

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144656.5 + phase: 0 /pseudo
(2301 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
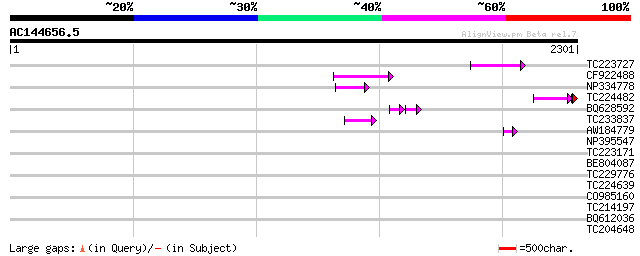
Sequences producing significant alignments: (bits) Value
TC223727 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 99 3e-20
CF922488 92 2e-18
NP334778 reverse transcriptase [Glycine max] 75 3e-13
TC224482 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 46 3e-07
BQ628592 49 1e-06
TC233837 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 50 1e-05
AW184779 48 4e-05
NP395547 reverse transcriptase [Glycine max] 42 0.002
TC223171 similar to UP|Q8RW43 (Q8RW43) TCP1 protein, partial (9%) 36 0.22
BE804087 35 0.29
TC229776 similar to GB|AAS10022.1|41619094|AY519552 MYB transcri... 35 0.50
TC224639 homologue to UP|Q43682 (Q43682) Extensin-like protein (... 33 1.9
CO985160 32 2.5
TC214197 similar to UP|Q6UNT2 (Q6UNT2) 60S ribosomal protein L5,... 31 5.5
BQ612036 similar to GP|7291176|gb|A CG15295-PA {Drosophila melan... 31 5.5
TC204648 homologue to UP|Q9M5J4 (Q9M5J4) Beta galactosidase, com... 31 7.2
>TC223727 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (9%)
Length = 843
Score = 98.6 bits (244), Expect = 3e-20
Identities = 76/225 (33%), Positives = 118/225 (51%)
Frame = +3
Query: 1870 SRETRSCIRH*RSC*WQAMVPRYQMLPSKARVSTWSIQ*RQEDSKKIVWQFLPERGHFV* 1929
S +TR+ + R A+V R+Q + K R+ + ED +K+ +FL ER H V*
Sbjct: 84 SWQTRALLPSRRGTGR*ALVFRHQAICHKQRIPARDCRQ**EDIEKVGGRFLHERKHTV* 263
Query: 1930 KKLRHGVAQMC*QT*GRLIDA*NTRRILRNSF*RTYNG*EDIESRLLLDDNGS*LLQACQ 1989
+K RH + +C G+ D + ++ N+ R G ED +SRLLL +G *LL C+
Sbjct: 264 EKPRHETSAVCGCQGGKSHDRGSP*GLVWNARQRACYGQEDPKSRLLLAYHGK*LLCPCE 443
Query: 1990 EVL*MSNLCQ*HSCTANCTQCPFLPVAILYVGH*HDWKN*TQSFKWAPLHSTGY*LFHQV 2049
E+ MS++ +* C ++C LP+A L+VG+ +* Q +W+ LH LFHQV
Sbjct: 444 EMPQMSSVRR*CQCPTASSECHVLPLAFLHVGNRCHRGH*AQGLEWSSLHPRSDRLFHQV 623
Query: 2050 G*SCILC*CDQAGGSQVYQESYYLPLWCP*PDHYR*WDESEQQDD 2094
G L + G QV+ E +LP+W D++ + + QDD
Sbjct: 624 GRGSFLYRRHEGCGGQVH*ERDHLPIWFAKEDYHGQRHQPK*QDD 758
>CF922488
Length = 741
Score = 92.4 bits (228), Expect = 2e-18
Identities = 74/241 (30%), Positives = 115/241 (47%)
Frame = +2
Query: 1315 EEGWQSQNVC*LP*P*QS*SKR*FPITSY*CTG*QHCKIQSVLPHGWFLRLQSDQDGARR 1374
E GW+ NV L S SK * T+Y + H + +L HGW RL D+D R
Sbjct: 8 ERGWEGVNVRGLSRSKLSQSKG*VSFTAYQRSRG*HDQFFPILLHGWIFRL*PDKDSTRG 187
Query: 1375 QREDVFHHTLGHFLLQSNAVRFDQCRSYLSKGYDYYLSRHDTQRD*GLCG*HDRQVNH*R 1434
+D FH+++G+ LL V ++C + G+ + HD Q D GL G*HDR++ +
Sbjct: 188 YGKDNFHYSMGNLLL*GYVVWAEECWGNIPAGHGGIILGHDAQGDRGLHG*HDREIKNGG 367
Query: 1435 TTC*VFAKDVPTAKKVQTSSESQQVYFWC*IRKSSGLHC*SERY*SRSRQGQSHKRNASS 1494
T FAK V +Q +ES++VY I+K + L+ * ER QG+S + +
Sbjct: 368 GTPCQFAKVV*ATT*IQVETESRKVYV*GKIQKVARLY**LERNRGGFEQGESDP*DGQA 547
Query: 1495 ANRKTSKGVSRTTELHLPFHFSYDYNMWADIQTPPQGSRVRLDRRLPESL**HQSILTRT 1554
R+ S ELH H + ++ A Q S ++ L S+* + ++ ++
Sbjct: 548 TYREASPRFLGEVELHRKIHIIVNCHLRASFHIVVQESVCQMGP*LLSSI*EN*TVFNKS 727
Query: 1555 S 1555
S
Sbjct: 728 S 730
>NP334778 reverse transcriptase [Glycine max]
Length = 431
Score = 75.5 bits (184), Expect = 3e-13
Identities = 50/139 (35%), Positives = 75/139 (52%)
Frame = +2
Query: 1321 QNVC*LP*P*QS*SKR*FPITSY*CTG*QHCKIQSVLPHGWFLRLQSDQDGARRQREDVF 1380
++VC L S SK *F T++ + H + +L HGW L SD+DG R +D F
Sbjct: 2 EDVCGLSRSKPSQSKG*FSFTAHRHSHG*HGQFCPILFHGWIFGL*SDKDGTRGYGKDNF 181
Query: 1381 HHTLGHFLLQSNAVRFDQCRSYLSKGYDYYLSRHDTQRD*GLCG*HDRQVNH*RTTC*VF 1440
H+++G+ LL + V ++ L G+ + HD Q D GLCG*+DR++ + T F
Sbjct: 182 HYSMGNLLL*GDVVWAEEFWGNLPSGHGGIIPGHDAQGDRGLCG*NDREIKNGGGTPSQF 361
Query: 1441 AKDVPTAKKVQTSSESQQV 1459
AK V +Q +ESQ+V
Sbjct: 362 AKFVWATT*IQVETESQEV 418
>TC224482 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (6%)
Length = 669
Score = 45.8 bits (107), Expect(2) = 3e-07
Identities = 43/160 (26%), Positives = 68/160 (41%)
Frame = +3
Query: 2125 KYKEDHPEDGGDL*RLA*DASLCITWISYFCTHINRGNPFLFGIWHGGSTSHRS*DSFNA 2184
+Y++D+P+D + LA DA + +T + F ++N GN L GIW GG +
Sbjct: 15 EYQKDYPKDDRVIQGLARDAPIRVTRLPDFSANVNWGNAVLIGIWDGGCVTV*GRSPVIK 194
Query: 2185 SFNGG*IIRG*VGSK*V*PIESDRRKAHVCSLSWLAVSKEDEAGF*QESSSP*IQRR*SC 2244
I VGS +* + A SW V ++E QE + C
Sbjct: 195 DLGRIRIKGIRVGSNAL*SAQPH*G*ALNGHESWALVPAKNEECIRQEGMLAQVP*GRPC 374
Query: 2245 AQKDLDFPTRL*RQVDA*L*RPVCCQESLLRRCHDSSNHG 2284
A++++ R++ L R CC+E RR + HG
Sbjct: 375 AKENVPCCQGPSREMGPELRRAFCCEEGFFRRSPGAYQHG 494
Score = 28.9 bits (63), Expect(2) = 3e-07
Identities = 11/21 (52%), Positives = 16/21 (75%)
Frame = +1
Query: 2281 SNHGCEELPRPVNTDAVKKYF 2301
+N EELP P+N+D VK+Y+
Sbjct: 484 TNMDGEELPSPMNSDVVKRYY 546
>BQ628592
Length = 423
Score = 48.9 bits (115), Expect(2) = 1e-06
Identities = 26/63 (41%), Positives = 37/63 (58%)
Frame = -3
Query: 1540 LPESL**HQSILTRTSDSHPSS*RKTTNHVPDCTRRFHGLCARTTRRNWKKGACHLLFEQ 1599
LP L +Q+ + +H + RKT + V D RR +G+ + RR W+KG HLLF+Q
Sbjct: 388 LPRGLRKNQTEPRESLGAHATCNRKTFSPVHDYVRRVYGMRVGSARRLWEKGTSHLLFKQ 209
Query: 1600 EVY 1602
EVY
Sbjct: 208 EVY 200
Score = 23.9 bits (50), Expect(2) = 1e-06
Identities = 18/61 (29%), Positives = 31/61 (50%)
Frame = -2
Query: 1608 LFHAREDLLRTSLGRQTPPSLYD*SYYLVDI*DGSDQVHL*ETRFDRKNCPVADVIIRI* 1667
L + +D+L +G + +++ SY++ +GS ++HL*E N VA I I
Sbjct: 185 LLNVGKDVLYLGMGVTSS*AVHAQSYHVAYFQNGSCEIHL*EAGPHGTNR*VAGTTI*IQ 6
Query: 1668 Y 1668
Y
Sbjct: 5 Y 3
>TC233837 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (6%)
Length = 402
Score = 50.1 bits (118), Expect = 1e-05
Identities = 39/130 (30%), Positives = 66/130 (50%)
Frame = +1
Query: 1357 LPHGWFLRLQSDQDGARRQREDVFHHTLGHFLLQSNAVRFDQCRSYLSKGYDYYLSRHDT 1416
L +GWFL + S+ DG R RED F H +G L S+ +R ++ LS + + +D
Sbjct: 4 LIYGWFLGV*SNIDGT*RCREDHFRHPMGDIQL*SDGLRAEKYWGNLSACHGGVVP*YDA 183
Query: 1417 QRD*GLCG*HDRQVNH*RTTC*VFAKDVPTAKKVQTSSESQQVYFWC*IRKSSGLHC*SE 1476
+ GL *HD QV+ * T V ++ ++ Q++ W + +++ ++ SE
Sbjct: 184 *GNRGLRR*HDCQVSD*DRTPCQSV*AVWKVAEIPIKTKPNQMHIWGEVGEAARIYRKSE 363
Query: 1477 RY*SRSRQGQ 1486
R RSR+ +
Sbjct: 364 RDRDRSRESE 393
>AW184779
Length = 432
Score = 48.1 bits (113), Expect = 4e-05
Identities = 26/60 (43%), Positives = 34/60 (56%), Gaps = 5/60 (8%)
Frame = +2
Query: 2005 ANCTQCPFLPVAIL-----YVGH*HDWKN*TQSFKWAPLHSTGY*LFHQVG*SCILC*CD 2059
A+ P +P+ +L YVGH D +* Q FKW LH + *L HQ+G S +C*CD
Sbjct: 233 ADNVNAPPIPLNVLAAPWPYVGHRRDRSH*AQGFKWTSLHFSRN*LLHQMGRSSFVC*CD 412
>NP395547 reverse transcriptase [Glycine max]
Length = 762
Score = 42.4 bits (98), Expect = 0.002
Identities = 38/107 (35%), Positives = 51/107 (47%)
Frame = +3
Query: 1319 QSQNVC*LP*P*QS*SKR*FPITSY*CTG*QHCKIQSVLPHGWFLRLQSDQDGARRQRED 1378
Q +NV *L S KR P + + *+ CK + G LRLQSD G+ R++
Sbjct: 156 QMENVY*L*EAQ*SHKKRPLPTSLHGSNA*ETCKAILLPFLGRILRLQSDCSGSSGSRKN 335
Query: 1379 VFHHTLGHFLLQSNAVRFDQCRSYLSKGYDYYLSRHDTQRD*GLCG* 1425
F+ + F L +AVRF C Y S+ YD H + * L G*
Sbjct: 336 SFYMSFQCFCLSPHAVRFM*CLYYFSEMYDGNF**HGREMY*SLYG* 476
>TC223171 similar to UP|Q8RW43 (Q8RW43) TCP1 protein, partial (9%)
Length = 411
Score = 35.8 bits (81), Expect = 0.22
Identities = 24/74 (32%), Positives = 32/74 (42%), Gaps = 12/74 (16%)
Frame = +3
Query: 388 QSTNISLPSHLLPMLFNHQIVSLDSHNILNKIL------------NNNIPNQHTSNAHIN 435
Q +SL S P+L +H +HN +L NNN N H +N H N
Sbjct: 198 QDLRLSLQSLQDPILLHH------NHNNNEHVLFAGTAFDNMVAWNNNSNNNHNNNNHNN 359
Query: 436 NNPINNNSLDHKKC 449
NN NNN+ + C
Sbjct: 360 NNNNNNNTASNDTC 401
>BE804087
Length = 160
Score = 35.4 bits (80), Expect = 0.29
Identities = 18/47 (38%), Positives = 25/47 (52%)
Frame = +1
Query: 2125 KYKEDHPEDGGDL*RLA*DASLCITWISYFCTHINRGNPFLFGIWHG 2171
+Y E + ED + RLA*D C ++S T+ GN L G+W G
Sbjct: 16 EY*EKYLEDDSVVQRLA*DVFFCPAYVSNLGTNFYWGNTLLLGLWEG 156
>TC229776 similar to GB|AAS10022.1|41619094|AY519552 MYB transcription factor
{Arabidopsis thaliana;} , partial (8%)
Length = 841
Score = 34.7 bits (78), Expect = 0.50
Identities = 20/62 (32%), Positives = 34/62 (54%), Gaps = 1/62 (1%)
Frame = +1
Query: 389 STNISLPSHLLPMLFNHQIVSLDSHNILNKILNNNIPNQHTSNAHINNNP-INNNSLDHK 447
+ N ++ S +LP +F Q V L S+N I ++ + N TSN + +N+ NN S+ +
Sbjct: 43 NNNSTITSSMLPSIFPTQHVKLQSNNNNPSISSDGVQNWETSNLNNSNSTNKNNGSMQLQ 222
Query: 448 KC 449
C
Sbjct: 223 SC 228
>TC224639 homologue to UP|Q43682 (Q43682) Extensin-like protein (Fragment),
partial (48%)
Length = 702
Score = 32.7 bits (73), Expect = 1.9
Identities = 33/102 (32%), Positives = 45/102 (43%), Gaps = 4/102 (3%)
Frame = +2
Query: 20 HKPRLRL*LKLRLMPK-PKHLLHLLSELRSKHLP-LLSLNGQYVPTLRHTLLHNVPRLGS 77
H P + + L L +P P H+ L HLP L+S N P LRH H++ S
Sbjct: 17 HHPPMSISLHLHRLPLLPHHMFTNLHPHHHLHLPHLISTN----PLLRHLPRHHLHMFIS 184
Query: 78 HLLLLAKYSVPLLVKLR--CLLLNMQLMFLRLLRQFLKLL*H 117
HL L + P+ R LLL++ F+ L L L H
Sbjct: 185 HLPHLPLHHHPMSTSHRHLHLLLHLPHTFINLHLHLLHRLLH 310
>CO985160
Length = 777
Score = 32.3 bits (72), Expect = 2.5
Identities = 37/101 (36%), Positives = 53/101 (51%), Gaps = 1/101 (0%)
Frame = +3
Query: 30 LRLMPKPKHLLHLLSELRSKHLPLLSLNGQYVPTLR-HTLLHNVPRLGSHLLLLAKYSVP 88
L L+ + + LL LL +LR L LL L + L LL + +L LLLL +
Sbjct: 477 LLLLLQLRKLLLLLLQLRKLLLLLLQLRKLLLLLLELRKLLLLLLQLRKLLLLLLELRKL 656
Query: 89 LLVKLRCLLLNMQLMFLRLLRQFLKLL*HTQHLRFMLSRKI 129
LL+ +R LLL +QL + LL L LL HL +L+R++
Sbjct: 657 LLLHVR*LLLLLQLRKMLLL--VLHLLMLLLHLLLLLTRQL 773
>TC214197 similar to UP|Q6UNT2 (Q6UNT2) 60S ribosomal protein L5, partial
(67%)
Length = 904
Score = 31.2 bits (69), Expect = 5.5
Identities = 45/167 (26%), Positives = 72/167 (42%), Gaps = 13/167 (7%)
Frame = +1
Query: 45 ELRSKHLPLLSLNGQYVPTLRHTLLHNVPRLGSH-------LLLLAKYSVPLLVKLRCLL 97
E SKHL + + + L T+ N+ R G H +LL + + LV LR L
Sbjct: 37 EESSKHLKWMRSTKEMLRLLERTIQWNLLRAGGHSVLSLMLVLLGLQLATVSLVPLRELW 216
Query: 98 LNMQLMFLRLLRQFLKLL*HTQHLRFMLSRKITNPFSIQ------RV*RPMVKWRIYKRS 151
+ + + FL + R L L+ + R ++ R I N F + R R M I S
Sbjct: 217 MGVWI-FLIVTRGLLGLI---RRRRSLMLRCIANMFLVDMLLPI*RSWRKMNLRNISHTS 384
Query: 152 MMRCTVR*KLSAGKKGLEKLLMTSVWCRTSRYPTSSKSQISRSIKGT 198
+ +V *KL + + ++ SV SR + +S R+ +GT
Sbjct: 385 VNTSSVE*KLMVLRHCTRRFMLPSVLILLSR---NLRSSPRRNTRGT 516
>BQ612036 similar to GP|7291176|gb|A CG15295-PA {Drosophila melanogaster},
partial (5%)
Length = 288
Score = 31.2 bits (69), Expect = 5.5
Identities = 17/48 (35%), Positives = 24/48 (49%)
Frame = +1
Query: 417 NKILNNNIPNQHTSNAHINNNPINNNSLDHKKCKLTQSLLLTQSYSQD 464
N+I+NNN N +N + +NN NNN+ D+ Q YS D
Sbjct: 118 NEIINNN--NNGANNVNYHNNNFNNNNGDNNGFHENQDATTDTIYSSD 255
>TC204648 homologue to UP|Q9M5J4 (Q9M5J4) Beta galactosidase, complete
Length = 2817
Score = 30.8 bits (68), Expect = 7.2
Identities = 12/31 (38%), Positives = 19/31 (60%)
Frame = -1
Query: 1764 RLYKQHGRVRSMYHGYRRSHRSENQEPRHLW 1794
R Y ++ V+ HG+ ++HRSE + HLW
Sbjct: 351 RTYLRNSIVQEKVHGHSKTHRSE*HQGLHLW 259
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.358 0.158 0.577
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 112,759,325
Number of Sequences: 63676
Number of extensions: 1742791
Number of successful extensions: 20394
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 10985
Number of HSP's successfully gapped in prelim test: 754
Number of HSP's that attempted gapping in prelim test: 8435
Number of HSP's gapped (non-prelim): 13370
length of query: 2301
length of database: 12,639,632
effective HSP length: 112
effective length of query: 2189
effective length of database: 5,507,920
effective search space: 12056836880
effective search space used: 12056836880
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 67 (30.4 bits)
Medicago: description of AC144656.5


