

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144721.3 - phase: 0 /pseudo
(750 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
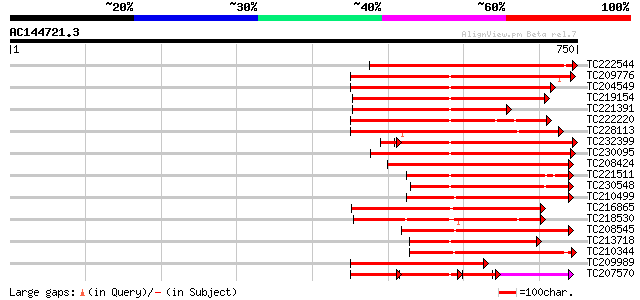
Score E
Sequences producing significant alignments: (bits) Value
TC222544 weakly similar to UP|O23743 (O23743) SFR1 protein precu... 429 e-120
TC209776 similar to UP|O65470 (O65470) Serine/threonine kinase-l... 341 8e-94
TC204549 similar to UP|Q9M0X5 (Q9M0X5) Receptor protein kinase-l... 329 2e-90
TC219154 similar to PIR|T05754|T05754 S-receptor kinase M4I22.1... 321 9e-88
TC221391 similar to UP|O81906 (O81906) Serine/threonine kinase-l... 310 1e-84
TC222220 weakly similar to UP|Q9ZP16 (Q9ZP16) Receptor-like prot... 299 3e-81
TC228113 similar to UP|Q9XED4 (Q9XED4) Receptor-like protein kin... 295 5e-80
TC232399 similar to UP|Q39086 (Q39086) Receptor kinase , partial... 288 6e-78
TC230095 weakly similar to UP|Q9XED4 (Q9XED4) Receptor-like prot... 259 4e-69
TC208424 weakly similar to UP|O65466 (O65466) Serine/threonine k... 256 2e-68
TC221511 similar to UP|Q40096 (Q40096) Receptor protein kinase, ... 249 2e-66
TC230548 weakly similar to UP|Q8LLI4 (Q8LLI4) S-locus receptor-l... 228 6e-60
TC210499 weakly similar to UP|Q9LDS6 (Q9LDS6) Serine/threonine k... 227 2e-59
TC216865 similar to UP|Q9LPG0 (Q9LPG0) T3F20.24 protein, partial... 225 6e-59
TC218530 similar to UP|Q9SGU0 (Q9SGU0) T6H22.8.1 protein, partia... 225 6e-59
TC208545 weakly similar to UP|Q9S972 (Q9S972) ARK2 product/recep... 224 1e-58
TC213718 similar to UP|Q70I29 (Q70I29) S-receptor kinase-like pr... 224 1e-58
TC210344 similar to UP|Q9AVE0 (Q9AVE0) SRKb, partial (21%) 223 3e-58
TC209989 similar to UP|Q9XED4 (Q9XED4) Receptor-like protein kin... 222 4e-58
TC207570 weakly similar to UP|O65470 (O65470) Serine/threonine k... 102 3e-57
>TC222544 weakly similar to UP|O23743 (O23743) SFR1 protein precursor ,
partial (27%)
Length = 861
Score = 429 bits (1104), Expect = e-120
Identities = 214/275 (77%), Positives = 240/275 (86%)
Frame = +3
Query: 476 EVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPNKSLDAFLFDPIQKKKLDWRKRSNIV 535
EVVVISKLQHRNLVRLLGCC+ER E+MLVYEFMPNKSLD+FLFDP+Q+K LDW+KR NI+
Sbjct: 3 EVVVISKLQHRNLVRLLGCCIERDEQMLVYEFMPNKSLDSFLFDPLQRKILDWKKRFNII 182
Query: 536 EGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMIPKISDFGLARIVKGGEGDEANTKRV 595
EGIARGI+YLHRDSRL+IIHRDLKASNILLDDEM PKISDFGLARIV+ G+ DEANTKRV
Sbjct: 183 EGIARGILYLHRDSRLRIIHRDLKASNILLDDEMHPKISDFGLARIVRSGDDDEANTKRV 362
Query: 596 VGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSGRRNNSFYQNEDSLSLVGFAWKLWL 655
VGTYGYMPPEYAM G+FSEKSDVY VLLLEIVSGRRN SFY NE SLSL +AWKLW
Sbjct: 363 VGTYGYMPPEYAMEGIFSEKSDVYXXXVLLLEIVSGRRNTSFYNNEQSLSLXXYAWKLWN 542
Query: 656 EENTISLIDREVWDASFESSMLRCMHIGLLCVQELPKERPSISTVVLMLISEITHLPPPG 715
E N S+ID E+ D FE S+LRC+HIGLLCVQEL KERP+ISTVV MLISEITHLPPP
Sbjct: 543 EGNIKSIIDLEIQDPMFEKSILRCIHIGLLCVQELTKERPTISTVVXMLISEITHLPPPR 722
Query: 716 KVAFVHNQNSRSTESSQQSHRSNSNNNVTLSDVIG 750
+ FV QN +S+ESSQ+S + NSNNNVT+S++ G
Sbjct: 723 QXXFVQKQNCQSSESSQKS-QFNSNNNVTISEIQG 824
>TC209776 similar to UP|O65470 (O65470) Serine/threonine kinase-like protein
, partial (47%)
Length = 1305
Score = 341 bits (874), Expect = 8e-94
Identities = 175/305 (57%), Positives = 227/305 (74%), Gaps = 7/305 (2%)
Frame = +2
Query: 451 IQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPN 510
+ GQE+AVKRLSK SGQG EEF NEV +++KLQHRNLVRLLG C+E EK+LVYEF+ N
Sbjct: 137 LPSGQEVAVKRLSKISGQGGEEFKNEVEIVAKLQHRNLVRLLGFCLEGEEKILVYEFVAN 316
Query: 511 KSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMI 570
KSLD LFDP ++K LDW +R IVEGIARGI YLH DSRLKIIHRDLKASN+LLD +M
Sbjct: 317 KSLDYILFDPEKQKSLDWTRRYKIVEGIARGIQYLHEDSRLKIIHRDLKASNVLLDGDMN 496
Query: 571 PKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVS 630
PKISDFG+ARI G + +ANT R+VGTYGYM PEYAM G +S KSDVYSFGVL+LEI+S
Sbjct: 497 PKISDFGMARIF-GVDQTQANTNRIVGTYGYMSPEYAMHGEYSAKSDVYSFGVLILEIIS 673
Query: 631 GRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLCVQEL 690
G+RN+SFY+ + + L+ +AWKLW +E + L+D+ + ++ + ++RC+HIGLLCVQE
Sbjct: 674 GKRNSSFYETDVAEDLLSYAWKLWKDEAPLELMDQSLRESYTRNEVIRCIHIGLLCVQED 853
Query: 691 PKERPSISTVVLMLISEITHLPPPGKVAFVHNQNSR-------STESSQQSHRSNSNNNV 743
P +RP++++VVLML S L P + AF N + + S + S S N++
Sbjct: 854 PIDRPTMASVVLMLDSYSVTLQVPNQPAFYINSRTEPNMPKGLKIDQSTTNSTSKSVNDM 1033
Query: 744 TLSDV 748
++S+V
Sbjct: 1034SVSEV 1048
>TC204549 similar to UP|Q9M0X5 (Q9M0X5) Receptor protein kinase-like protein,
partial (54%)
Length = 2032
Score = 329 bits (844), Expect = 2e-90
Identities = 165/273 (60%), Positives = 215/273 (78%), Gaps = 1/273 (0%)
Frame = +3
Query: 451 IQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPN 510
+ GQ +AVKRLSK+SGQG EEF NEVVV++KLQHRNLVRLLG C++ EK+LVYE++PN
Sbjct: 885 LSSGQVVAVKRLSKSSGQGGEEFKNEVVVVAKLQHRNLVRLLGFCLQGEEKILVYEYVPN 1064
Query: 511 KSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMI 570
KSLD LFDP ++++LDW +R I+ GIARGI YLH DSRL+IIHRDLKASNILLD +M
Sbjct: 1065 KSLDYILFDPEKQRELDWGRRYKIIGGIARGIQYLHEDSRLRIIHRDLKASNILLDGDMN 1244
Query: 571 PKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVS 630
PKISDFG+ARI G + + NT R+VGTYGYM PEYAM G FS KSDVYSFGVLL+EI+S
Sbjct: 1245 PKISDFGMARIF-GVDQTQGNTSRIVGTYGYMAPEYAMHGEFSVKSDVYSFGVLLMEILS 1421
Query: 631 GRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLCVQEL 690
G++N+SFYQ + + L+ +AW+LW + + L+D + ++ ++ ++R +HIGLLCVQE
Sbjct: 1422 GKKNSSFYQTDGAEDLLSYAWQLWKDGTPLELMDPILRESYNQNEVIRSIHIGLLCVQED 1601
Query: 691 PKERPSISTVVLMLISEITHLPPPGKVA-FVHN 722
P +RP+++T+VLML S LP P + A FVH+
Sbjct: 1602 PADRPTMATIVLMLDSNTVTLPTPTQPAFFVHS 1700
>TC219154 similar to PIR|T05754|T05754 S-receptor kinase M4I22.110 precursor
- Arabidopsis thaliana {Arabidopsis thaliana;} ,
partial (31%)
Length = 1363
Score = 321 bits (822), Expect = 9e-88
Identities = 156/260 (60%), Positives = 204/260 (78%)
Frame = +1
Query: 454 GQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPNKSL 513
GQEIAVKRLS SGQGI EF+ EV +I+KLQHRNLV+LLGCC++ EK+LVYE++ N SL
Sbjct: 415 GQEIAVKRLSSLSGQGITEFITEVKLIAKLQHRNLVKLLGCCIKGQEKLLVYEYVVNGSL 594
Query: 514 DAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMIPKI 573
++F+FD I+ K LDW +R NI+ GIARG++YLH+DSRL+IIHRDLKASN+LLD+++ PKI
Sbjct: 595 NSFIFDQIKSKLLDWPRRFNIILGIARGLLYLHQDSRLRIIHRDLKASNVLLDEKLNPKI 774
Query: 574 SDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSGRR 633
SDFG+AR GG+ E NT RVVGTYGYM PEYA G FS KSDV+SFG+LLLEIV G +
Sbjct: 775 SDFGMARAF-GGDQTEGNTNRVVGTYGYMAPEYAFDGNFSIKSDVFSFGILLLEIVCGIK 951
Query: 634 NNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLCVQELPKE 693
N S +L+LVG+AW LW E+N + LID + D+ +LRC+H+ LLCVQ+ P++
Sbjct: 952 NKSLCHENQTLNLVGYAWALWKEQNALQLIDSGIKDSCVIPEVLRCIHVSLLCVQQYPED 1131
Query: 694 RPSISTVVLMLISEITHLPP 713
RP++++V+ ML SE+ + P
Sbjct: 1132RPTMTSVIQMLGSEMDMVEP 1191
>TC221391 similar to UP|O81906 (O81906) Serine/threonine kinase-like protein
, partial (25%)
Length = 642
Score = 310 bits (795), Expect = 1e-84
Identities = 151/211 (71%), Positives = 181/211 (85%)
Frame = +2
Query: 454 GQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPNKSL 513
G+E+AVKRLS+ S QG+EEF NE+V+I+KLQHRNLVRLLGCC++ EK+LVYE++PNKSL
Sbjct: 2 GEEVAVKRLSRKSSQGLEEFKNEMVLIAKLQHRNLVRLLGCCIQGEEKILVYEYLPNKSL 181
Query: 514 DAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMIPKI 573
D FLFDP+++ +LDW KR I+EGIARG++YLHRDSRL+IIHRDLKASNILLD+ M PKI
Sbjct: 182 DCFLFDPVKQTQLDWAKRFEIIEGIARGLLYLHRDSRLRIIHRDLKASNILLDESMNPKI 361
Query: 574 SDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSGRR 633
SDFGLARI GG +EANT RVVGTYGYM PEYAM GLFS KSDVYSFGVLLLEI+SGR+
Sbjct: 362 SDFGLARIF-GGNQNEANTNRVVGTYGYMSPEYAMEGLFSIKSDVYSFGVLLLEIMSGRK 538
Query: 634 NNSFYQNEDSLSLVGFAWKLWLEENTISLID 664
N SF +DS SL+G+AW LW E+ + L+D
Sbjct: 539 NTSFRDTDDS-SLIGYAWHLWSEQRVMELVD 628
>TC222220 weakly similar to UP|Q9ZP16 (Q9ZP16) Receptor-like protein kinase,
RLK3 precursor, partial (35%)
Length = 1304
Score = 299 bits (766), Expect = 3e-81
Identities = 153/269 (56%), Positives = 201/269 (73%), Gaps = 3/269 (1%)
Frame = +3
Query: 451 IQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPN 510
+ DG+EIAVK+LSK+SGQG EF NE+++I+KLQHRNLV LLG C+E EKML+YEF+ N
Sbjct: 282 LPDGREIAVKKLSKSSGQGANEFKNEILLIAKLQHRNLVTLLGFCLEEHEKMLIYEFVSN 461
Query: 511 KSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMI 570
KSLD FLFD + K+L+W +R I+EGIA+GI YLH SRLK+IHRDLK SN+LLD M
Sbjct: 462 KSLDYFLFDSHRSKQLNWSERYKIIEGIAQGISYLHEHSRLKVIHRDLKPSNVLLDSNMN 641
Query: 571 PKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVS 630
PKISDFG+ARIV + + T R+VGTYGYM PEYAM G FSEKSDV+SFGV++LEI+S
Sbjct: 642 PKISDFGMARIV-AIDQLQGKTNRIVGTYGYMSPEYAMHGQFSEKSDVFSFGVIVLEIIS 818
Query: 631 GRRN-NSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASF--ESSMLRCMHIGLLCV 687
+RN S + + D L+ + W+ W++E +++ D+ + +A F S +++C+ IGLLCV
Sbjct: 819 AKRNTRSVFSDHD--DLLSYTWEQWMDEAPLNIFDQSI-EAEFCDHSEVVKCIQIGLLCV 989
Query: 688 QELPKERPSISTVVLMLISEITHLPPPGK 716
QE P +RP I+ V+ L S IT LP P K
Sbjct: 990 QEKPDDRPKITQVISYLNSSITELPLPKK 1076
>TC228113 similar to UP|Q9XED4 (Q9XED4) Receptor-like protein kinase homolog
RK20-1, partial (53%)
Length = 1492
Score = 295 bits (755), Expect = 5e-80
Identities = 156/318 (49%), Positives = 210/318 (65%), Gaps = 36/318 (11%)
Frame = +1
Query: 451 IQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPN 510
+ DGQ IAVKRLS+ S QG EF NEV++++KLQHRNLVRLLG C+E E++L+YE++PN
Sbjct: 211 LSDGQMIAVKRLSRESSQGDTEFKNEVLLVAKLQHRNLVRLLGFCLEGKERLLIYEYVPN 390
Query: 511 KSLDAFLF------------------------------------DPIQKKKLDWRKRSNI 534
KSLD F+F DP +K +L+W R I
Sbjct: 391 KSLDYFIFGRS**VHN*TVALYFTSGSGQRLNIHIPLKLMAVYADPTKKAQLNWEMRYKI 570
Query: 535 VEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMIPKISDFGLARIVKGGEGDEANTKR 594
+ G+ARG++YLH DS L+IIHRDLKASNILL++EM PKI+DFG+AR+V + +ANT R
Sbjct: 571 ITGVARGLLYLHEDSHLRIIHRDLKASNILLNEEMNPKIADFGMARLVLMDQ-TQANTNR 747
Query: 595 VVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSGRRNNSFYQNEDSLSLVGFAWKLW 654
+VGTYGYM PEYAM G FS KSDV+SFGVL+LEI+SG +N+ E+ L+ FAW+ W
Sbjct: 748 IVGTYGYMAPEYAMHGQFSMKSDVFSFGVLVLEIISGHKNSGIRHGENVEDLLSFAWRNW 927
Query: 655 LEENTISLIDREVWDASFESSMLRCMHIGLLCVQELPKERPSISTVVLMLISEITHLPPP 714
E + ++D + + S + MLRC+HIGLLCVQE +RP+++T++LML S LP P
Sbjct: 928 REGTAVKIVDPSLNNNS-RNEMLRCIHIGLLCVQENLADRPTMTTIMLMLNSYSLSLPIP 1104
Query: 715 GKVAFVHNQNSRSTESSQ 732
K AF + + S ++Q
Sbjct: 1105SKPAFYVSSRTGSISATQ 1158
>TC232399 similar to UP|Q39086 (Q39086) Receptor kinase , partial (23%)
Length = 1167
Score = 288 bits (737), Expect = 6e-78
Identities = 141/243 (58%), Positives = 187/243 (76%), Gaps = 2/243 (0%)
Frame = +2
Query: 510 NKSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEM 569
N +L+ + DP + K LDWRKR I+EGI RG++YLHRDSRLKIIHRDLKASN+LL + +
Sbjct: 299 NATLNVYFLDPSKSKLLDWRKRCGIIEGIGRGLLYLHRDSRLKIIHRDLKASNVLLYEAL 478
Query: 570 IPKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIV 629
PKISDFG+ARI GG D+ANT RVVGTYGYM PEYAM GLFSEKSDV+SFGVL++EIV
Sbjct: 479 NPKISDFGMARIF-GGTEDQANTNRVVGTYGYMSPEYAMQGLFSEKSDVFSFGVLVIEIV 655
Query: 630 SGRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLCVQE 689
SGRRN+ FY ++++LSL+GFAW W E N +S+ID E++D + +LRC+HIGLLCVQE
Sbjct: 656 SGRRNSRFYDDDNALSLLGFAWIQWREGNILSVIDPEIYDVTHHKDILRCIHIGLLCVQE 835
Query: 690 LPKERPSISTVVLMLISEITHLPPPGKVAFVHNQNSRS--TESSQQSHRSNSNNNVTLSD 747
+RP+++ V+ ML SE+ LPPP + AFV +QN + + SS++ + S N ++++D
Sbjct: 836 RAVDRPTMAAVISMLNSEVAFLPPPDQPAFVQSQNMLNLVSVSSEERQKLCSINGISITD 1015
Query: 748 VIG 750
+ G
Sbjct: 1016IRG 1024
Score = 46.2 bits (108), Expect = 5e-05
Identities = 19/28 (67%), Positives = 22/28 (77%)
Frame = +3
Query: 491 LLGCCVERGEKMLVYEFMPNKSLDAFLF 518
L GCC E EKML+YE+M NKSLD F+F
Sbjct: 3 LFGCCAEGDEKMLIYEYMLNKSLDVFIF 86
>TC230095 weakly similar to UP|Q9XED4 (Q9XED4) Receptor-like protein kinase
homolog RK20-1, partial (30%)
Length = 819
Score = 259 bits (661), Expect = 4e-69
Identities = 132/271 (48%), Positives = 188/271 (68%)
Frame = +3
Query: 478 VVISKLQHRNLVRLLGCCVERGEKMLVYEFMPNKSLDAFLFDPIQKKKLDWRKRSNIVEG 537
++I+KLQH+NLVRL+G C E EK+L+YE++PNKSLD FLFD + ++L W +R IV+G
Sbjct: 6 LLIAKLQHKNLVRLVGFCQEDREKILIYEYVPNKSLDHFLFDSQKHRQLTWSERFKIVKG 185
Query: 538 IARGIMYLHRDSRLKIIHRDLKASNILLDDEMIPKISDFGLARIVKGGEGDEANTKRVVG 597
IARGI+YLH DSRLKIIHRD+K SN+LLD + PKISDFG+AR+V + + T RVVG
Sbjct: 186 IARGILYLHEDSRLKIIHRDIKPSNVLLDYGINPKISDFGMARMV-ATDQIQGCTNRVVG 362
Query: 598 TYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSGRRNNSFYQNEDSLSLVGFAWKLWLEE 657
TYGYM PEYAM G FSEKSDV+SFGV++L+I+SG++N+ +++ L+ +AW W +E
Sbjct: 363 TYGYMSPEYAMHGQFSEKSDVFSFGVMVLDIISGKKNSCSFESCRVDDLLSYAWNNWRDE 542
Query: 658 NTISLIDREVWDASFESSMLRCMHIGLLCVQELPKERPSISTVVLMLISEITHLPPPGKV 717
+ L+D + ++ + + +CM IGLLCVQE P +RP++ T+V L + +P P +
Sbjct: 543 SPYQLLDSTLLESYVPNEVEKCMQIGLLCVQENPDDRPTMGTIVSYLSNPSFEMPFPLEP 722
Query: 718 AFVHNQNSRSTESSQQSHRSNSNNNVTLSDV 748
AF + R + +S N+ + S V
Sbjct: 723 AFFMHGRMRRHSAEHESSSGYYTNHPSSSSV 815
>TC208424 weakly similar to UP|O65466 (O65466) Serine/threonine kinase-like
protein , partial (34%)
Length = 913
Score = 256 bits (655), Expect = 2e-68
Identities = 127/248 (51%), Positives = 173/248 (69%), Gaps = 1/248 (0%)
Frame = +2
Query: 500 EKMLVYEFMPNKSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLK 559
EK+L+YE++ NKSLD FLFDP+++K+LDW +R I+ GIARGI YLH DS+L+IIHRDLK
Sbjct: 2 EKILIYEYITNKSLDHFLFDPVKQKELDWSRRYKIIVGIARGIQYLHEDSQLRIIHRDLK 181
Query: 560 ASNILLDDEMIPKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVY 619
ASN+LLD+ M PKISDFG+A+I + + + NT R+VGTYGYM PEYAM G FS KSDV+
Sbjct: 182 ASNVLLDENMNPKISDFGMAKIFQADQ-TQVNTGRIVGTYGYMSPEYAMRGQFSVKSDVF 358
Query: 620 SFGVLLLEIVSGRRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRC 679
SFGVL+LEIVSG++N FYQ+ + L+ AWK W + + L+D + + + + RC
Sbjct: 359 SFGVLVLEIVSGKKNTDFYQSNHADDLLSHAWKNWTLQTPLELLDPTLRGSYSRNEVNRC 538
Query: 680 MHIGLLCVQELPKERPSISTVVLMLIS-EITHLPPPGKVAFVHNQNSRSTESSQQSHRSN 738
+HIGLLCVQE P +RPS++T+ LML S +T P + + + S +S
Sbjct: 539 IHIGLLCVQENPSDRPSMATIALMLNSYSVTMSMPQQPASLLRGRGPNRLNQGMDSDQST 718
Query: 739 SNNNVTLS 746
+N + T S
Sbjct: 719 TNQSTTCS 742
>TC221511 similar to UP|Q40096 (Q40096) Receptor protein kinase, partial
(21%)
Length = 765
Score = 249 bits (637), Expect = 2e-66
Identities = 124/221 (56%), Positives = 164/221 (74%)
Frame = +3
Query: 526 LDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMIPKISDFGLARIVKGG 585
LDW KR NI+ GIA+G++YLH+DSRL+IIHRDLKASN+LLD E+ PKISDFG+ARI G
Sbjct: 3 LDWSKRFNIICGIAKGLLYLHQDSRLRIIHRDLKASNVLLDSELNPKISDFGMARIF-GV 179
Query: 586 EGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSGRRNNSFYQNEDSLS 645
+ E NTKR+VGTYGYM PEYA GLFS KSDV+SFGVLLLEI+SG+R+ +Y S +
Sbjct: 180 DQQEGNTKRIVGTYGYMAPEYATDGLFSVKSDVFSFGVLLLEIISGKRSRGYYNQNHSQN 359
Query: 646 LVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLCVQELPKERPSISTVVLMLI 705
L+G AWKLW E + LID+ + D+S S ML C+H+ LLCVQ+ P++RP +S+V+LML+
Sbjct: 360 LIGHAWKLWKEGRPLELIDKSIEDSSSLSQMLHCIHVSLLCVQQNPEDRPGMSSVLLMLV 539
Query: 706 SEITHLPPPGKVAFVHNQNSRSTESSQQSHRSNSNNNVTLS 746
SE+ LP P + F + S +SS + +S N +T++
Sbjct: 540 SEL-ELPEPKQPGF-FGKYSGEADSSTSKQQLSSTNEITIT 656
>TC230548 weakly similar to UP|Q8LLI4 (Q8LLI4) S-locus receptor-like kinase
RLK14, partial (20%)
Length = 692
Score = 228 bits (582), Expect = 6e-60
Identities = 119/216 (55%), Positives = 155/216 (71%)
Frame = +3
Query: 531 RSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMIPKISDFGLARIVKGGEGDEA 590
R I+ GIARG++YLH+DSRL+IIHRDLKASN+LLD+EM PKISDFGLAR+ GG+ E
Sbjct: 3 RFGIINGIARGLLYLHQDSRLRIIHRDLKASNVLLDNEMNPKISDFGLARMC-GGDQIEG 179
Query: 591 NTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSGRRNNSFYQNEDSLSLVGFA 650
T RVVGTYGYM PEYA G+FS KSDV+SFGVLLLEIVSG++N+ + D +L+G A
Sbjct: 180 ETSRVVGTYGYMAPEYAFDGIFSIKSDVFSFGVLLLEIVSGKKNSRLFYPNDYNNLIGHA 359
Query: 651 WKLWLEENTISLIDREVWDASFESSMLRCMHIGLLCVQELPKERPSISTVVLMLISEITH 710
W LW E N + ID + D+ LRC+HIGLLCVQ P +RP++++VV++L +E
Sbjct: 360 WMLWKEGNPMQFIDSSLEDSCILYEALRCIHIGLLCVQHHPNDRPNMASVVVLLSNE-NA 536
Query: 711 LPPPGKVAFVHNQNSRSTESSQQSHRSNSNNNVTLS 746
LP P +++ S ESS ++ S S N+VT+S
Sbjct: 537 LPLPKDPSYLSKDISTERESSSKNFTSVSINDVTMS 644
>TC210499 weakly similar to UP|Q9LDS6 (Q9LDS6) Serine/threonine kinase-like
protein , partial (27%)
Length = 717
Score = 227 bits (578), Expect = 2e-59
Identities = 116/221 (52%), Positives = 154/221 (69%)
Frame = +3
Query: 526 LDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMIPKISDFGLARIVKGG 585
LDW+KR NI+EGI++GI+YLH+ SRLKIIHRDLKASNILLD+ M PKISDFGLAR+
Sbjct: 15 LDWKKRFNIIEGISQGILYLHKYSRLKIIHRDLKASNILLDENMNPKISDFGLARMFMQQ 194
Query: 586 EGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSGRRNNSFYQNEDSLS 645
E T R+VGTYGYM PEYAM G FS KSDVYSFGVLLLEIVSGR+N SFY + L+
Sbjct: 195 EST-GTTSRIVGTYGYMSPEYAMEGTFSTKSDVYSFGVLLLEIVSGRKNTSFYDVDHLLN 371
Query: 646 LVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLCVQELPKERPSISTVVLMLI 705
L+G AW+LW + ++ L+D + D+ + RC+H+GLLCV+ +RP++S V+ ML
Sbjct: 372 LIGHAWELWNQGESLQLLDPSLNDSFDPDEVKRCIHVGLLCVEHYANDRPTMSNVISMLT 551
Query: 706 SEITHLPPPGKVAFVHNQNSRSTESSQQSHRSNSNNNVTLS 746
+E + P + AF + + ++S + +S + T S
Sbjct: 552 NESAPVTLPRRPAFYVERKNFDGKTSSKELCVDSTDEFTAS 674
>TC216865 similar to UP|Q9LPG0 (Q9LPG0) T3F20.24 protein, partial (28%)
Length = 1358
Score = 225 bits (573), Expect = 6e-59
Identities = 122/257 (47%), Positives = 171/257 (66%), Gaps = 1/257 (0%)
Frame = +1
Query: 453 DGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPNKS 512
DG IAVK+LS S QG EF+NE+ +IS LQH +LV+L GCCVE + +LVYE+M N S
Sbjct: 58 DGTLIAVKQLSSKSRQGNREFLNEIGMISALQHPHLVKLYGCCVEGDQLLLVYEYMENNS 237
Query: 513 LDAFLFDPIQKK-KLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMIP 571
L LF + + KLDW R I GIARG+ YLH +SRLKI+HRD+KA+N+LLD ++ P
Sbjct: 238 LARALFGAEEHQIKLDWTTRYKICVGIARGLAYLHEESRLKIVHRDIKATNVLLDQDLNP 417
Query: 572 KISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSG 631
KISDFGLA++ + E + + R+ GT+GYM PEYAM G ++K+DVYSFG++ LEI++G
Sbjct: 418 KISDFGLAKLDE--EDNTHISTRIAGTFGYMAPEYAMHGYLTDKADVYSFGIVALEIING 591
Query: 632 RRNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLCVQELP 691
R N Q E+S S++ +A L + + + L+DR + + L + + LLC
Sbjct: 592 RSNTIHRQKEESFSVLEWAHLLREKGDIMDLVDRRLGLEFNKEEALVMIKVALLCTNVTA 771
Query: 692 KERPSISTVVLMLISEI 708
RP++S+VV ML +I
Sbjct: 772 ALRPTMSSVVSMLEGKI 822
>TC218530 similar to UP|Q9SGU0 (Q9SGU0) T6H22.8.1 protein, partial (25%)
Length = 1190
Score = 225 bits (573), Expect = 6e-59
Identities = 125/256 (48%), Positives = 170/256 (65%), Gaps = 3/256 (1%)
Frame = +3
Query: 456 EIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPNKSLDA 515
EIAVK+LS S QG +F+ E+ IS +QHRNLV+L GCC+E +++LVYE++ NKSLD
Sbjct: 3 EIAVKQLSVGSHQGKSQFITEIATISAVQHRNLVKLYGCCIEGSKRLLVYEYLENKSLDQ 182
Query: 516 FLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMIPKISD 575
LF L+W R +I G+ARG+ YLH +SRL+I+HRD+KASNILLD E+IPKISD
Sbjct: 183 ALFGKCLT--LNWSTRYDICLGVARGLTYLHEESRLRIVHRDVKASNILLDYELIPKISD 356
Query: 576 FGLARIVKGGEGDEANT---KRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSGR 632
FGLA++ D+ T V GT GY+ PEYAM G +EK+DV+SFGV+ LE+VSGR
Sbjct: 357 FGLAKLY-----DDKKTHISTGVAGTIGYLAPEYAMRGHLTEKADVFSFGVVALELVSGR 521
Query: 633 RNNSFYQNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLCVQELPK 692
N+ + + L+ +AW+L + I L+D + + + E + R + I LLC Q P
Sbjct: 522 PNSDSSLEGEKVYLLEWAWQLHEKNCIIDLVDDRLSEFN-EEEVKRVVGIALLCTQTSPT 698
Query: 693 ERPSISTVVLMLISEI 708
RPS+S VV ML +I
Sbjct: 699 LRPSMSRVVAMLSGDI 746
>TC208545 weakly similar to UP|Q9S972 (Q9S972) ARK2 product/receptor-like
serine/threonine protein kinase ARK2, partial (20%)
Length = 1253
Score = 224 bits (571), Expect = 1e-58
Identities = 119/228 (52%), Positives = 153/228 (66%)
Frame = +2
Query: 519 DPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMIPKISDFGL 578
D + K LDW+KR NI+EGI++G++YLH+ SRLK+IHRDLKASNILLD+ M PKISDFGL
Sbjct: 326 DGTRSKLLDWKKRFNIIEGISQGLLYLHKYSRLKVIHRDLKASNILLDENMNPKISDFGL 505
Query: 579 ARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSGRRNNSFY 638
AR+ E NT R+VGTYGYM PEYAM G+FS KSDVYSFGVLLLEIVSGRRN SFY
Sbjct: 506 ARMFTRQEST-TNTSRIVGTYGYMSPEYAMEGVFSVKSDVYSFGVLLLEIVSGRRNTSFY 682
Query: 639 QNEDSLSLVGFAWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLCVQELPKERPSIS 698
+ L+L+G AW+LW E + LID + ++ + RC+HIGLLCV++ RP +S
Sbjct: 683 DGDRFLNLIGHAWELWNEGACLKLIDPSLTESPDLDEVQRCIHIGLLCVEQNANNRPLMS 862
Query: 699 TVVLMLISEITHLPPPGKVAFVHNQNSRSTESSQQSHRSNSNNNVTLS 746
++ ML S + P + AF + S ++S +T S
Sbjct: 863 QIISML-SNKNPITLPQRPAFYFGSETFDGIISSTEFCTDSTKAITTS 1003
>TC213718 similar to UP|Q70I29 (Q70I29) S-receptor kinase-like protein 2,
partial (20%)
Length = 525
Score = 224 bits (571), Expect = 1e-58
Identities = 108/174 (62%), Positives = 140/174 (80%)
Frame = +1
Query: 530 KRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMIPKISDFGLARIVKGGEGDE 589
KR I++GIARG++YLH+DSRL+IIHRDLK SNILLD++M PKISDFGLAR GG+ E
Sbjct: 7 KRLQIIDGIARGLLYLHQDSRLRIIHRDLKVSNILLDNDMNPKISDFGLARTF-GGDQAE 183
Query: 590 ANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSGRRNNSFYQNEDSLSLVGF 649
ANT RV+GTYGYMPPEYA+ G FS KSDV+SFGV++LEI+SGR+N +F +E L+L+
Sbjct: 184 ANTNRVMGTYGYMPPEYALHGRFSIKSDVFSFGVIVLEIISGRKNRNFQDSEHHLNLLSH 363
Query: 650 AWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLCVQELPKERPSISTVVLM 703
AW+LW+EE + LID + D +LRC+H+GLLCVQ+ P+ RP++S+VVLM
Sbjct: 364 AWRLWIEEKPLELIDDLLDDPVSPHEILRCIHVGLLCVQQTPENRPNMSSVVLM 525
>TC210344 similar to UP|Q9AVE0 (Q9AVE0) SRKb, partial (21%)
Length = 870
Score = 223 bits (567), Expect = 3e-58
Identities = 122/220 (55%), Positives = 152/220 (68%)
Frame = +2
Query: 530 KRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMIPKISDFGLARIVKGGEGDE 589
KR +I+ GIARGI+YLH DSRLKIIHRDLKASN+LLD +M PKISDFG+ARI G EG E
Sbjct: 5 KRLDIINGIARGILYLHEDSRLKIIHRDLKASNVLLDYDMNPKISDFGMARIFAGSEG-E 181
Query: 590 ANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSGRRNNSFYQNEDSLSLVGF 649
ANT +VGTYGYM PEYAM GL+S KSDV+ FGVLLLEI++G+RN FY + + SL+ +
Sbjct: 182 ANTATIVGTYGYMAPEYAMEGLYSIKSDVFGFGVLLLEIITGKRNAGFYHSNKTPSLLSY 361
Query: 650 AWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLCVQELPKERPSISTVVLMLISEIT 709
AW LW E + LID D+ LR MHIGLLCVQE +RP++S+VVLML +E
Sbjct: 362 AWHLWNEGKGLELIDPMSVDSCPGDEFLRYMHIGLLCVQEDAYDRPTMSSVVLMLKNESA 541
Query: 710 HLPPPGKVAFVHNQNSRSTESSQQSHRSNSNNNVTLSDVI 749
L P + F + + + Q S N +TLSD++
Sbjct: 542 TLGQPERPPFSLGRFNANEPDCQDC----SLNFLTLSDIV 649
>TC209989 similar to UP|Q9XED4 (Q9XED4) Receptor-like protein kinase homolog
RK20-1, partial (32%)
Length = 738
Score = 222 bits (566), Expect = 4e-58
Identities = 110/183 (60%), Positives = 141/183 (76%)
Frame = +3
Query: 451 IQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPN 510
+ DGQEIA+KRLS S QG EF NE+++ +LQHRNLVRLLG C R E++L+YEF+PN
Sbjct: 192 LSDGQEIAIKRLSINSNQGETEFKNEILLTGRLQHRNLVRLLGFCFARRERLLIYEFVPN 371
Query: 511 KSLDAFLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMI 570
KSLD F+FDP ++ L+ R I+ GIARG++YLH DSRL ++HRDLK SNILLD E+
Sbjct: 372 KSLDYFIFDPNKRVNLN*EIRYKIIRGIARGLLYLHEDSRLNVVHRDLKTSNILLDGELN 551
Query: 571 PKISDFGLARIVKGGEGDEANTKRVVGTYGYMPPEYAMGGLFSEKSDVYSFGVLLLEIVS 630
PKISDFG+AR+ + + EA+T +VGT+GYM PEY G FS KSDV+SFGV++LEIV
Sbjct: 552 PKISDFGMARLFEINQ-TEASTTTIVGTFGYMAPEYIKHGQFSIKSDVFSFGVMILEIVC 728
Query: 631 GRR 633
G+R
Sbjct: 729 GQR 737
>TC207570 weakly similar to UP|O65470 (O65470) Serine/threonine kinase-like
protein , partial (30%)
Length = 1285
Score = 102 bits (254), Expect(3) = 3e-57
Identities = 54/83 (65%), Positives = 62/83 (74%)
Frame = +1
Query: 516 FLFDPIQKKKLDWRKRSNIVEGIARGIMYLHRDSRLKIIHRDLKASNILLDDEMIPKISD 575
F D +K LDW KR NI+ GIA+G++YLH+ SRLK+IHRDLKASNILLD EM KISD
Sbjct: 295 FKTDSARKDLLDWEKRLNIIGGIAQGLLYLHKYSRLKVIHRDLKASNILLDHEMNAKISD 474
Query: 576 FGLARIVKGGEGDEANTKRVVGT 598
FG+ARI G E NT RVVGT
Sbjct: 475 FGMARIF-GVRVSEENTNRVVGT 540
Score = 93.2 bits (230), Expect(3) = 3e-57
Identities = 42/68 (61%), Positives = 57/68 (83%)
Frame = +2
Query: 451 IQDGQEIAVKRLSKASGQGIEEFMNEVVVISKLQHRNLVRLLGCCVERGEKMLVYEFMPN 510
+ D QE+A+KRLSK+SGQG+ EF NE +++KLQH NLV+LLG C++R E++LVYE+M N
Sbjct: 14 LSDQQEVAIKRLSKSSGQGLIEFTNEAKLMAKLQHTNLVKLLGFCIQRDERILVYEYMSN 193
Query: 511 KSLDAFLF 518
KSLD +LF
Sbjct: 194 KSLDFYLF 217
Score = 67.8 bits (164), Expect = 2e-11
Identities = 36/110 (32%), Positives = 60/110 (53%), Gaps = 2/110 (1%)
Frame = +1
Query: 639 QNEDSLSLVGF--AWKLWLEENTISLIDREVWDASFESSMLRCMHIGLLCVQELPKERPS 696
+ ED L ++ F AW+LW + LID + ++ + RC+HIGLLCVQ+ +RP+
Sbjct: 826 KGEDLLWMLEFL*AWQLWNAGRALELIDSTLNGLCSQNEVFRCIHIGLLCVQDQATDRPT 1005
Query: 697 ISTVVLMLISEITHLPPPGKVAFVHNQNSRSTESSQQSHRSNSNNNVTLS 746
+ +V L ++ LP P + A+ N+ +E +S N+VT+S
Sbjct: 1006MVDIVSFLSNDTIQLPQPMQPAYFINEVVEESELPYNQQEFHSENDVTIS 1155
Score = 66.6 bits (161), Expect(3) = 3e-57
Identities = 30/50 (60%), Positives = 41/50 (82%)
Frame = +2
Query: 600 GYMPPEYAMGGLFSEKSDVYSFGVLLLEIVSGRRNNSFYQNEDSLSLVGF 649
GYM PEYAM G+ S K+DV+SFGVLLLEI+S ++NNS Y ++ L+L+G+
Sbjct: 629 GYMAPEYAMKGVVSIKTDVFSFGVLLLEILSSKKNNSRYHSDHPLNLIGY 778
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.358 0.159 0.601
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 38,283,956
Number of Sequences: 63676
Number of extensions: 607174
Number of successful extensions: 9100
Number of sequences better than 10.0: 927
Number of HSP's better than 10.0 without gapping: 4998
Number of HSP's successfully gapped in prelim test: 295
Number of HSP's that attempted gapping in prelim test: 3221
Number of HSP's gapped (non-prelim): 5570
length of query: 750
length of database: 12,639,632
effective HSP length: 104
effective length of query: 646
effective length of database: 6,017,328
effective search space: 3887193888
effective search space used: 3887193888
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 63 (28.9 bits)
Medicago: description of AC144721.3


