protein,
partial (15%)
Length = 511
Score = 27.7 bits (60), Expect = 2.2
Identities = 10/27 (37%), Positives = 15/27 (55%)
Frame = +3
Query: 5 MLCLFNVCSILYIVLFLNNDEVFWRPW 31
+ CLF+VC I+ + N E+ W W
Sbjct: 408 LFCLFSVCCIVE*ISLCNKAELLWIVW 488
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.324 0.137 0.431
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,898,101
Number of Sequences: 63676
Number of extensions: 142127
Number of successful extensions: 1351
Number of sequences better than 10.0: 64
Number of HSP's better than 10.0 without gapping: 1331
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1345
length of query: 151
length of database: 12,639,632
effective HSP length: 89
effective length of query: 62
effective length of database: 6,972,468
effective search space: 432293016
effective search space used: 432293016
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 54 (25.4 bits)
Medicago: description of AC144726.3


