

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144724.5 - phase: 0 /pseudo
(534 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
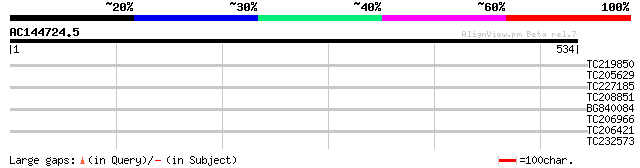
Score E
Sequences producing significant alignments: (bits) Value
TC219850 weakly similar to UP|Q9FKI5 (Q9FKI5) Emb|CAB69839.1, pa... 31 1.6
TC205629 similar to UP|Q9LEX1 (Q9LEX1) CaLB protein (At3g61050),... 30 2.7
TC227185 homologue to UP|Q6RUF6 (Q6RUF6) Fructose-bisphosphate a... 30 3.5
TC208851 UP|Q7PG82 (Q7PG82) ENSANGP00000024412 (Fragment), parti... 30 3.5
BG840084 29 4.6
TC206966 similar to UP|Q94EI3 (Q94EI3) AT5g17550/K10A8_30, parti... 29 4.6
TC206421 homologue to UP|Q93XV7 (Q93XV7) Hydroxypyruvate reducta... 29 6.0
TC232573 28 7.8
>TC219850 weakly similar to UP|Q9FKI5 (Q9FKI5) Emb|CAB69839.1, partial (53%)
Length = 1153
Score = 30.8 bits (68), Expect = 1.6
Identities = 14/35 (40%), Positives = 20/35 (57%)
Frame = -1
Query: 465 IFNYLQKDLRCTTSNFFPTMELSMSDLIGICIIKK 499
IF Y K L+ SN+ P +S SDL +C I++
Sbjct: 1063 IFXYTSKCLKKQASNYSPVSNISASDLATVCKIER 959
>TC205629 similar to UP|Q9LEX1 (Q9LEX1) CaLB protein (At3g61050), partial (92%)
Length = 1766
Score = 30.0 bits (66), Expect = 2.7
Identities = 18/69 (26%), Positives = 24/69 (34%)
Frame = +3
Query: 369 WTWC*ACRG*IPWVRY*WIGRLSLCNSCMKEKW*RCWGKVASRRNTVISTHFWGTNKVGW 428
W WC CR C SC K W CW + +R+ S + W + W
Sbjct: 1230 WYWC-CCR----------------CRSCGKWYW--CWSRACWKRHRCWSWYCWEWSWSRW 1352
Query: 429 EWIGGGLNI 437
W G +
Sbjct: 1353 *WTEQGREV 1379
>TC227185 homologue to UP|Q6RUF6 (Q6RUF6) Fructose-bisphosphate aldolase ,
complete
Length = 1435
Score = 29.6 bits (65), Expect = 3.5
Identities = 17/59 (28%), Positives = 25/59 (41%), Gaps = 8/59 (13%)
Frame = -1
Query: 401 W*RCWGKVASRRNTVISTHFW--------GTNKVGWEWIGGGLNIISLRVMKLQCQKSF 451
W R W A N V+S H W G+N VG E +++ + + C+ SF
Sbjct: 826 WHRSWDSPAECLNCVLSNHLWWYLWAIRAGSNHVGLEKSSFKKDMLLIECLVNSCKHSF 650
>TC208851 UP|Q7PG82 (Q7PG82) ENSANGP00000024412 (Fragment), partial (25%)
Length = 482
Score = 29.6 bits (65), Expect = 3.5
Identities = 16/40 (40%), Positives = 23/40 (57%)
Frame = +2
Query: 186 KKKERTRQDLRSGKGSATSTTTRWRKEGPRAYASSVGVSI 225
+++ER R+ R S STT R GPR+ A+ V VS+
Sbjct: 65 RERERERERERERASSKPSTTGESRGRGPRSSATPVTVSL 184
>BG840084
Length = 381
Score = 29.3 bits (64), Expect = 4.6
Identities = 9/19 (47%), Positives = 13/19 (68%)
Frame = +2
Query: 420 FWGTNKVGWEWIGGGLNII 438
FW N GWEW G G++++
Sbjct: 227 FWYCNGTGWEWPGDGVDLL 283
>TC206966 similar to UP|Q94EI3 (Q94EI3) AT5g17550/K10A8_30, partial (87%)
Length = 1018
Score = 29.3 bits (64), Expect = 4.6
Identities = 15/32 (46%), Positives = 17/32 (52%), Gaps = 2/32 (6%)
Frame = +3
Query: 43 WKILSKNWQH*NRVEMLKNLWRP--LNFCPRR 72
WKI S N + LWRP +NFCPRR
Sbjct: 384 WKIGSNNLSSLLALRTWNLLWRP*CINFCPRR 479
>TC206421 homologue to UP|Q93XV7 (Q93XV7) Hydroxypyruvate reductase , complete
Length = 1550
Score = 28.9 bits (63), Expect = 6.0
Identities = 18/66 (27%), Positives = 29/66 (43%), Gaps = 3/66 (4%)
Frame = +1
Query: 434 GLNIISLRVMKLQCQKSFL---QPWNSTLQFLRIIFNYLQKDLRCTTSNFFPTMELSMSD 490
GLN S+R++ Q L +PW LQ + + Y + CT F S++
Sbjct: 1114 GLNHSSMRMLNHQLHPQALLMQKPWVCQLQSYKHTWFYPTRAFNCTNDQFLSVELGSVAS 1293
Query: 491 LIGICI 496
L+ C+
Sbjct: 1294 LLFFCV 1311
>TC232573
Length = 497
Score = 28.5 bits (62), Expect = 7.8
Identities = 13/30 (43%), Positives = 17/30 (56%)
Frame = -3
Query: 386 WIGRLSLCNSCMKEKW*RCWGKVASRRNTV 415
W+ S SC+ K RCW KVAS + T+
Sbjct: 336 WLLHWSQGISCVAHKRIRCWKKVASSQTTI 247
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.357 0.157 0.630
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 29,040,890
Number of Sequences: 63676
Number of extensions: 447457
Number of successful extensions: 5010
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 2712
Number of HSP's successfully gapped in prelim test: 192
Number of HSP's that attempted gapping in prelim test: 2228
Number of HSP's gapped (non-prelim): 3030
length of query: 534
length of database: 12,639,632
effective HSP length: 102
effective length of query: 432
effective length of database: 6,144,680
effective search space: 2654501760
effective search space used: 2654501760
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.7 bits)
S2: 61 (28.1 bits)
Medicago: description of AC144724.5


