

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144617.10 + phase: 0
(337 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
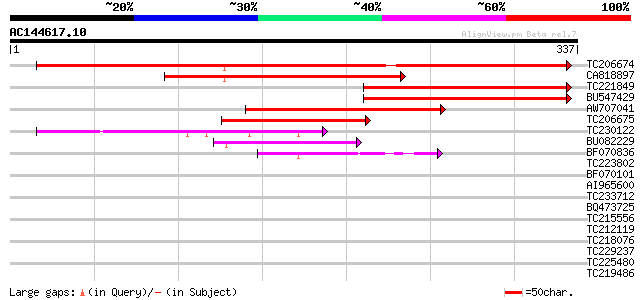
Sequences producing significant alignments: (bits) Value
TC206674 similar to UP|FMO3_HUMAN (P31513) Dimethylaniline monoo... 291 3e-79
CA818897 similar to GP|16555354|gb flavin-containing monooxygena... 179 2e-45
TC221849 163 1e-40
BU547429 158 4e-39
AW707041 149 1e-36
TC206675 84 9e-17
TC230122 61 8e-10
BU082229 similar to GP|9757850|dbj dimethylaniline monooxygenase... 48 7e-06
BF070836 similar to GP|21740855|emb OSJNBb0072N21.4 {Oryza sativ... 48 7e-06
TC223802 similar to UP|Q9FNA2 (Q9FNA2) Polyamine oxidase, partia... 38 0.007
BF070101 similar to GP|9759603|dbj| dimethylaniline monooxygenas... 37 0.013
AI965600 35 0.037
TC233712 similar to UP|O23024 (O23024) T1G11.14 protein (Flavin-... 34 0.11
BQ473725 33 0.24
TC215556 weakly similar to GB|AAP37810.1|30725576|BT008451 At3g4... 30 1.2
TC212119 weakly similar to UP|Q9LDR3 (Q9LDR3) Arabidopsis thalia... 30 1.2
TC218076 similar to UP|Q9MUE2 (Q9MUE2) S-adenosyl-L-methionine M... 30 1.5
TC229237 similar to UP|Q6TF29 (Q6TF29) Rapid alkalinization fact... 30 2.0
TC225480 UP|Q9XE94 (Q9XE94) Geranylgeranyl hydrogenase, complete 29 3.4
TC219486 28 4.5
>TC206674 similar to UP|FMO3_HUMAN (P31513) Dimethylaniline monooxygenase
[N-oxide-forming] 3 (Hepatic flavin-containing
monooxygenase 3) (FMO 3) (Dimethylaniline oxidase 3)
(FMO form 2) (FMO II) , partial (4%)
Length = 1470
Score = 291 bits (745), Expect = 3e-79
Identities = 143/323 (44%), Positives = 205/323 (63%), Gaps = 5/323 (1%)
Frame = +3
Query: 17 IIVGAGPSGIAVAACLSEQGVPSLILERSDCIASLWQNRTYDRLKLHLPKHFCELPMMSF 76
II+GAG SGIA A CL++Q +P ++LER DC ASLWQ TYDRL LHL K CELP + F
Sbjct: 57 IIIGAGTSGIATAGCLTKQSIPYIMLEREDCFASLWQKYTYDRLHLHLRKQVCELPHLPF 236
Query: 77 PQTFPKYPTKHQFISYMESYADHFHIHPRFNQTVLSAEFDSTSQIWMVRT---KEGDF-Q 132
P+++P Y + QFI Y+ +Y +HF I P + + V E+D IW V+ + G+ +
Sbjct: 237 PKSYPHYVPRKQFIDYLGNYVNHFEIKPLYQRAVELVEYDGWKGIWRVKAQNRRSGELEE 416
Query: 133 YFSPWLIVATGENAEPVFPTIHGMEHFHGPVVHTSDYKSGSEYKNKKVLVIGCGNSGMEV 192
Y +L+VA+GE AEP P I G+E F+G V+H++ YK+G+E+KNK VLV+G GNSGME+
Sbjct: 417 YAGKYLVVASGETAEPRLPQIQGLESFNGKVIHSTAYKNGNEFKNKHVLVVGSGNSGMEI 596
Query: 193 SLDLCRHNAMPHLVARNSVHILPRDMFGFSTYGIAMGLYKWLPLKLVDKFLLLVSSFFLG 252
+LDL A P ++ R+ VH L RDM +++ + +L L V+K L++VS G
Sbjct: 597 ALDLSNFGAKPSIIVRSPVHFLSRDMMYYASL-----MLNYLSLSTVEKVLVMVSKVVYG 761
Query: 253 NTNHYGIKRPKTGPIELKLATGKTPVLDVGQIAQIKSGNIKVMEG-VKEITRNGAKFLDG 311
+ + YGI P GP +K+ K P++DVG + +IKS I+V+ +K I N F DG
Sbjct: 762 DLSEYGIPYPSEGPFTMKMKYAKFPIIDVGTVKKIKSREIQVLPAEIKSIRGNEVLFQDG 941
Query: 312 QEKEFDAIILATGYKSNVPSWLK 334
+ FD+I+ TG+K + WLK
Sbjct: 942 KSYTFDSIVFCTGFKRSTQKWLK 1010
>CA818897 similar to GP|16555354|gb flavin-containing monooxygenase YUCCA2
{Arabidopsis thaliana}, partial (26%)
Length = 445
Score = 179 bits (454), Expect = 2e-45
Identities = 88/148 (59%), Positives = 105/148 (70%), Gaps = 5/148 (3%)
Frame = +1
Query: 93 MESYADHFHIHPRFNQTVLSAEFDSTSQIWMVRT-----KEGDFQYFSPWLIVATGENAE 147
+++YADHF I P F+QTV+SAEFD Q+W V+T KE +Y WLIVATGE AE
Sbjct: 1 LKAYADHFDIKPVFSQTVVSAEFDHVCQLWRVKTRGVIKKEDTAEYVCQWLIVATGECAE 180
Query: 148 PVFPTIHGMEHFHGPVVHTSDYKSGSEYKNKKVLVIGCGNSGMEVSLDLCRHNAMPHLVA 207
V P I GM F G +VHTS YKSGS + K VLV+GCGNSGMEV LDLC HNA P LV
Sbjct: 181 EVVPQIEGMGEFEGQIVHTSKYKSGSMFCGKNVLVVGCGNSGMEVCLDLCNHNARPSLVV 360
Query: 208 RNSVHILPRDMFGFSTYGIAMGLYKWLP 235
R++VHILP+ M G ST+G++M L KW P
Sbjct: 361 RDTVHILPQQMLGKSTFGLSMFLLKWFP 444
>TC221849
Length = 452
Score = 163 bits (412), Expect = 1e-40
Identities = 74/124 (59%), Positives = 97/124 (77%)
Frame = +1
Query: 211 VHILPRDMFGFSTYGIAMGLYKWLPLKLVDKFLLLVSSFFLGNTNHYGIKRPKTGPIELK 270
VHILP+ M G ST+G++M L KW P++ VD+FLLL+S LG+T +G++RPK GP+ELK
Sbjct: 73 VHILPQQMLGKSTFGLSMFLLKWFPIRFVDQFLLLMSHLMLGDTAQFGLRRPKLGPLELK 252
Query: 271 LATGKTPVLDVGQIAQIKSGNIKVMEGVKEITRNGAKFLDGQEKEFDAIILATGYKSNVP 330
GKTPVLDVG + QIK+G IKV G+K + RN +F+DG+ + FDA++LATGYKSNVP
Sbjct: 253 NLYGKTPVLDVGTLTQIKNGKIKVCRGIKRLARNAVEFVDGKVENFDAMVLATGYKSNVP 432
Query: 331 SWLK 334
SWLK
Sbjct: 433 SWLK 444
>BU547429
Length = 665
Score = 158 bits (399), Expect = 4e-39
Identities = 72/124 (58%), Positives = 95/124 (76%)
Frame = -2
Query: 211 VHILPRDMFGFSTYGIAMGLYKWLPLKLVDKFLLLVSSFFLGNTNHYGIKRPKTGPIELK 270
VHILP+ M G ST+G++M L KW P++ VD+FLLL+S +T +G++RPK GP+ELK
Sbjct: 580 VHILPQQMXGKSTFGLSMFLLKWFPIRFVDQFLLLMSHLMXXDTAQFGLRRPKLGPLELK 401
Query: 271 LATGKTPVLDVGQIAQIKSGNIKVMEGVKEITRNGAKFLDGQEKEFDAIILATGYKSNVP 330
GKTPVLDVG + QIK+G IKV G+K + RN +F+DG+ + FDA++LATGYKSNVP
Sbjct: 400 NLYGKTPVLDVGTLTQIKNGKIKVCRGIKRLARNAVEFVDGKVENFDAMVLATGYKSNVP 221
Query: 331 SWLK 334
SWLK
Sbjct: 220 SWLK 209
>AW707041
Length = 365
Score = 149 bits (377), Expect = 1e-36
Identities = 68/119 (57%), Positives = 89/119 (74%)
Frame = +1
Query: 141 ATGENAEPVFPTIHGMEHFHGPVVHTSDYKSGSEYKNKKVLVIGCGNSGMEVSLDLCRHN 200
ATGENAE V P I G+ F G V+H DYKSG ++ KKVLV+GCGNSGME+SLDLC H+
Sbjct: 7 ATGENAECVMPEIEGLSEFKGDVIHACDYKSGERFRGKKVLVVGCGNSGMELSLDLCNHH 186
Query: 201 AMPHLVARNSVHILPRDMFGFSTYGIAMGLYKWLPLKLVDKFLLLVSSFFLGNTNHYGI 259
+ P +V R+SVH+LPR++FG S + +A+ L +WLPL LVDK LL+++ F LGN G+
Sbjct: 187 SSPSMVVRSSVHVLPREVFGISIFELAVMLLQWLPLWLVDKILLILAWFVLGNIEKLGL 363
>TC206675
Length = 957
Score = 84.0 bits (206), Expect = 9e-17
Identities = 38/89 (42%), Positives = 61/89 (67%), Gaps = 1/89 (1%)
Frame = +2
Query: 127 KEGDFQYFSPW-LIVATGENAEPVFPTIHGMEHFHGPVVHTSDYKSGSEYKNKKVLVIGC 185
+ G+ + ++ W L++A+GE AEP P I G+E F+G V+H++ Y +G E+K+K VLV+
Sbjct: 152 RSGEVEEYAGWYLVLASGETAEPRVPQIEGLESFNGKVIHSTGYNNGKEFKDKLVLVV*S 331
Query: 186 GNSGMEVSLDLCRHNAMPHLVARNSVHIL 214
GNSGME++LDL A P ++ R+ + L
Sbjct: 332 GNSGMEIALDLSDFGAKPSIIVRSPIGTL 418
Score = 30.4 bits (67), Expect = 1.2
Identities = 12/36 (33%), Positives = 20/36 (55%)
Frame = +1
Query: 299 KEITRNGAKFLDGQEKEFDAIILATGYKSNVPSWLK 334
K I + F DG+ + FD+I+ T +K + WL+
Sbjct: 652 KSIRGDEVLFRDGKSRPFDSIVFCTRFKRSTQKWLQ 759
>TC230122
Length = 779
Score = 60.8 bits (146), Expect = 8e-10
Identities = 54/211 (25%), Positives = 86/211 (40%), Gaps = 38/211 (18%)
Frame = +3
Query: 17 IIVGAGPSGIAVAACLSEQGVPSLILERSDCIASLWQNRTYDRLKLHLPKHFCELPMMSF 76
+I+GAG SG+ + E G ++ E D + LW++ T + KL K + +
Sbjct: 150 VIIGAGISGLVACKYVIEFGFNPIVFEVDDGVGGLWRH-TMESTKLQNNKQMFQFMDFPW 326
Query: 77 PQTFPK-YPTKHQFISYMESYADHFHIHP--RFNQTVLSAEF------------------ 115
P + + P+ Q + Y+ SYA+HF + P RFN V+ ++
Sbjct: 327 PPSVKEDNPSHKQVLDYVNSYAEHFSLIPYIRFNSKVIDIDYVGGESSEEMKSWELWGGN 506
Query: 116 ------DSTSQIWMVRTKEGDFQYFSPWLIVA-----TGENAEPVFPTIHGMEHFHGPVV 164
T I + TK + +V +G P FP G E F+G V+
Sbjct: 507 GRPFCSKGTWHIAVQHTKNLSIEVHEAEFVVLCIGKYSGFPNIPEFPPGKGPEVFNGKVM 686
Query: 165 HTSDYK------SGSEYKNKKVLVIGCGNSG 189
H+ DY + K K+V +IG SG
Sbjct: 687 HSMDYSNLDNDTAAELVKGKRVTIIGSLKSG 779
>BU082229 similar to GP|9757850|dbj dimethylaniline monooxygenase
(N-oxide-forming)-like protein {Arabidopsis thaliana},
partial (22%)
Length = 423
Score = 47.8 bits (112), Expect = 7e-06
Identities = 27/92 (29%), Positives = 50/92 (54%), Gaps = 4/92 (4%)
Frame = +2
Query: 122 WMVRTK----EGDFQYFSPWLIVATGENAEPVFPTIHGMEHFHGPVVHTSDYKSGSEYKN 177
W+VR+K E + + ++VATG + P P I GM + +H+ Y+S ++
Sbjct: 83 WVVRSKDKNSEEEVEQVFDAVVVATGHYSNPRLPCIQGMAIWKRKQMHSHIYRSPEPFRG 262
Query: 178 KKVLVIGCGNSGMEVSLDLCRHNAMPHLVARN 209
+ V+V+G SG E+S++L + HL +++
Sbjct: 263 EIVVVVGNSFSGQEISMELVKVVKELHLSSKS 358
>BF070836 similar to GP|21740855|emb OSJNBb0072N21.4 {Oryza sativa (japonica
cultivar-group)}, partial (4%)
Length = 403
Score = 47.8 bits (112), Expect = 7e-06
Identities = 38/117 (32%), Positives = 53/117 (44%), Gaps = 7/117 (5%)
Frame = +1
Query: 148 PVFPTIHGMEHFHGPVVHTSDYK------SGSEYKNKKVLVIGCGNSGMEVSLDLCRHNA 201
P FP G E F+G V+H DY + K K+V +IG SG++++ N
Sbjct: 10 PEFPPGKGPEVFNGRVMHYMDYSNLDNETAAELIKGKRVTIIGSQKSGLDLAAQCSNANG 189
Query: 202 MPHLVARNSVHILPRDMFGFSTYGIAMGLYK-WLPLKLVDKFLLLVSSFFLGNTNHY 257
VA N +H L +D+FG+ T LY WL FLL +TNH+
Sbjct: 190 EKFFVA-NFLHKLQKDLFGYPT-----SLYS*WL------YFLLNFPLLKFHSTNHF 324
>TC223802 similar to UP|Q9FNA2 (Q9FNA2) Polyamine oxidase, partial (24%)
Length = 425
Score = 37.7 bits (86), Expect = 0.007
Identities = 22/49 (44%), Positives = 29/49 (58%), Gaps = 1/49 (2%)
Frame = +2
Query: 17 IIVGAGPSGIAVAACLSEQGVPSL-ILERSDCIASLWQNRTYDRLKLHL 64
IIVGAG SGI+ A L+E GV L ILE S+CI + + + + L
Sbjct: 17 IIVGAGVSGISAAKLLAENGVKDLVILEASNCIGGRIRKENFGGVSVEL 163
>BF070101 similar to GP|9759603|dbj| dimethylaniline monooxygenase
(N-oxide-forming)-like protein {Arabidopsis thaliana},
partial (9%)
Length = 382
Score = 37.0 bits (84), Expect = 0.013
Identities = 22/79 (27%), Positives = 41/79 (51%), Gaps = 3/79 (3%)
Frame = +2
Query: 40 LILERSDCIASLWQNRTYDRLKLHLPKHFCELPMMSFPQTFPK-YPTKHQFISYMESYAD 98
L L+ + + LW++ T + KL K + ++P + + P+ Q + Y+ SYA+
Sbjct: 143 LSLKLRNGVGGLWRH-TIESTKLQNKKQMYQFLDFAWPSSVKEDNPSHEQVLDYLNSYAE 319
Query: 99 HFHIHP--RFNQTVLSAEF 115
HF + P RFN V+ ++
Sbjct: 320 HFSLIPYIRFNSKVIDIDY 376
>AI965600
Length = 346
Score = 35.4 bits (80), Expect = 0.037
Identities = 14/26 (53%), Positives = 18/26 (68%)
Frame = +2
Query: 257 YGIKRPKTGPIELKLATGKTPVLDVG 282
YG+ RP GP +K+ GK PV+DVG
Sbjct: 266 YGVARPNEGPFYMKVKYGKYPVIDVG 343
>TC233712 similar to UP|O23024 (O23024) T1G11.14 protein (Flavin-containing
monooxygenase YUCCA3) (At1g04610), partial (21%)
Length = 741
Score = 33.9 bits (76), Expect = 0.11
Identities = 13/16 (81%), Positives = 15/16 (93%)
Frame = +3
Query: 319 IILATGYKSNVPSWLK 334
++LATGY SNVPSWLK
Sbjct: 18 VVLATGYHSNVPSWLK 65
>BQ473725
Length = 307
Score = 32.7 bits (73), Expect = 0.24
Identities = 17/54 (31%), Positives = 29/54 (53%), Gaps = 6/54 (11%)
Frame = +3
Query: 148 PVFPTIHGMEHFHGPVVHTSDY------KSGSEYKNKKVLVIGCGNSGMEVSLD 195
P FP G E F+G V+H+ D+ + K K+V +IG SG++++ +
Sbjct: 135 PEFPPGKGPEVFNGKVMHSMDFFNLDNETAAELIKGKRVTIIGSQKSGLDLAAE 296
>TC215556 weakly similar to GB|AAP37810.1|30725576|BT008451 At3g44190
{Arabidopsis thaliana;} , partial (89%)
Length = 1238
Score = 30.4 bits (67), Expect = 1.2
Identities = 15/43 (34%), Positives = 23/43 (52%)
Frame = +2
Query: 288 KSGNIKVMEGVKEITRNGAKFLDGQEKEFDAIILATGYKSNVP 330
K GN+ V V IT DGQ+ +D +++ATG+ +P
Sbjct: 269 KKGNLVVSSAVN-ITETAVVTEDGQQIAYDYLVIATGHTEPIP 394
>TC212119 weakly similar to UP|Q9LDR3 (Q9LDR3) Arabidopsis thaliana genomic
DNA, chromosome 3, P1 clone: MJK13 (MJK13.8 protein)
(At3g15420), partial (25%)
Length = 493
Score = 30.4 bits (67), Expect = 1.2
Identities = 11/24 (45%), Positives = 16/24 (65%)
Frame = -2
Query: 57 YDRLKLHLPKHFCELPMMSFPQTF 80
+DR+KLHLP + C +P P +F
Sbjct: 372 HDRVKLHLPPYSC*MPKSQLPNSF 301
>TC218076 similar to UP|Q9MUE2 (Q9MUE2) S-adenosyl-L-methionine
Mg-protoporphyrin IX methyltranserase , partial (59%)
Length = 829
Score = 30.0 bits (66), Expect = 1.5
Identities = 16/52 (30%), Positives = 24/52 (45%)
Frame = -2
Query: 48 IASLWQNRTYDRLKLHLPKHFCELPMMSFPQTFPKYPTKHQFISYMESYADH 99
++SLW ++ + L LHLP+ + P P K Q + S ADH
Sbjct: 555 LSSLWSSQPFAMLSLHLPR----------SEDMPLLPLKDQGTTLQPSLADH 430
>TC229237 similar to UP|Q6TF29 (Q6TF29) Rapid alkalinization factor 1,
partial (51%)
Length = 605
Score = 29.6 bits (65), Expect = 2.0
Identities = 17/56 (30%), Positives = 28/56 (49%), Gaps = 2/56 (3%)
Frame = -2
Query: 180 VLVIGCGNS--GMEVSLDLCRHNAMPHLVARNSVHILPRDMFGFSTYGIAMGLYKW 233
VL + C NS G+ + L LC+ A+P+ + N FG Y + + ++KW
Sbjct: 238 VLFLHCQNSSVGLRIHLRLCQAQALPNALGANP--------FGHHIYFMEIEVHKW 95
>TC225480 UP|Q9XE94 (Q9XE94) Geranylgeranyl hydrogenase, complete
Length = 1746
Score = 28.9 bits (63), Expect = 3.4
Identities = 12/27 (44%), Positives = 19/27 (69%)
Frame = +1
Query: 18 IVGAGPSGIAVAACLSEQGVPSLILER 44
+VG GP+G A A L++ GV + ++ER
Sbjct: 166 VVGGGPAGGAAAETLAKGGVETFLIER 246
>TC219486
Length = 1900
Score = 28.5 bits (62), Expect = 4.5
Identities = 17/53 (32%), Positives = 26/53 (48%)
Frame = -2
Query: 66 KHFCELPMMSFPQTFPKYPTKHQFISYMESYADHFHIHPRFNQTVLSAEFDST 118
KHF P+ + +T + P K Q ++Y DH + HP N T+ + F T
Sbjct: 834 KHFITNPLNTNDET--RRPNKTQ-----QTYIDHMNPHPHRNATLQNESFSFT 697
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.322 0.139 0.433
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,466,565
Number of Sequences: 63676
Number of extensions: 281205
Number of successful extensions: 1241
Number of sequences better than 10.0: 45
Number of HSP's better than 10.0 without gapping: 1224
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1236
length of query: 337
length of database: 12,639,632
effective HSP length: 98
effective length of query: 239
effective length of database: 6,399,384
effective search space: 1529452776
effective search space used: 1529452776
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC144617.10


