

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
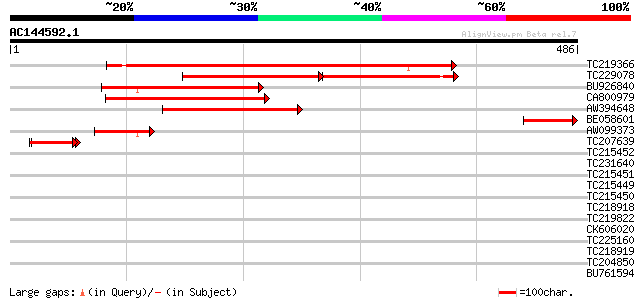
Query= AC144592.1 - phase: 0 /pseudo
(486 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC219366 similar to UP|Q9FMR1 (Q9FMR1) Rac GTPase activating pro... 369 e-102
TC229078 similar to UP|O82458 (O82458) Rac GTPase activating pro... 180 5e-76
BU926840 similar to PIR|T01383|T01 GTPase-activating protein hom... 199 2e-51
CA800979 similar to GP|3695059|gb| rac GTPase activating protein... 188 5e-48
AW394648 similar to GP|3695059|gb| rac GTPase activating protein... 187 7e-48
BE058601 similar to GP|3695061|gb|A rac GTPase activating protei... 85 6e-17
AW099373 similar to GP|3695063|gb|A rac GTPase activating protei... 50 2e-06
TC207639 similar to UP|O48592 (O48592) GT2 protein, partial (24%) 42 6e-04
TC215452 similar to UP|P93488 (P93488) NAP1Ps, partial (29%) 39 0.005
TC231640 homologue to UP|Q918P0 (Q918P0) Latency associated anti... 39 0.007
TC215451 weakly similar to UP|Q94K07 (Q94K07) Nucleosome assembl... 39 0.007
TC215449 similar to UP|P93488 (P93488) NAP1Ps, partial (97%) 38 0.009
TC215450 similar to UP|P93488 (P93488) NAP1Ps, partial (30%) 38 0.012
TC218918 UP|Q39892 (Q39892) Nucleosome assembly protein 1, complete 37 0.015
TC219822 homologue to UP|CENB_HUMAN (P07199) Major centromere au... 37 0.015
CK606020 37 0.020
TC225160 UP|Q948X9 (Q948X9) Beta-conglycinin alpha-subunit, comp... 37 0.020
TC218919 homologue to UP|Q39892 (Q39892) Nucleosome assembly pro... 37 0.026
TC204850 similar to GB|AAM16178.1|20334844|AY094022 AT3g12390/T2... 37 0.026
BU761594 similar to GP|22654971|gb At1g80930/F23A5_23 {Arabidops... 36 0.034
>TC219366 similar to UP|Q9FMR1 (Q9FMR1) Rac GTPase activating protein,
partial (52%)
Length = 1397
Score = 369 bits (946), Expect = e-102
Identities = 189/316 (59%), Positives = 236/316 (73%), Gaps = 16/316 (5%)
Frame = +1
Query: 84 ILDIVMAALKKSIVTCSVEREDVSSL-DISWPTEVRHVSHVTFDRFNGFLGLPSEFQPEV 142
+L +++ L+KS + R+++ S+ DI PT VRHV+HVTFDRFNGFLGLP EF+PEV
Sbjct: 7 LLALLLTLLRKSF---QLPRKNIGSIMDIGSPTNVRHVAHVTFDRFNGFLGLPVEFEPEV 177
Query: 143 PTRVPSASVKVFGVSAKSMQCSYDDRGNSVPTILLRMQKQLYSEGGLKAEGIFRITAENS 202
P R PSAS VFGVS +SMQ SYD RGNSVPTILL MQ+ LY +GGL+ EGIFRI A+N
Sbjct: 178 PRRPPSASASVFGVSTESMQLSYDSRGNSVPTILLLMQRHLYVQGGLQVEGIFRINADNG 357
Query: 203 QEAFVRDQLNKGVVPHGIDVHCLSGLIKAWFRELPTGVLDSLTPEQVMQCNTEDDCTNLV 262
QE RDQLN GVVP GIDVHCL+GLIKAWFRELPTG+LDSL+PEQVMQC TED+C LV
Sbjct: 358 QEEHARDQLNLGVVPEGIDVHCLAGLIKAWFRELPTGILDSLSPEQVMQCQTEDECAELV 537
Query: 263 KLLPSTEAALLDWAINLMADVVENEQFNKMNARNVAMVFAPNMTQMADPLTALIHAVQVM 322
+ LP TEA+LLDWAINLMADVV++E NKMNA N+AMVFAPNMTQMADP++AL++AVQVM
Sbjct: 538 RHLPHTEASLLDWAINLMADVVQHENVNKMNAHNIAMVFAPNMTQMADPISALMYAVQVM 717
Query: 323 NFLKTLILKMLREREESI---------------DNARLLSHSMDFASCNDEFPPFSFNKE 367
NFLKTLIL+ +RER++S+ +N R+L +E +F E
Sbjct: 718 NFLKTLILRTVRERKDSVVESCPRFYLQPSVDNENRRILESFRQDTPAENEEAQENFVLE 897
Query: 368 ESSCEQIEDACDKNSS 383
+++ ++ ++ NS+
Sbjct: 898 KTALDRSPESLQNNST 945
>TC229078 similar to UP|O82458 (O82458) Rac GTPase activating protein 1,
partial (70%)
Length = 1579
Score = 180 bits (457), Expect(2) = 5e-76
Identities = 87/121 (71%), Positives = 102/121 (83%)
Frame = +1
Query: 149 ASVKVFGVSAKSMQCSYDDRGNSVPTILLRMQKQLYSEGGLKAEGIFRITAENSQEAFVR 208
+S VFGVS +SMQ S+D RGNSVPTILL MQ+ LY++GGL+AEGIFRI AEN QE FVR
Sbjct: 61 SSANVFGVSTESMQLSFDARGNSVPTILLLMQRHLYAQGGLQAEGIFRIDAENGQEEFVR 240
Query: 209 DQLNKGVVPHGIDVHCLSGLIKAWFRELPTGVLDSLTPEQVMQCNTEDDCTNLVKLLPST 268
+ LN+GVVP GIDVHCL+GLIKAWFRELPTGVLD +PEQVMQ +E++C LV+LLP T
Sbjct: 241 E*LNRGVVPDGIDVHCLAGLIKAWFRELPTGVLDPFSPEQVMQSQSEEECAQLVRLLPPT 420
Query: 269 E 269
E
Sbjct: 421 E 423
Score = 122 bits (307), Expect(2) = 5e-76
Identities = 67/116 (57%), Positives = 81/116 (69%)
Frame = +2
Query: 269 EAALLDWAINLMADVVENEQFNKMNARNVAMVFAPNMTQMADPLTALIHAVQVMNFLKTL 328
++ALLDWAINLMADV + E NKMNARN+AMVFAPNMTQMADPLTAL++AVQVMNFLKTL
Sbjct: 422 KSALLDWAINLMADVAQMENLNKMNARNIAMVFAPNMTQMADPLTALMYAVQVMNFLKTL 601
Query: 329 ILKMLREREESIDNARLLSHSMDFASCNDEFPPFSFNKEESSCEQIEDACDKNSST 384
++K LREREESI + + F + +KE+S E D D + T
Sbjct: 602 VVKALREREESIVKSNPVPDLNSFDDDGNHSNSEMLDKEDS--ENGNDCSDDDEDT 763
>BU926840 similar to PIR|T01383|T01 GTPase-activating protein homolog T4I9.2
- Arabidopsis thaliana, partial (32%)
Length = 443
Score = 199 bits (506), Expect = 2e-51
Identities = 98/144 (68%), Positives = 121/144 (83%), Gaps = 5/144 (3%)
Frame = +3
Query: 79 QNQFAILDIVMAALKKSIVTCSVER-EDVSS----LDISWPTEVRHVSHVTFDRFNGFLG 133
QNQ +++ +++AA++KS+V+C V+ EDV S ++I WPT V+H++HVTFDRFNGFLG
Sbjct: 12 QNQLSLVALLLAAIRKSMVSCRVDPPEDVISTVHHMEIGWPTNVQHITHVTFDRFNGFLG 191
Query: 134 LPSEFQPEVPTRVPSASVKVFGVSAKSMQCSYDDRGNSVPTILLRMQKQLYSEGGLKAEG 193
LP EFQ E+P RVPSASV VFGVSA+SMQCSYD +GNSVPTILL MQ +LYS+GGLKAEG
Sbjct: 192 LPYEFQVEIPARVPSASVSVFGVSAESMQCSYDPKGNSVPTILLLMQDRLYSQGGLKAEG 371
Query: 194 IFRITAENSQEAFVRDQLNKGVVP 217
IFRI ENSQE VRDQLN+G+VP
Sbjct: 372 IFRINPENSQEEHVRDQLNRGIVP 443
>CA800979 similar to GP|3695059|gb| rac GTPase activating protein 1 {Lotus
japonicus}, partial (25%)
Length = 421
Score = 188 bits (477), Expect = 5e-48
Identities = 89/140 (63%), Positives = 111/140 (78%)
Frame = +2
Query: 83 AILDIVMAALKKSIVTCSVEREDVSSLDISWPTEVRHVSHVTFDRFNGFLGLPSEFQPEV 142
++L I++ L+KS++ C+ E +++I WPT VRHV+HVTFDRFNGFLGLP EF+PEV
Sbjct: 2 SLLAILVTLLRKSLIACNKSEEGHGAMEIGWPTNVRHVAHVTFDRFNGFLGLPREFEPEV 181
Query: 143 PTRVPSASVKVFGVSAKSMQCSYDDRGNSVPTILLRMQKQLYSEGGLKAEGIFRITAENS 202
TR PSAS VFGVS +SMQ SYD RGNSVPTILL MQ+ LY+ GGL+ EGIFRI A+NS
Sbjct: 182 STRPPSASATVFGVSTESMQLSYDTRGNSVPTILLLMQRHLYALGGLQEEGIFRINADNS 361
Query: 203 QEAFVRDQLNKGVVPHGIDV 222
QE VRDQLN+G+VP +D+
Sbjct: 362 QEESVRDQLNRGLVPEDVDI 421
>AW394648 similar to GP|3695059|gb| rac GTPase activating protein 1 {Lotus
japonicus}, partial (24%)
Length = 361
Score = 187 bits (476), Expect = 7e-48
Identities = 91/120 (75%), Positives = 103/120 (85%)
Frame = +2
Query: 132 LGLPSEFQPEVPTRVPSASVKVFGVSAKSMQCSYDDRGNSVPTILLRMQKQLYSEGGLKA 191
LGLP EF+PEVP R PSAS VFGVS +SMQ S+D RGNSVPTILL MQ+ LY++GGL+A
Sbjct: 2 LGLPVEFEPEVPRRPPSASANVFGVSTESMQLSFDARGNSVPTILLLMQRHLYAQGGLQA 181
Query: 192 EGIFRITAENSQEAFVRDQLNKGVVPHGIDVHCLSGLIKAWFRELPTGVLDSLTPEQVMQ 251
EGIFRI AEN QE FVR+QLN+G+VP GIDVHCL+GLIKAWFRELP VLD L+PEQVMQ
Sbjct: 182 EGIFRINAENGQEEFVREQLNRGIVPDGIDVHCLAGLIKAWFRELPNWVLDPLSPEQVMQ 361
>BE058601 similar to GP|3695061|gb|A rac GTPase activating protein 2 {Lotus
japonicus}, partial (12%)
Length = 307
Score = 85.1 bits (209), Expect = 6e-17
Identities = 38/46 (82%), Positives = 42/46 (90%)
Frame = +2
Query: 441 YRRRYDSEHWSRLRNGVRKLCRHPVFQLSKPSKKPASLGIVNTREG 486
YR YDSEHW +LRNGVR+LCRHPVFQLSKP+KK ASLG+VNTREG
Sbjct: 2 YRGIYDSEHWVKLRNGVRRLCRHPVFQLSKPTKKRASLGVVNTREG 139
>AW099373 similar to GP|3695063|gb|A rac GTPase activating protein 3 {Lotus
japonicus}, partial (12%)
Length = 416
Score = 50.4 bits (119), Expect = 2e-06
Identities = 26/58 (44%), Positives = 38/58 (64%), Gaps = 6/58 (10%)
Frame = +2
Query: 73 GSNNNHQNQFAILDIVMAALKKSIVTCSVER-EDVSS-----LDISWPTEVRHVSHVT 124
G QN + + +++AAL+KS+V CSV+ +DV S ++I WPT V+HVSHVT
Sbjct: 242 GGGEEEQNPASPVAVLLAALRKSMVACSVDSPDDVISAVHHPMEIGWPTNVKHVSHVT 415
>TC207639 similar to UP|O48592 (O48592) GT2 protein, partial (24%)
Length = 1689
Score = 42.0 bits (97), Expect = 6e-04
Identities = 16/41 (39%), Positives = 29/41 (70%)
Frame = +1
Query: 18 SEFNPAAQVTNNKKKEEQQEQEEEDEEEELESDDNDNDDDD 58
S++ A NN + +++E+EEEDEEEE E ++++N++ D
Sbjct: 868 SQYEQGAAKENNNNERKEREEEEEDEEEEEEEEEDENEEGD 990
Score = 42.0 bits (97), Expect = 6e-04
Identities = 17/42 (40%), Positives = 29/42 (68%)
Frame = +1
Query: 19 EFNPAAQVTNNKKKEEQQEQEEEDEEEELESDDNDNDDDDEI 60
E A + NN++KE ++E+E+E+EEEE E D+N+ D + +
Sbjct: 877 EQGAAKENNNNERKEREEEEEDEEEEEEEEEDENEEGDLESV 1002
Score = 31.2 bits (69), Expect = 1.1
Identities = 11/41 (26%), Positives = 23/41 (55%)
Frame = +1
Query: 19 EFNPAAQVTNNKKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
++ P +Q KE + +E EEEE + ++ + +++DE
Sbjct: 853 QWRPPSQYEQGAAKENNNNERKEREEEEEDEEEEEEEEEDE 975
>TC215452 similar to UP|P93488 (P93488) NAP1Ps, partial (29%)
Length = 609
Score = 38.9 bits (89), Expect = 0.005
Identities = 17/42 (40%), Positives = 26/42 (61%)
Frame = +3
Query: 33 EEQQEQEEEDEEEELESDDNDNDDDDEIYSSNNASRQFTKGS 74
E+ ++ EE+D+E+E + DD+D+DDDDE S K S
Sbjct: 147 EDDEDIEEDDDEDEEDDDDDDDDDDDEEESKTKKKSSAPKKS 272
Score = 37.0 bits (84), Expect = 0.020
Identities = 14/27 (51%), Positives = 21/27 (76%)
Frame = +3
Query: 33 EEQQEQEEEDEEEELESDDNDNDDDDE 59
+E E EED++E+ E DD+D+DDDD+
Sbjct: 144 DEDDEDIEEDDDEDEEDDDDDDDDDDD 224
Score = 35.8 bits (81), Expect = 0.044
Identities = 12/28 (42%), Positives = 24/28 (84%)
Frame = +3
Query: 32 KEEQQEQEEEDEEEELESDDNDNDDDDE 59
++++ +E++DE+EE + DD+D+DDD+E
Sbjct: 147 EDDEDIEEDDDEDEEDDDDDDDDDDDEE 230
Score = 32.3 bits (72), Expect = 0.49
Identities = 13/40 (32%), Positives = 24/40 (59%)
Frame = +3
Query: 35 QQEQEEEDEEEELESDDNDNDDDDEIYSSNNASRQFTKGS 74
+ ++++ED EE+ + D+ D+DDDD+ S+ K S
Sbjct: 138 EDDEDDEDIEEDDDEDEEDDDDDDDDDDDEEESKTKKKSS 257
Score = 31.2 bits (69), Expect = 1.1
Identities = 9/27 (33%), Positives = 22/27 (81%)
Frame = +3
Query: 33 EEQQEQEEEDEEEELESDDNDNDDDDE 59
E+ ++ E+ +E+++ + +D+D+DDDD+
Sbjct: 138 EDDEDDEDIEEDDDEDEEDDDDDDDDD 218
Score = 29.6 bits (65), Expect = 3.2
Identities = 8/27 (29%), Positives = 23/27 (84%)
Frame = +3
Query: 33 EEQQEQEEEDEEEELESDDNDNDDDDE 59
+E + E+++++E++E DD+++++DD+
Sbjct: 120 DEFGDLEDDEDDEDIEEDDDEDEEDDD 200
>TC231640 homologue to UP|Q918P0 (Q918P0) Latency associated antigen,
partial (7%)
Length = 466
Score = 38.5 bits (88), Expect = 0.007
Identities = 14/35 (40%), Positives = 27/35 (77%)
Frame = +1
Query: 25 QVTNNKKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
Q +K+EE++E+EEE+EEEE E ++ + ++++E
Sbjct: 22 QEEEEEKEEEEEEEEEEEEEEEEEEEEEEKEEEEE 126
Score = 37.0 bits (84), Expect = 0.020
Identities = 14/35 (40%), Positives = 26/35 (74%)
Frame = +1
Query: 30 KKKEEQQEQEEEDEEEELESDDNDNDDDDEIYSSN 64
++KEE++E+EEE+EEEE E ++ + +++E N
Sbjct: 34 EEKEEEEEEEEEEEEEEEEEEEEEEKEEEEEEEGN 138
Score = 35.8 bits (81), Expect = 0.044
Identities = 12/34 (35%), Positives = 28/34 (82%)
Frame = +1
Query: 26 VTNNKKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
V ++ ++EE++++EEE+EEEE E ++ + ++++E
Sbjct: 7 VLSDTQEEEEEKEEEEEEEEEEEEEEEEEEEEEE 108
Score = 35.4 bits (80), Expect = 0.058
Identities = 12/33 (36%), Positives = 26/33 (78%)
Frame = +1
Query: 27 TNNKKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
T +++E+++E+EEE+EEEE E ++ + ++ +E
Sbjct: 19 TQEEEEEKEEEEEEEEEEEEEEEEEEEEEEKEE 117
Score = 35.4 bits (80), Expect = 0.058
Identities = 11/34 (32%), Positives = 29/34 (84%)
Frame = +1
Query: 26 VTNNKKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
+++ +++EE++E+EEE+EEEE E ++ + +++++
Sbjct: 10 LSDTQEEEEEKEEEEEEEEEEEEEEEEEEEEEEK 111
Score = 35.0 bits (79), Expect = 0.075
Identities = 13/45 (28%), Positives = 28/45 (61%)
Frame = +1
Query: 21 NPAAQVTNNKKKEEQQEQEEEDEEEELESDDNDNDDDDEIYSSNN 65
N + ++++E++E+EEE+EEEE E ++ + + ++E N
Sbjct: 4 NVLSDTQEEEEEKEEEEEEEEEEEEEEEEEEEEEEKEEEEEEEGN 138
Score = 33.5 bits (75), Expect = 0.22
Identities = 12/44 (27%), Positives = 28/44 (63%)
Frame = +1
Query: 17 VSEFNPAAQVTNNKKKEEQQEQEEEDEEEELESDDNDNDDDDEI 60
+S+ + +++EE++E+EEE+EEEE E + + +++ +
Sbjct: 10 LSDTQEEEEEKEEEEEEEEEEEEEEEEEEEEEEKEEEEEEEGNV 141
Score = 32.7 bits (73), Expect = 0.37
Identities = 26/116 (22%), Positives = 50/116 (42%), Gaps = 24/116 (20%)
Frame = +1
Query: 11 NIAGFSVSEFNPAAQVTNNKKKEEQQEQEEEDEEEELESD----------DNDNDDDDEI 60
N+ + E + +++EE++E+EEE+EEE+ E + D D D +E+
Sbjct: 4 NVLSDTQEEEEEKEEEEEEEEEEEEEEEEEEEEEEKEEEEEEEGNVLVIIDEDKHDTEEL 183
Query: 61 --------------YSSNNASRQFTKGSNNNHQNQFAILDIVMAALKKSIVTCSVE 102
+SSNN + ++Q ++ V+ L+ S ++ S E
Sbjct: 184 NKKCEDFIKRMKATFSSNNLELRAVDSFYFDNQKLPVAVN*VVVDLRTSCISLSFE 351
>TC215451 weakly similar to UP|Q94K07 (Q94K07) Nucleosome assembly protein,
partial (20%)
Length = 777
Score = 38.5 bits (88), Expect = 0.007
Identities = 14/27 (51%), Positives = 23/27 (84%)
Frame = +2
Query: 33 EEQQEQEEEDEEEELESDDNDNDDDDE 59
E+ ++ EE+D+E+E + DD+D+DDDDE
Sbjct: 98 EDDEDIEEDDDEDEEDDDDDDDDDDDE 178
Score = 37.0 bits (84), Expect = 0.020
Identities = 14/27 (51%), Positives = 21/27 (76%)
Frame = +2
Query: 33 EEQQEQEEEDEEEELESDDNDNDDDDE 59
+E E EED++E+ E DD+D+DDDD+
Sbjct: 95 DEDDEDIEEDDDEDEEDDDDDDDDDDD 175
Score = 35.8 bits (81), Expect = 0.044
Identities = 12/28 (42%), Positives = 24/28 (84%)
Frame = +2
Query: 32 KEEQQEQEEEDEEEELESDDNDNDDDDE 59
++++ +E++DE+EE + DD+D+DDD+E
Sbjct: 98 EDDEDIEEDDDEDEEDDDDDDDDDDDEE 181
Score = 32.3 bits (72), Expect = 0.49
Identities = 10/27 (37%), Positives = 23/27 (85%)
Frame = +2
Query: 33 EEQQEQEEEDEEEELESDDNDNDDDDE 59
+E E++++++EE+ + DD+D+DD++E
Sbjct: 104 DEDIEEDDDEDEEDDDDDDDDDDDEEE 184
Score = 31.2 bits (69), Expect = 1.1
Identities = 9/27 (33%), Positives = 22/27 (81%)
Frame = +2
Query: 33 EEQQEQEEEDEEEELESDDNDNDDDDE 59
E+ ++ E+ +E+++ + +D+D+DDDD+
Sbjct: 89 EDDEDDEDIEEDDDEDEEDDDDDDDDD 169
Score = 31.2 bits (69), Expect = 1.1
Identities = 10/25 (40%), Positives = 20/25 (80%)
Frame = +2
Query: 35 QQEQEEEDEEEELESDDNDNDDDDE 59
+ ++++ED EE+ + D+ D+DDDD+
Sbjct: 89 EDDEDDEDIEEDDDEDEEDDDDDDD 163
Score = 30.4 bits (67), Expect = 1.9
Identities = 12/36 (33%), Positives = 22/36 (60%)
Frame = +2
Query: 37 EQEEEDEEEELESDDNDNDDDDEIYSSNNASRQFTK 72
E +E+DE+ E + D+++ DDDD+ ++ TK
Sbjct: 89 EDDEDDEDIEEDDDEDEEDDDDDDDDDDDEEESKTK 196
Score = 29.6 bits (65), Expect = 3.2
Identities = 8/27 (29%), Positives = 23/27 (84%)
Frame = +2
Query: 33 EEQQEQEEEDEEEELESDDNDNDDDDE 59
+E + E+++++E++E DD+++++DD+
Sbjct: 71 DEFGDLEDDEDDEDIEEDDDEDEEDDD 151
>TC215449 similar to UP|P93488 (P93488) NAP1Ps, partial (97%)
Length = 1495
Score = 38.1 bits (87), Expect = 0.009
Identities = 17/42 (40%), Positives = 26/42 (61%)
Frame = +2
Query: 33 EEQQEQEEEDEEEELESDDNDNDDDDEIYSSNNASRQFTKGS 74
E+ ++ EE+D+EEE E +D+D+D+DDE S K S
Sbjct: 1052 EDDEDIEEDDDEEEEEDEDDDDDEDDEEESKTKKKSSAPKKS 1177
Score = 35.8 bits (81), Expect = 0.044
Identities = 12/28 (42%), Positives = 24/28 (84%)
Frame = +2
Query: 32 KEEQQEQEEEDEEEELESDDNDNDDDDE 59
++++ +E++DEEEE + DD+D++DD+E
Sbjct: 1052 EDDEDIEEDDDEEEEEDEDDDDDEDDEE 1135
Score = 35.0 bits (79), Expect = 0.075
Identities = 13/27 (48%), Positives = 21/27 (77%)
Frame = +2
Query: 33 EEQQEQEEEDEEEELESDDNDNDDDDE 59
+E E EED++EE E D++D+DD+D+
Sbjct: 1049 DEDDEDIEEDDDEEEEEDEDDDDDEDD 1129
Score = 33.5 bits (75), Expect = 0.22
Identities = 14/40 (35%), Positives = 28/40 (70%)
Frame = +2
Query: 28 NNKKKEEQQEQEEEDEEEELESDDNDNDDDDEIYSSNNAS 67
+++ E+ +E ++E+EEE+ E DD+D DD++E + +S
Sbjct: 1046 DDEDDEDIEEDDDEEEEED-EDDDDDEDDEEESKTKKKSS 1162
Score = 30.4 bits (67), Expect = 1.9
Identities = 9/27 (33%), Positives = 22/27 (81%)
Frame = +2
Query: 33 EEQQEQEEEDEEEELESDDNDNDDDDE 59
E+ ++ E+ +E+++ E +++++DDDDE
Sbjct: 1043 EDDEDDEDIEEDDDEEEEEDEDDDDDE 1123
Score = 28.9 bits (63), Expect = 5.4
Identities = 8/27 (29%), Positives = 22/27 (80%)
Frame = +2
Query: 33 EEQQEQEEEDEEEELESDDNDNDDDDE 59
+E + E+++++E++E DD++ +++DE
Sbjct: 1025 DEFGDLEDDEDDEDIEEDDDEEEEEDE 1105
Score = 28.1 bits (61), Expect = 9.2
Identities = 10/40 (25%), Positives = 24/40 (60%)
Frame = +2
Query: 35 QQEQEEEDEEEELESDDNDNDDDDEIYSSNNASRQFTKGS 74
+ ++++ED EE+ + ++ +++DDD+ S+ K S
Sbjct: 1043 EDDEDDEDIEEDDDEEEEEDEDDDDDEDDEEESKTKKKSS 1162
>TC215450 similar to UP|P93488 (P93488) NAP1Ps, partial (30%)
Length = 672
Score = 37.7 bits (86), Expect = 0.012
Identities = 14/27 (51%), Positives = 23/27 (84%)
Frame = +3
Query: 33 EEQQEQEEEDEEEELESDDNDNDDDDE 59
E+ ++ EE+D+EEE E +D+D+D+DDE
Sbjct: 219 EDDEDIEEDDDEEEEEDEDDDDDEDDE 299
Score = 35.8 bits (81), Expect = 0.044
Identities = 12/28 (42%), Positives = 24/28 (84%)
Frame = +3
Query: 32 KEEQQEQEEEDEEEELESDDNDNDDDDE 59
++++ +E++DEEEE + DD+D++DD+E
Sbjct: 219 EDDEDIEEDDDEEEEEDEDDDDDEDDEE 302
Score = 35.4 bits (80), Expect = 0.058
Identities = 14/37 (37%), Positives = 24/37 (64%)
Frame = +3
Query: 33 EEQQEQEEEDEEEELESDDNDNDDDDEIYSSNNASRQ 69
+E E EED++EE E D++D+DD+D+ S ++
Sbjct: 216 DEDDEDIEEDDDEEEEEDEDDDDDEDDEEESKTKKKK 326
Score = 32.7 bits (73), Expect = 0.37
Identities = 13/32 (40%), Positives = 25/32 (77%)
Frame = +3
Query: 28 NNKKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
+++ E+ +E ++E+EEE+ E DD+D DD++E
Sbjct: 213 DDEDDEDIEEDDDEEEEED-EDDDDDEDDEEE 305
Score = 30.4 bits (67), Expect = 1.9
Identities = 9/27 (33%), Positives = 22/27 (81%)
Frame = +3
Query: 33 EEQQEQEEEDEEEELESDDNDNDDDDE 59
E+ ++ E+ +E+++ E +++++DDDDE
Sbjct: 210 EDDEDDEDIEEDDDEEEEEDEDDDDDE 290
Score = 28.9 bits (63), Expect = 5.4
Identities = 8/27 (29%), Positives = 22/27 (80%)
Frame = +3
Query: 33 EEQQEQEEEDEEEELESDDNDNDDDDE 59
+E + E+++++E++E DD++ +++DE
Sbjct: 192 DEFGDLEDDEDDEDIEEDDDEEEEEDE 272
>TC218918 UP|Q39892 (Q39892) Nucleosome assembly protein 1, complete
Length = 1507
Score = 37.4 bits (85), Expect = 0.015
Identities = 15/37 (40%), Positives = 26/37 (69%)
Frame = +1
Query: 23 AAQVTNNKKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
AAQ + E+ +++EE+++E+E E DD D DD++E
Sbjct: 973 AAQGDEFEDLEDDEDEEEDEDEDEDEEDDEDEDDEEE 1083
Score = 35.4 bits (80), Expect = 0.058
Identities = 12/26 (46%), Positives = 22/26 (84%)
Frame = +1
Query: 33 EEQQEQEEEDEEEELESDDNDNDDDD 58
E+++E E+EDE+EE + D++D ++DD
Sbjct: 1012 EDEEEDEDEDEDEEDDEDEDDEEEDD 1089
Score = 35.4 bits (80), Expect = 0.058
Identities = 16/44 (36%), Positives = 28/44 (63%)
Frame = +1
Query: 16 SVSEFNPAAQVTNNKKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
+VS F A + + E E EEEDE+E+ + +D++++DD+E
Sbjct: 949 AVSWFTGEAAQGDEFEDLEDDEDEEEDEDEDEDEEDDEDEDDEE 1080
>TC219822 homologue to UP|CENB_HUMAN (P07199) Major centromere autoantigen B
(Centromere protein B) (CENP-B), partial (5%)
Length = 710
Score = 37.4 bits (85), Expect = 0.015
Identities = 16/49 (32%), Positives = 31/49 (62%)
Frame = +1
Query: 21 NPAAQVTNNKKKEEQQEQEEEDEEEELESDDNDNDDDDEIYSSNNASRQ 69
N A + +++EE++E+EEE+ EEE E ++ + ++++E S N Q
Sbjct: 397 NYAPALAEPEEEEEEEEEEEEENEEEEEEEEGEEEEEEEARSMVNELAQ 543
>CK606020
Length = 340
Score = 37.0 bits (84), Expect = 0.020
Identities = 12/33 (36%), Positives = 28/33 (84%)
Frame = +1
Query: 28 NNKKKEEQQEQEEEDEEEELESDDNDNDDDDEI 60
+ +++EE++E+EEE+EEEE E ++ + ++++E+
Sbjct: 196 DEEEEEEEEEEEEEEEEEEEEIEEEEEEEEEEV 294
Score = 36.2 bits (82), Expect = 0.034
Identities = 12/30 (40%), Positives = 26/30 (86%)
Frame = +1
Query: 30 KKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
+++EE++E+EEE+EEEE+E ++ + +++ E
Sbjct: 208 EEEEEEEEEEEEEEEEEIEEEEEEEEEEVE 297
Score = 35.0 bits (79), Expect = 0.075
Identities = 13/41 (31%), Positives = 28/41 (67%)
Frame = +1
Query: 19 EFNPAAQVTNNKKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
E +P+ +++EE++E+EEE+EEE E ++ + ++ +E
Sbjct: 178 EIDPSLDEEEEEEEEEEEEEEEEEEEEIEEEEEEEEEEVEE 300
Score = 31.2 bits (69), Expect = 1.1
Identities = 12/38 (31%), Positives = 25/38 (65%)
Frame = +1
Query: 22 PAAQVTNNKKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
P + V N+ + + ++EEE+EEEE E ++ + +++ E
Sbjct: 151 PDSPVANDDEIDPSLDEEEEEEEEEEEEEEEEEEEEIE 264
Score = 30.8 bits (68), Expect = 1.4
Identities = 12/30 (40%), Positives = 23/30 (76%)
Frame = +1
Query: 30 KKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
+++EE+ E+EEE+EEEE+E + + + + E
Sbjct: 241 EEEEEEIEEEEEEEEEEVEEEVEEEEVEGE 330
Score = 30.4 bits (67), Expect = 1.9
Identities = 12/30 (40%), Positives = 23/30 (76%)
Frame = +1
Query: 30 KKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
+++EE++E+EEE EEEE E ++ ++ +E
Sbjct: 223 EEEEEEEEEEEEIEEEEEEEEEEVEEEVEE 312
Score = 30.0 bits (66), Expect = 2.4
Identities = 11/30 (36%), Positives = 24/30 (79%)
Frame = +1
Query: 30 KKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
+++EE++E+EEE+ EEE E ++ + +++ E
Sbjct: 220 EEEEEEEEEEEEEIEEEEEEEEEEVEEEVE 309
Score = 29.6 bits (65), Expect = 3.2
Identities = 12/30 (40%), Positives = 23/30 (76%)
Frame = +1
Query: 30 KKKEEQQEQEEEDEEEELESDDNDNDDDDE 59
+++EE++E EEE+EEEE E ++ +++ E
Sbjct: 235 EEEEEEEEIEEEEEEEEEEVEEEVEEEEVE 324
Score = 29.3 bits (64), Expect = 4.1
Identities = 10/24 (41%), Positives = 20/24 (82%)
Frame = +1
Query: 33 EEQQEQEEEDEEEELESDDNDNDD 56
EE++E+EEE+ EEE+E ++ + ++
Sbjct: 262 EEEEEEEEEEVEEEVEEEEVEGEE 333
Score = 28.5 bits (62), Expect = 7.0
Identities = 11/22 (50%), Positives = 19/22 (86%)
Frame = +1
Query: 30 KKKEEQQEQEEEDEEEELESDD 51
+++EE++E EEE EEEE+E ++
Sbjct: 268 EEEEEEEEVEEEVEEEEVEGEE 333
>TC225160 UP|Q948X9 (Q948X9) Beta-conglycinin alpha-subunit, complete
Length = 2152
Score = 37.0 bits (84), Expect = 0.020
Identities = 15/54 (27%), Positives = 32/54 (58%)
Frame = +3
Query: 22 PAAQVTNNKKKEEQQEQEEEDEEEELESDDNDNDDDDEIYSSNNASRQFTKGSN 75
P + + K+ EE++++E+EDEE++ ES+++++ + + F GSN
Sbjct: 570 PFPRPPHQKESEERKQEEDEDEEQQRESEESESSESQRELRRHKNKNPFLFGSN 731
>TC218919 homologue to UP|Q39892 (Q39892) Nucleosome assembly protein 1,
partial (33%)
Length = 729
Score = 36.6 bits (83), Expect = 0.026
Identities = 12/26 (46%), Positives = 22/26 (84%)
Frame = +1
Query: 33 EEQQEQEEEDEEEELESDDNDNDDDD 58
E+++E E+EDE+EE + D++D ++DD
Sbjct: 199 EDEEEDEDEDEDEEYDEDEDDEEEDD 276
Score = 32.0 bits (71), Expect = 0.64
Identities = 14/26 (53%), Positives = 19/26 (72%)
Frame = +1
Query: 34 EQQEQEEEDEEEELESDDNDNDDDDE 59
E E EEEDE+E+ E ++ D D+DDE
Sbjct: 190 EDDEDEEEDEDED-EDEEYDEDEDDE 264
Score = 28.1 bits (61), Expect = 9.2
Identities = 10/27 (37%), Positives = 20/27 (74%)
Frame = +1
Query: 33 EEQQEQEEEDEEEELESDDNDNDDDDE 59
E + +++EDEEE+ + D+++ D+DE
Sbjct: 175 EFEDLEDDEDEEEDEDEDEDEEYDEDE 255
>TC204850 similar to GB|AAM16178.1|20334844|AY094022 AT3g12390/T2E22_130
{Arabidopsis thaliana;} , partial (85%)
Length = 878
Score = 36.6 bits (83), Expect = 0.026
Identities = 32/146 (21%), Positives = 69/146 (46%), Gaps = 2/146 (1%)
Frame = +1
Query: 24 AQVTNNKKKEEQQEQEEEDEEEELESDDNDNDDDDEIYSSNNASRQFTKGSNNNHQNQFA 83
AQ +K + + E+D++EE++ DD+D DDD + +AS + +K + + +++ A
Sbjct: 82 AQHLEQQKIHDDEPVVEDDDDEEVDDDDDDEDDDHDEGLEGDASGR-SKQTRSEKKSRKA 258
Query: 84 ILDIVMAALKKSIVTCSVEREDVSSLDISWPTEVRHVSHVTFDRFN--GFLGLPSEFQPE 141
+L + M + + +V++ IS P + + T+ F L S+ Q +
Sbjct: 259 MLKLGMKPV-TGVSRVTVKKSKNILFVISKPDVFKSPTSDTYIIFGEAKIEDLSSQLQTQ 435
Query: 142 VPTRVPSASVKVFGVSAKSMQCSYDD 167
+ + ++ G+ +S + DD
Sbjct: 436 AAEQFKAPNLSNDGLKPESSAVAQDD 513
>BU761594 similar to GP|22654971|gb At1g80930/F23A5_23 {Arabidopsis
thaliana}, partial (13%)
Length = 397
Score = 36.2 bits (82), Expect = 0.034
Identities = 17/49 (34%), Positives = 30/49 (60%), Gaps = 5/49 (10%)
Frame = +1
Query: 16 SVSEFNPAAQVTNNKKKEEQQE-----QEEEDEEEELESDDNDNDDDDE 59
S+ F P N+K+ E+ + +E ED+EE L+S+ +D+DD+D+
Sbjct: 106 SLDIFKPDPNFLENEKRYEELKKSMLGEESEDDEEGLDSESDDDDDEDD 252
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.314 0.129 0.369
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 19,155,035
Number of Sequences: 63676
Number of extensions: 246781
Number of successful extensions: 3532
Number of sequences better than 10.0: 203
Number of HSP's better than 10.0 without gapping: 1934
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2736
length of query: 486
length of database: 12,639,632
effective HSP length: 101
effective length of query: 385
effective length of database: 6,208,356
effective search space: 2390217060
effective search space used: 2390217060
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 61 (28.1 bits)
Medicago: description of AC144592.1


