

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
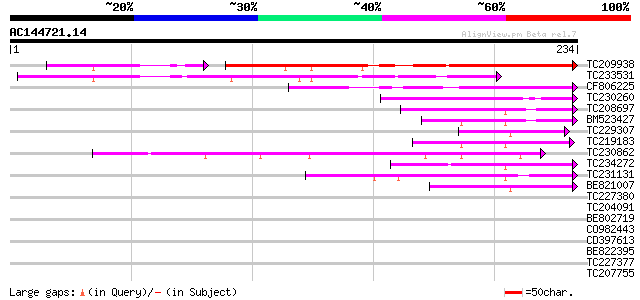
Query= AC144721.14 - phase: 0
(234 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC209938 similar to UP|Q84U29 (Q84U29) Zinc finger-like protein ... 145 4e-42
TC233531 similar to UP|Q84U29 (Q84U29) Zinc finger-like protein ... 115 1e-26
CF806225 103 9e-23
TC230260 49 2e-06
TC208697 43 1e-04
BM523427 43 1e-04
TC229307 similar to UP|Q6K9X4 (Q6K9X4) Ubiquitin-protein ligase-... 43 1e-04
TC219183 42 2e-04
TC230862 42 2e-04
TC234272 42 3e-04
TC231131 similar to GB|BAB71852.1|16902294|AB062739 kinesin-rela... 41 5e-04
BE821007 40 7e-04
TC227380 weakly similar to UP|Q7EYM0 (Q7EYM0) Zinc finger protei... 40 0.001
TC204091 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1... 40 0.001
BE802719 40 0.001
CO982443 39 0.002
CD397613 39 0.002
BE822395 39 0.002
TC227377 similar to UP|Q7EYM0 (Q7EYM0) Zinc finger protein famil... 39 0.002
TC207755 similar to UP|Q6K9X4 (Q6K9X4) Ubiquitin-protein ligase-... 39 0.002
>TC209938 similar to UP|Q84U29 (Q84U29) Zinc finger-like protein (Fragment),
partial (73%)
Length = 1104
Score = 145 bits (367), Expect(2) = 4e-42
Identities = 86/159 (54%), Positives = 100/159 (62%), Gaps = 14/159 (8%)
Frame = +3
Query: 90 RRTSDESTHQQQQNEDIPTNSDP--------SEFLPGGEFSD----ETAQVSMSLMDLLE 137
R SDE T Q +S+P + GG+ SD A V +SLMDLLE
Sbjct: 303 RHDSDELTQQNDVVAGEANDSEPIARFGSPVEQVAAGGDSSDGENNHEAVVGVSLMDLLE 482
Query: 138 ESEIELE--RISDVVDEGDDVEENNEKREEEEEREEEGEGEGSNVKVEHNCCVCMVRDKG 195
E+E E++ R+SD DDV E E+RE EGE E V VE+NCCVCMVR KG
Sbjct: 483 EAEQEMDLYRVSD----DDDVAEY-------EKREGEGEEEKGGV-VEYNCCVCMVRHKG 626
Query: 196 AAFIPCGHTFCRMCSRELWVSRGNCPLCNHFILEVLDIF 234
AAFIPCGHTFCRMCSRE+WVSRGNCPLCN+ ILE+LDIF
Sbjct: 627 AAFIPCGHTFCRMCSREIWVSRGNCPLCNNLILEILDIF 743
Score = 42.7 bits (99), Expect(2) = 4e-42
Identities = 30/74 (40%), Positives = 39/74 (52%), Gaps = 7/74 (9%)
Frame = +2
Query: 16 TSNTRSSLASLIQNDAVL-------SNQKNTNQTLLDIIRENEPNNVVVNVSVTNSSSSS 68
+S +R SLASL+ +D V S + + ++TL DIIRE +PNN SS
Sbjct: 17 SSGSRDSLASLMMDDVVFKKMETEASGELSRSRTLQDIIREEKPNN------------SS 160
Query: 69 NNNVVKDRKSWKAF 82
N DR SWKAF
Sbjct: 161N---ATDRNSWKAF 193
>TC233531 similar to UP|Q84U29 (Q84U29) Zinc finger-like protein (Fragment),
partial (68%)
Length = 926
Score = 115 bits (289), Expect = 1e-26
Identities = 96/258 (37%), Positives = 124/258 (47%), Gaps = 58/258 (22%)
Frame = +2
Query: 4 RQRITLYDQMTSTSNTRSSLASLIQNDAVL-----------------SNQKNTNQTLLDI 46
R+RIT YDQM + +++R SLASL+ D V S + + ++TL DI
Sbjct: 173 RRRITFYDQMKAGNSSRYSLASLMLGDVVFKKVEAEEAEEAEAEAEASCELSRSRTLQDI 352
Query: 47 IRENEPNNVVVNVSVTNSSSSSNNNVVKDRKSWKAFKELLLFKR---------------- 90
IRE +P+N SSNN KDR SW+AF L KR
Sbjct: 353 IREEKPSN------------SSNNT--KDRNSWRAFTAKLRLKRAIGSAWSSTTTITNHS 490
Query: 91 ----------RTSDESTHQQQQNEDIPTNSDPSEFLPG-----------GEFSD----ET 125
T +S QQN+ + ++ S+ + G SD
Sbjct: 491 NDDRNLPNIPETRHDSDELTQQNDIVVEGANDSDPIARFGSPVEHVEAVGNSSDGENNHE 670
Query: 126 AQVSMSLMDLLEESEIELERISDVVDEGDDVEENNEKREEEEEREEEGEGEGSNVKVEHN 185
A V +SLMDLLEE+E E++ + V D+ DDV E EKREEE+ EE G VE+N
Sbjct: 671 AVVGVSLMDLLEEAEQEMD-LYRVSDDDDDVAEY-EKREEEKGGVEEEIGS----VVEYN 832
Query: 186 CCVCMVRDKGAAFIPCGH 203
CCVCMVR KGAAFIPCGH
Sbjct: 833 CCVCMVRHKGAAFIPCGH 886
Score = 31.6 bits (70), Expect = 0.32
Identities = 14/31 (45%), Positives = 16/31 (51%)
Frame = +3
Query: 186 CCVCMVRDKGAAFIPCGHTFCRMCSRELWVS 216
C C + K G FCR CSRE+WVS
Sbjct: 840 CAWCGI--KARRLFRAGTXFCRTCSREIWVS 926
>CF806225
Length = 612
Score = 103 bits (256), Expect = 9e-23
Identities = 59/119 (49%), Positives = 70/119 (58%)
Frame = +1
Query: 116 LPGGEFSDETAQVSMSLMDLLEESEIELERISDVVDEGDDVEENNEKREEEEEREEEGEG 175
+P E A +SLM LLEE+E G D E+E+E+EE G
Sbjct: 268 VPQEEGGGVRAPWRVSLMRLLEETE------------GGDAAT----AEKEKEKEENVLG 399
Query: 176 EGSNVKVEHNCCVCMVRDKGAAFIPCGHTFCRMCSRELWVSRGNCPLCNHFILEVLDIF 234
S CCVCM R KGAA IPCGHTFCR+CSRELW++RG+CPLCN ILE+LDIF
Sbjct: 400 NDSV------CCVCMGRKKGAALIPCGHTFCRVCSRELWLNRGSCPLCNRSILEILDIF 558
>TC230260
Length = 783
Score = 49.3 bits (116), Expect = 2e-06
Identities = 26/81 (32%), Positives = 39/81 (48%)
Frame = +3
Query: 154 DDVEENNEKREEEEEREEEGEGEGSNVKVEHNCCVCMVRDKGAAFIPCGHTFCRMCSREL 213
++++E+ E+ER E+ E K C VC+ + +PCGH CR CS
Sbjct: 381 ENLKESQAALVLEQERVEKATKEADTAKAAWVCRVCLSSEVDITIVPCGHVLCRRCSSA- 557
Query: 214 WVSRGNCPLCNHFILEVLDIF 234
VSR CP C + + + IF
Sbjct: 558 -VSR--CPFCRLQVTKAIRIF 611
>TC208697
Length = 769
Score = 43.1 bits (100), Expect = 1e-04
Identities = 22/74 (29%), Positives = 36/74 (47%), Gaps = 1/74 (1%)
Frame = +3
Query: 162 KREEEEEREEEGEGEGSNVKVEHNCCVCMVRDKGAAFIPCGH-TFCRMCSRELWVSRGNC 220
+ + E+ + +G + + C +C+ ++ A F+PCGH C CS L NC
Sbjct: 147 QNNDVEKADSLSDGAKKDRLMPDLCVICLEQEYNAVFVPCGHMCCCTACSSHL----TNC 314
Query: 221 PLCNHFILEVLDIF 234
PLC I +V+ F
Sbjct: 315 PLCRRQIEKVVKTF 356
>BM523427
Length = 419
Score = 43.1 bits (100), Expect = 1e-04
Identities = 24/69 (34%), Positives = 34/69 (48%), Gaps = 5/69 (7%)
Frame = +1
Query: 171 EEGEGEGSNVKVEHN----CCVCMVRDKGAAFIPCGH-TFCRMCSRELWVSRGNCPLCNH 225
E+ +G VK + C +C+ ++ A F+PCGH C CS L NCPLC
Sbjct: 16 EKADGLSDGVKKDRLMPDLCVICLEQEYNAVFVPCGHMCCCTTCSSHL----TNCPLCRR 183
Query: 226 FILEVLDIF 234
I +V+ F
Sbjct: 184 QIEKVVKTF 210
>TC229307 similar to UP|Q6K9X4 (Q6K9X4) Ubiquitin-protein ligase-like,
partial (15%)
Length = 983
Score = 42.7 bits (99), Expect = 1e-04
Identities = 18/47 (38%), Positives = 26/47 (55%), Gaps = 1/47 (2%)
Frame = +1
Query: 186 CCVCMVRDKGAAFIPCGHTF-CRMCSRELWVSRGNCPLCNHFILEVL 231
CC+C + + CGH C C+ EL SR NCP+C ++EV+
Sbjct: 451 CCICCESNIDSLLYRCGHLCTCSKCANELLQSRRNCPMCQAPVVEVI 591
>TC219183
Length = 742
Score = 42.4 bits (98), Expect = 2e-04
Identities = 23/73 (31%), Positives = 32/73 (43%), Gaps = 6/73 (8%)
Frame = +1
Query: 167 EEREEEGEGEGSNVKVEHN-----CCVCMVRDKGAAFIPCGH-TFCRMCSRELWVSRGNC 220
E RE G G S + N C +CM K A +PC H C C+ + C
Sbjct: 103 ELRELYGIGNSSAADFDDNDPGKECVICMTEPKDTAVLPCRHMCMCSECANAHRLQSNKC 282
Query: 221 PLCNHFILEVLDI 233
P+C I E+++I
Sbjct: 283 PICRQSIEELIEI 321
>TC230862
Length = 868
Score = 42.0 bits (97), Expect = 2e-04
Identities = 44/218 (20%), Positives = 91/218 (41%), Gaps = 31/218 (14%)
Frame = +1
Query: 35 NQKNTNQTLLDIIRENEPNNVVVNVSVTNSSSSSNNNVVKDRKSW-KAFKELLLFKRRTS 93
+ + NQ LL+ + E + N+ + VS + + ++N ++ +++ K +++ +
Sbjct: 49 DMQTQNQNLLNQVIERDDYNIKL-VSDSVKTKQAHNTLMSQKQALAKQLQQINTSIENSK 225
Query: 94 DESTHQQQQ-----NEDIPTNSDPSEFLPGGEFS-------DETAQVSMSLMDLLEESEI 141
TH ++Q ++ I N + EF+ ++ ++ S + E+
Sbjct: 226 TRITHSEEQMKAILSDAIKCNQEEKHLAVTLEFAKWELADAEKELKLLKSAVSSSEKEYD 405
Query: 142 ELERISDVVDEGDDVEENNEKREEEEERE--------EEGEGEGSNVKVEHN-------- 185
++++ ++ ++ + E + K+ EEE RE GE + K+E
Sbjct: 406 QIQKDTEAIEMELESERSLRKKLEEELRELNCKIDELTSETGETTIQKLEKEIRICKNMI 585
Query: 186 -CCVCMVRDKGAAFIPCGHTFCRMC-SRELWVSRGNCP 221
C VC R K + C H FC C R L + CP
Sbjct: 586 KCTVCTDRPKEVVIVKCYHLFCNPCIQRNLELRHRKCP 699
>TC234272
Length = 554
Score = 41.6 bits (96), Expect = 3e-04
Identities = 21/78 (26%), Positives = 38/78 (47%), Gaps = 1/78 (1%)
Frame = +3
Query: 158 ENNEKREEEEEREEEGEGEGSNVKVEHNCCVCMVRDKGAAFIPCGH-TFCRMCSRELWVS 216
+ + K+E + ++EE + K + NCC+C + CGH C C+ EL +
Sbjct: 78 QQSMKQEVQTVKKEEKKSNNRTPK-KGNCCICYEMKVDSVLYRCGHMCTCLKCANELQWN 254
Query: 217 RGNCPLCNHFILEVLDIF 234
G C +C I +V+ ++
Sbjct: 255 SGKCSICRAKIEDVVRVY 308
>TC231131 similar to GB|BAB71852.1|16902294|AB062739 kinesin-related protein
{Arabidopsis thaliana;} , partial (15%)
Length = 725
Score = 40.8 bits (94), Expect = 5e-04
Identities = 34/118 (28%), Positives = 51/118 (42%), Gaps = 6/118 (5%)
Frame = +3
Query: 123 DETAQVSMSLMDLLEESEIELERISDV-VDEGDDVEEN----NEKREEEEEREEEGEGEG 177
DE A + + +E I E+I DV + E + E+ K +E RE+E + G
Sbjct: 282 DEEAHTNDLKTNDIESDIIPKEQILDVSIPENEITNEDPLVVRLKARMKEMREKEFKHLG 461
Query: 178 SNVKVEHNCCVCMVRDKGAAFIPCGH-TFCRMCSRELWVSRGNCPLCNHFILEVLDIF 234
+ H C VC A +PC H C+ CS ++ CP+C I + L F
Sbjct: 462 NGDANSHVCKVCFQSSTAAILLPCRHFCLCKSCS----LACSECPICRTNISDRLFAF 623
>BE821007
Length = 702
Score = 40.4 bits (93), Expect = 7e-04
Identities = 20/62 (32%), Positives = 30/62 (48%), Gaps = 1/62 (1%)
Frame = -1
Query: 174 EGEGSNVKVEHNCCVCMVRDKGAAFIPCGHTF-CRMCSRELWVSRGNCPLCNHFILEVLD 232
EGE S +C +C+ A IPCGH C C E+ + CP+C I +V+
Sbjct: 441 EGEKSAGGSSSSCVICLDAPAEGACIPCGHVAGCMSCLNEVKSKKWGCPVCRAKIDQVIK 262
Query: 233 IF 234
++
Sbjct: 261 LY 256
>TC227380 weakly similar to UP|Q7EYM0 (Q7EYM0) Zinc finger protein
family-like, partial (70%)
Length = 770
Score = 39.7 bits (91), Expect = 0.001
Identities = 20/90 (22%), Positives = 38/90 (42%), Gaps = 2/90 (2%)
Frame = +2
Query: 143 LERISDVVDEGDDVEENNEKREEEEEREEEGEGEGSNVKVEHN--CCVCMVRDKGAAFIP 200
L ++ + V + +D ++ E +R++E + S++ +E C +CM +
Sbjct: 110 LVQLQESVADTEDKKQKAVCMERYRKRDDEEHRQPSDIDIEREEECGICMEMNSKIVLPD 289
Query: 201 CGHTFCRMCSRELWVSRGNCPLCNHFILEV 230
C H C C E +CP C + V
Sbjct: 290 CNHVMCLTCYHEWRTRSQSCPFCRDSLKRV 379
>TC204091 similar to UP|Q8LLE6 (Q8LLE6) RNA-binding protein AKIP1, partial
(70%)
Length = 2075
Score = 39.7 bits (91), Expect = 0.001
Identities = 17/35 (48%), Positives = 25/35 (70%)
Frame = +2
Query: 138 ESEIELERISDVVDEGDDVEENNEKREEEEEREEE 172
E E+E E + + V+E ++ EE +E+ EEEEE EEE
Sbjct: 137 EEEVEEEEVEEEVEEEEEEEEEDEEEEEEEEEEEE 241
Score = 37.4 bits (85), Expect = 0.006
Identities = 22/45 (48%), Positives = 28/45 (61%)
Frame = +2
Query: 137 EESEIELERISDVVDEGDDVEENNEKREEEEEREEEGEGEGSNVK 181
EE E E+E +V +E ++ EE E+ EEEEE EEE E E VK
Sbjct: 128 EEVEEEVEE-EEVEEEVEEEEEEEEEDEEEEEEEEEEEEENDVVK 259
Score = 34.3 bits (77), Expect = 0.050
Identities = 18/44 (40%), Positives = 28/44 (62%)
Frame = +2
Query: 142 ELERISDVVDEGDDVEENNEKREEEEEREEEGEGEGSNVKVEHN 185
E E + + V+E ++VEE E+ EEEEE +EE E E + E++
Sbjct: 122 EYEEVEEEVEE-EEVEEEVEEEEEEEEEDEEEEEEEEEEEEEND 250
Score = 32.3 bits (72), Expect = 0.19
Identities = 15/35 (42%), Positives = 22/35 (62%)
Frame = +2
Query: 136 LEESEIELERISDVVDEGDDVEENNEKREEEEERE 170
+EE E+E E + +E +D EE E+ EEEEE +
Sbjct: 146 VEEEEVEEEVEEEEEEEEEDEEEEEEEEEEEEEND 250
>BE802719
Length = 385
Score = 39.7 bits (91), Expect = 0.001
Identities = 22/70 (31%), Positives = 31/70 (43%), Gaps = 6/70 (8%)
Frame = +2
Query: 167 EEREEEGEGEGSNVKVEHN-----CCVCMVRDKGAAFIPCGH-TFCRMCSRELWVSRGNC 220
E RE G G + + N C +CM K A +PC H C C++ L + C
Sbjct: 176 ELRELYGIGSSTVADFDDNDPGKECVICMTEPKDTAVLPCRHMCMCGDCAKALRLQSNKC 355
Query: 221 PLCNHFILEV 230
P+C I E+
Sbjct: 356 PICRQPIEEL 385
>CO982443
Length = 901
Score = 39.3 bits (90), Expect = 0.002
Identities = 16/47 (34%), Positives = 25/47 (53%), Gaps = 1/47 (2%)
Frame = -2
Query: 186 CCVCMVRDKGAAFIPCGH-TFCRMCSRELWVSRGNCPLCNHFILEVL 231
CC+C + + CGH C C+ +L SR CP+C ++EV+
Sbjct: 600 CCICCESNIDSLLYRCGHMCTCSKCANDLLQSRRKCPMCQAPVVEVI 460
>CD397613
Length = 469
Score = 39.3 bits (90), Expect = 0.002
Identities = 16/47 (34%), Positives = 25/47 (53%), Gaps = 1/47 (2%)
Frame = -3
Query: 186 CCVCMVRDKGAAFIPCGH-TFCRMCSRELWVSRGNCPLCNHFILEVL 231
CC+C + + CGH C C+ +L SR CP+C ++EV+
Sbjct: 398 CCICCESNIDSLLYRCGHMCTCSKCANDLLQSRRKCPMCQAPVVEVI 258
>BE822395
Length = 637
Score = 38.9 bits (89), Expect = 0.002
Identities = 21/72 (29%), Positives = 34/72 (47%), Gaps = 1/72 (1%)
Frame = -1
Query: 164 EEEEEREEEGEGEGSNVKVEHNCCVCMVRDKGAAFIPCGHTF-CRMCSRELWVSRGNCPL 222
E+ + E+ G GS+ C +C+ A IPCGH C C E+ + CP+
Sbjct: 487 EKLPKGEKNAGGSGSS------CVICLDAPAEGACIPCGHVAGCMSCLNEVKSKKWGCPV 326
Query: 223 CNHFILEVLDIF 234
C I +V+ ++
Sbjct: 325 CRAKIDQVIKLY 290
>TC227377 similar to UP|Q7EYM0 (Q7EYM0) Zinc finger protein family-like,
partial (98%)
Length = 1244
Score = 38.9 bits (89), Expect = 0.002
Identities = 21/81 (25%), Positives = 33/81 (39%), Gaps = 5/81 (6%)
Frame = +3
Query: 155 DVEENNEKR---EEEEEREEEGEGEGSNVKVEHN--CCVCMVRDKGAAFIPCGHTFCRMC 209
D E+ +K E R++E + S++ +E C +CM + C H C C
Sbjct: 507 DTEDKKQKAVCMERYRRRDDEEYRQSSDIDIEREDECGICMDMNSKIVLPNCNHAMCLKC 686
Query: 210 SRELWVSRGNCPLCNHFILEV 230
RE +CP C + V
Sbjct: 687 YREWRTISQSCPFCRDSLKRV 749
>TC207755 similar to UP|Q6K9X4 (Q6K9X4) Ubiquitin-protein ligase-like, partial
(23%)
Length = 1438
Score = 38.9 bits (89), Expect = 0.002
Identities = 19/47 (40%), Positives = 23/47 (48%), Gaps = 1/47 (2%)
Frame = +3
Query: 186 CCVCMVRDKGAAFIPCGHTF-CRMCSRELWVSRGNCPLCNHFILEVL 231
CCVC + CGH C C+ EL G CPLC I+EV+
Sbjct: 1062 CCVCCDNHIDSLLYRCGHMCTCSKCANELIRGGGKCPLCRAPIVEVV 1202
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.311 0.128 0.360
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,530,246
Number of Sequences: 63676
Number of extensions: 158598
Number of successful extensions: 2527
Number of sequences better than 10.0: 322
Number of HSP's better than 10.0 without gapping: 1984
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2254
length of query: 234
length of database: 12,639,632
effective HSP length: 94
effective length of query: 140
effective length of database: 6,654,088
effective search space: 931572320
effective search space used: 931572320
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 57 (26.6 bits)
Medicago: description of AC144721.14


