

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144726.11 + phase: 0
(515 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
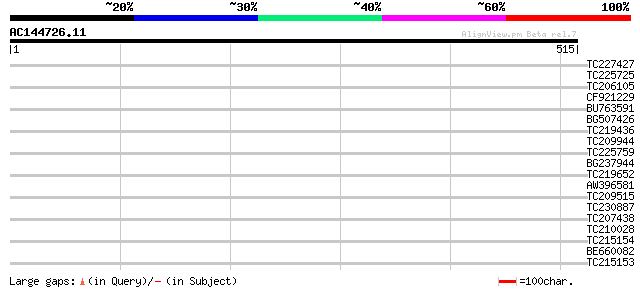
Score E
Sequences producing significant alignments: (bits) Value
TC227427 similar to UP|Q94JU5 (Q94JU5) At1g02330/T6A9_12, partia... 33 0.24
TC225725 similar to UP|Q9FR29 (Q9FR29) Transcription factor WRKY... 33 0.31
TC206105 weakly similar to UP|Q7RH28 (Q7RH28) Mature-parasite-in... 32 0.52
CF921229 32 0.52
BU763591 32 0.68
BG507426 similar to GP|15384989|emb Xaa-Pro aminopeptidase 1 {Ly... 30 2.0
TC219436 similar to GB|AAP37868.1|30725692|BT008509 At5g08200 {A... 30 2.0
TC209944 weakly similar to UP|Q9NP08 (Q9NP08) H6 homeodomain pro... 30 2.6
TC225759 similar to UP|Q9SMK5 (Q9SMK5) Plasma membrane intrinsic... 30 3.4
BG237944 29 4.4
TC219652 similar to UP|Q39878 (Q39878) Mitotic cyclin a2-type, p... 29 4.4
AW396581 29 5.8
TC209515 similar to UP|Q8VWH9 (Q8VWH9) Stress responsive gene 6,... 29 5.8
TC230887 similar to UP|Q9ZUM1 (Q9ZUM1) Expressed protein (At2g02... 28 7.6
TC207438 homologue to UP|Q08145 (Q08145) GTP-binding protein, pa... 28 7.6
TC210028 similar to UP|Q9LIR7 (Q9LIR7) Arabidopsis thaliana geno... 28 7.6
TC215154 similar to UP|Q6SZ89 (Q6SZ89) Translational elongation ... 28 9.9
BE660082 28 9.9
TC215153 similar to UP|Q6SZ89 (Q6SZ89) Translational elongation ... 28 9.9
>TC227427 similar to UP|Q94JU5 (Q94JU5) At1g02330/T6A9_12, partial (85%)
Length = 1381
Score = 33.5 bits (75), Expect = 0.24
Identities = 29/106 (27%), Positives = 44/106 (41%), Gaps = 9/106 (8%)
Frame = +3
Query: 243 RVIQAACQIGEAIEH-EGRIYRFMEKTKVKKATSHKSDSD--------SVPAPVKGENLT 293
R I AA Q ++ E +Y+ E KVK+ S +S + +P K +N+
Sbjct: 459 RKIDAADQAENELKRAEDELYKIPEHLKVKRRNSEESSTQWTTGIAEVQLPIEYKLKNIE 638
Query: 294 AEENEKLAKEEKRLRNKVASLIKKQKKQQALGIVKGRDQAKPWGQE 339
E K +EKRL + S A +GRD A+ +E
Sbjct: 639 ETEAAKKLLQEKRLMGRTKSDFSIPSSYSADYFQRGRDYAEKLRRE 776
>TC225725 similar to UP|Q9FR29 (Q9FR29) Transcription factor WRKY4, partial
(51%)
Length = 1012
Score = 33.1 bits (74), Expect = 0.31
Identities = 21/87 (24%), Positives = 46/87 (52%), Gaps = 6/87 (6%)
Frame = +2
Query: 259 GRIYRFMEKTKVKKATSHK-----SDSDSVPAPVKGENLTAEENEKLAKEEKR-LRNKVA 312
G + +M K K+ +S K S+++S+P V G + ++ +E+ K++K ++ K++
Sbjct: 203 GNLMEYMRKNPDKEHSSSKKRKSESNNNSIPMGVNGTSESSSTDEESCKKQKEDIKTKIS 382
Query: 313 SLIKKQKKQQALGIVKGRDQAKPWGQE 339
+ + + IVK Q + +GQ+
Sbjct: 383 RVYMRTEASDKSLIVKDGYQWRKYGQK 463
>TC206105 weakly similar to UP|Q7RH28 (Q7RH28) Mature-parasite-infected
erythrocyte surface antigen, partial (5%)
Length = 858
Score = 32.3 bits (72), Expect = 0.52
Identities = 21/68 (30%), Positives = 35/68 (50%)
Frame = +1
Query: 103 EVVQKLLDQMVIEEEKKNPQLEEKKKNKNNSYKYKMLRRRQIKIETEAWEEAAREYQELL 162
EV ++ + VI EEKK ++ E+KK + K + + K ETE EE E+
Sbjct: 655 EVTEEKKEAEVIVEEKKEVEVTEEKKEVEVTEGKKEVEVIEEKKETEVTEEKKEVEVEVR 834
Query: 163 EDMREQKL 170
E+ +E ++
Sbjct: 835 EEKKESEV 858
>CF921229
Length = 680
Score = 32.3 bits (72), Expect = 0.52
Identities = 21/68 (30%), Positives = 35/68 (50%)
Frame = -3
Query: 103 EVVQKLLDQMVIEEEKKNPQLEEKKKNKNNSYKYKMLRRRQIKIETEAWEEAAREYQELL 162
EV ++ + VI EEKK ++ E+KK + K + + K ETE EE E+
Sbjct: 531 EVTEEKKEAEVIVEEKKEVEVTEEKKEVEVTEGKKEVEVIEEKKETEVTEEKKEVEVEVR 352
Query: 163 EDMREQKL 170
E+ +E ++
Sbjct: 351 EEKKESEV 328
>BU763591
Length = 389
Score = 32.0 bits (71), Expect = 0.68
Identities = 21/67 (31%), Positives = 34/67 (50%)
Frame = -1
Query: 103 EVVQKLLDQMVIEEEKKNPQLEEKKKNKNNSYKYKMLRRRQIKIETEAWEEAAREYQELL 162
EV ++ + VI EEKK ++ E+KK + K + + K ETE EE E+
Sbjct: 203 EVTEEKKEAEVIVEEKKEVEVTEEKKEVEVTEGKKEVEVIEEKKETEVTEEKKEVEVEVR 24
Query: 163 EDMREQK 169
E+ +E +
Sbjct: 23 EEKKESE 3
>BG507426 similar to GP|15384989|emb Xaa-Pro aminopeptidase 1 {Lycopersicon
esculentum}, partial (11%)
Length = 436
Score = 30.4 bits (67), Expect = 2.0
Identities = 18/42 (42%), Positives = 26/42 (61%)
Frame = -1
Query: 67 PFSFSSISLNHFRTYGSVAAEAIESTDTEDDCSGSEEVVQKL 108
PF+ ++SL FRT SV+A+A + DTE S + VV +L
Sbjct: 190 PFACLAVSLGIFRT--SVSAKATLAVDTESSSSVASNVVSEL 71
>TC219436 similar to GB|AAP37868.1|30725692|BT008509 At5g08200 {Arabidopsis
thaliana;} , partial (14%)
Length = 761
Score = 30.4 bits (67), Expect = 2.0
Identities = 22/67 (32%), Positives = 30/67 (43%), Gaps = 4/67 (5%)
Frame = +3
Query: 89 IESTDTEDDCSGSEEVVQKLLDQMVIEEEKKNPQL----EEKKKNKNNSYKYKMLRRRQI 144
+ ST + D E + +L +E E+K + EKKK K K KM RRR
Sbjct: 555 LRSTHRQQDLGSETEKHEIVLKGKWVEREQKGQRKMCSWREKKKRKGKKGKLKMDRRR-- 728
Query: 145 KIETEAW 151
K+ E W
Sbjct: 729 KLYKERW 749
>TC209944 weakly similar to UP|Q9NP08 (Q9NP08) H6 homeodomain protein,
partial (5%)
Length = 589
Score = 30.0 bits (66), Expect = 2.6
Identities = 14/57 (24%), Positives = 33/57 (57%)
Frame = -2
Query: 118 KKNPQLEEKKKNKNNSYKYKMLRRRQIKIETEAWEEAAREYQELLEDMREQKLAPNL 174
+K P+L+ ++ +++ + K+ +R++ E W + ARE++ +E R Q+ P +
Sbjct: 276 EKTPELK*EETAESSRRRMKVKEQRRV----ETWTKKAREWRFSVEQQRTQETTPEI 118
>TC225759 similar to UP|Q9SMK5 (Q9SMK5) Plasma membrane intrinsic
polypeptide, partial (74%)
Length = 1051
Score = 29.6 bits (65), Expect = 3.4
Identities = 26/102 (25%), Positives = 44/102 (42%)
Frame = +1
Query: 68 FSFSSISLNHFRTYGSVAAEAIESTDTEDDCSGSEEVVQKLLDQMVIEEEKKNPQLEEKK 127
F F +S V A T TE++ S VV++ ++V+EEEKK + EE+K
Sbjct: 475 FVFEKVSTFIVTEEKDVEAPPGVETKTEEETSS---VVKER--EIVVEEEKKKEKKEEEK 639
Query: 128 KNKNNSYKYKMLRRRQIKIETEAWEEAAREYQELLEDMREQK 169
+ K + + + + E+ A E ++E K
Sbjct: 640 PQVIETSDEKKVEEKPAETSAKGEEKPAEEAAAVVEQAEPPK 765
>BG237944
Length = 486
Score = 29.3 bits (64), Expect = 4.4
Identities = 18/54 (33%), Positives = 30/54 (55%), Gaps = 4/54 (7%)
Frame = -1
Query: 111 QMVIEEEKKNPQLE----EKKKNKNNSYKYKMLRRRQIKIETEAWEEAAREYQE 160
Q I+E+KK +E +KKK + + + + R R+ KIET +E +E +E
Sbjct: 282 QKKIKEKKKRSMIEMRRRKKKKIEES*ERRNLSRERESKIETRFGDELKKEARE 121
>TC219652 similar to UP|Q39878 (Q39878) Mitotic cyclin a2-type, partial (30%)
Length = 670
Score = 29.3 bits (64), Expect = 4.4
Identities = 23/75 (30%), Positives = 34/75 (44%), Gaps = 5/75 (6%)
Frame = +1
Query: 12 RRFILPSQSS-----TSLRFSHQHNFPEKILRHPFSRFSQVGIFTSEYKLGRSNFSFSEH 66
RRFIL SQSS L F + ++ + F +F I S L R + SEH
Sbjct: 85 RRFILASQSSYKVSYVELEFLANYLAELTLVEYSFLQFLPSLIAASAVLLARWTLNQSEH 264
Query: 67 PFSFSSISLNHFRTY 81
P+ + ++ H+ Y
Sbjct: 265 PW---NSTMEHYTNY 300
>AW396581
Length = 401
Score = 28.9 bits (63), Expect = 5.8
Identities = 22/68 (32%), Positives = 37/68 (54%), Gaps = 2/68 (2%)
Frame = -1
Query: 266 EKTKVKKATSHKSDSDSVPAPVKGENLTAEENEKLAKEEKRLRNKVASLIKKQKKQQALG 325
+K K KK K+D++ V KG+ E+NE+ KEEK+ + KK++K +
Sbjct: 251 KKKKEKKVKDDKTDAEKV----KGKEDDGEDNEE--KEEKKKK-------KKKEKDDKID 111
Query: 326 I--VKGRD 331
+ VKG++
Sbjct: 110 VEKVKGKE 87
>TC209515 similar to UP|Q8VWH9 (Q8VWH9) Stress responsive gene 6, Srg6
(Stress responsive gene 6 protein, Srg6), partial (55%)
Length = 1386
Score = 28.9 bits (63), Expect = 5.8
Identities = 27/145 (18%), Positives = 63/145 (42%), Gaps = 4/145 (2%)
Frame = +2
Query: 253 EAIEHEGRIYRFMEKTKVKKATSHKSDSDSVPAPVKGENLTAEENEKLAKEEKRLRNKVA 312
+A++ + + E+ ++ D D P P + E+ ++++ ++KRL
Sbjct: 86 DALKEDEEDDNYQEEPEIADEFDSDFDQDE-PVPEEEEDPNKNDDDERMHKKKRLIFPGK 262
Query: 313 SLIKKQKKQQALGIV----KGRDQAKPWGQEAQVKVGSRLIQLLIETAYLEPPANKFGDG 368
+L KK+KK++ + + K + + G+ + + G R+I+ T+ + A +
Sbjct: 263 TLAKKKKKKKTISKLESSPKENEDEEHSGKVVEDEGGERMIRKSTRTSVIVRQAER---- 430
Query: 369 GPDIFPAFKHTLKTISNDSNNGSRR 393
I A + T+K + +R
Sbjct: 431 -DAIRAALQATIKPVKRKKEGEEKR 502
>TC230887 similar to UP|Q9ZUM1 (Q9ZUM1) Expressed protein (At2g02170/F5O4.6),
partial (29%)
Length = 1085
Score = 28.5 bits (62), Expect = 7.6
Identities = 15/56 (26%), Positives = 28/56 (49%)
Frame = +1
Query: 284 PAPVKGENLTAEENEKLAKEEKRLRNKVASLIKKQKKQQALGIVKGRDQAKPWGQE 339
PAP +N T E+++ + KR ++ IK +++ ALG+ G+ W +
Sbjct: 214 PAPTPLDNTTDEDSQFPVENGKRNLSEEEMKIKTRREIAALGVQLGKMNIAAWASK 381
>TC207438 homologue to UP|Q08145 (Q08145) GTP-binding protein, partial (95%)
Length = 853
Score = 28.5 bits (62), Expect = 7.6
Identities = 15/33 (45%), Positives = 18/33 (54%)
Frame = -2
Query: 5 LAKRASSRRFILPSQSSTSLRFSHQHNFPEKIL 37
+ + SS +LPS TSL FS H FP IL
Sbjct: 570 IKNKPSSSARLLPSSVLTSLIFSKSHLFPTNIL 472
>TC210028 similar to UP|Q9LIR7 (Q9LIR7) Arabidopsis thaliana genomic DNA,
chromosome 3, BAC clone:F14O13, partial (3%)
Length = 780
Score = 28.5 bits (62), Expect = 7.6
Identities = 21/46 (45%), Positives = 26/46 (55%), Gaps = 7/46 (15%)
Frame = +1
Query: 277 KSDSDSVPAPVK-----GENLTAEENEKLAKE--EKRLRNKVASLI 315
KS SDS A VK GEN T+EEN KL + ++ +RN S I
Sbjct: 592 KSPSDSGSAEVKHHLNDGENATSEENSKLFGDVFQEPIRNAKDSTI 729
>TC215154 similar to UP|Q6SZ89 (Q6SZ89) Translational elongation factor 1
subunit Bbeta, complete
Length = 1109
Score = 28.1 bits (61), Expect = 9.9
Identities = 21/73 (28%), Positives = 30/73 (40%)
Frame = +3
Query: 271 KKATSHKSDSDSVPAPVKGENLTAEENEKLAKEEKRLRNKVASLIKKQKKQQALGIVKGR 330
K A + D D V +L EE E EEK+ + A+ +K K++ G
Sbjct: 351 KAAVAEDDDDDDV-------DLFGEETE----EEKKAAEERAAAVKASTKKKESGKSSVL 497
Query: 331 DQAKPWGQEAQVK 343
KPW E +K
Sbjct: 498 LDVKPWDDETDMK 536
>BE660082
Length = 846
Score = 28.1 bits (61), Expect = 9.9
Identities = 16/49 (32%), Positives = 27/49 (54%), Gaps = 4/49 (8%)
Frame = +2
Query: 119 KNPQLEEKKKNKNNSYKYKMLRRRQIKIET----EAWEEAAREYQELLE 163
KN +LEE+ N +++ +L + +++ E E EA +E Q LLE
Sbjct: 425 KNKKLEEEYSNLKKNHEATLLEKCRLETEVLKLKEQLSEAEKEIQRLLE 571
>TC215153 similar to UP|Q6SZ89 (Q6SZ89) Translational elongation factor 1
subunit Bbeta, complete
Length = 1185
Score = 28.1 bits (61), Expect = 9.9
Identities = 21/73 (28%), Positives = 30/73 (40%)
Frame = +3
Query: 271 KKATSHKSDSDSVPAPVKGENLTAEENEKLAKEEKRLRNKVASLIKKQKKQQALGIVKGR 330
K A + D D V +L EE E EEK+ + A+ +K K++ G
Sbjct: 390 KAAVAEDDDDDDV-------DLFGEETE----EEKKAAEERAAAVKASAKKKESGKSSVL 536
Query: 331 DQAKPWGQEAQVK 343
KPW E +K
Sbjct: 537 LDVKPWDDETDMK 575
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.316 0.132 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,124,170
Number of Sequences: 63676
Number of extensions: 275835
Number of successful extensions: 1809
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 1763
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1800
length of query: 515
length of database: 12,639,632
effective HSP length: 101
effective length of query: 414
effective length of database: 6,208,356
effective search space: 2570259384
effective search space used: 2570259384
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC144726.11


