

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
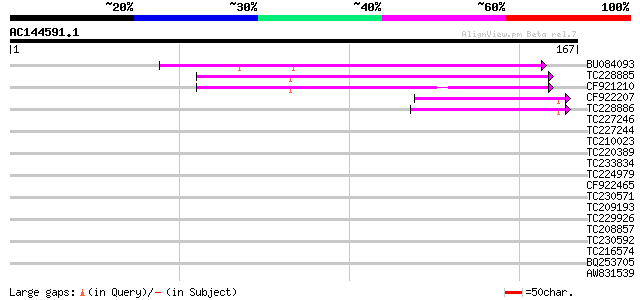
Query= AC144591.1 - phase: 0
(167 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BU084093 65 2e-11
TC228885 55 1e-08
CF921210 51 2e-07
CF922207 40 5e-04
TC228886 40 7e-04
TC227246 similar to UP|Q941A9 (Q941A9) At1g26300/F28B23_4, parti... 36 0.008
TC227244 similar to UP|Q941A9 (Q941A9) At1g26300/F28B23_4, parti... 35 0.013
TC210023 weakly similar to UP|Q868B4 (Q868B4) C. elegans GRL-25 ... 34 0.037
TC220389 33 0.064
TC233834 similar to UP|Q9FGZ0 (Q9FGZ0) Arabidopsis thaliana geno... 32 0.11
TC224979 similar to UP|Q09083 (Q09083) Hydroxyproline-rich glyco... 32 0.11
CF922465 32 0.14
TC230571 similar to UP|Q41645 (Q41645) Extensin (Fragment), part... 32 0.14
TC209193 weakly similar to UP|O64640 (O64640) Expressed protein ... 32 0.19
TC229926 similar to UP|Q93VB2 (Q93VB2) AT3g62770/F26K9_200, part... 32 0.19
TC208857 weakly similar to UP|Q84KG8 (Q84KG8) Fatty acid delta-6... 31 0.24
TC230592 similar to UP|Q902U4 (Q902U4) Rev protein, partial (16%) 31 0.24
TC216574 similar to UP|RK4_SPIOL (O49937) 50S ribosomal protein ... 31 0.32
BQ253705 weakly similar to GP|27817931|db P0453E05.6 {Oryza sati... 31 0.32
AW831539 31 0.32
>BU084093
Length = 421
Score = 64.7 bits (156), Expect = 2e-11
Identities = 40/121 (33%), Positives = 56/121 (46%), Gaps = 7/121 (5%)
Frame = -1
Query: 45 NRVSSLMVTKDLPQFQQRPQPP-----RQPSFTQQPRQQAPRT--RFDPIPMKYAEFLPS 97
N SS V P ++ PQ P R + R PR F P+PM Y + LPS
Sbjct: 364 NASSSTPVQPKAPAQREAPQVPTPNTTRPVGNSNTTRNFPPRPFQEFTPLPMTYEDLLPS 185
Query: 98 LLKGNYVQTRPPPPRPEKLPAGFRADLSCVFHQGAPGHDVERCYALRNAVQDLVEAKILS 157
L+ + P P + + +C +H G PGH VE+C AL+ VQ L++A L+
Sbjct: 184 LIANHLAVVTPGRVLQPPFPKWYDPNATCKYHGGVPGHSVEKCLALKYKVQHLMDAGWLT 5
Query: 158 F 158
F
Sbjct: 4 F 2
>TC228885
Length = 901
Score = 55.5 bits (132), Expect = 1e-08
Identities = 36/107 (33%), Positives = 47/107 (43%), Gaps = 2/107 (1%)
Frame = -3
Query: 56 LPQFQQRPQPPRQPSFTQQPRQQAPR--TRFDPIPMKYAEFLPSLLKGNYVQTRPPPPRP 113
LP FQ R P Q T Q A R F PIP+ YA+ LP LL + V
Sbjct: 707 LPHFQ*RTPPLAQTKNTNQEMNFAARKPVEFTPIPVSYADLLPYLLDNSMVAITLAKVHQ 528
Query: 114 EKLPAGFRADLSCVFHQGAPGHDVERCYALRNAVQDLVEAKILSFTD 160
+ ++ C H APG +E AL+ VQ L++A L F +
Sbjct: 527 PPFLREYDSNAMCACHGEAPGRSIEHYRALKRKVQGLIDAGWLKFEE 387
>CF921210
Length = 790
Score = 51.2 bits (121), Expect = 2e-07
Identities = 37/107 (34%), Positives = 49/107 (45%), Gaps = 2/107 (1%)
Frame = +2
Query: 56 LPQFQQRPQPPRQPSFTQQPRQQAPR--TRFDPIPMKYAEFLPSLLKGNYVQTRPPPPRP 113
LP FQ R P Q T Q A R F PIP+ YA+ L LL + V
Sbjct: 251 LPHFQ*RTPPLAQTKNTNQEVNFAARKPVEFTPIPVSYADLLSYLLDNSMVAITLAKVHQ 430
Query: 114 EKLPAGFRADLSCVFHQGAPGHDVERCYALRNAVQDLVEAKILSFTD 160
L G+ ++ +C GA H +E C AL+ VQ L++A L F +
Sbjct: 431 PPLF*GYDSNATC---GGALRHSIEHCRALKRKVQGLIDAGWLKFEE 562
>CF922207
Length = 616
Score = 40.0 bits (92), Expect = 5e-04
Identities = 17/49 (34%), Positives = 29/49 (58%), Gaps = 3/49 (6%)
Frame = -1
Query: 120 FRADLSCVFHQGAPGHDVERCYALRNAVQDLVEAKILSFTD---LNPNL 165
+ ++ +C +H GAPGH +E C ++ V L++ L F + LNPN+
Sbjct: 454 YDSNATCAYHGGAPGHSIEHCMTPKHKV*SLIDTG*LKFEENRLLNPNI 308
>TC228886
Length = 748
Score = 39.7 bits (91), Expect = 7e-04
Identities = 17/50 (34%), Positives = 29/50 (58%), Gaps = 3/50 (6%)
Frame = +3
Query: 119 GFRADLSCVFHQGAPGHDVERCYALRNAVQDLVEAKILSFTD---LNPNL 165
G+ ++ +C +H GA GH +E C ++ V L++ L F + LNPN+
Sbjct: 150 GYDSNATCAYHGGASGHSIEHCMTPKHKV*SLIDTGWLKFEENRLLNPNI 299
>TC227246 similar to UP|Q941A9 (Q941A9) At1g26300/F28B23_4, partial (41%)
Length = 577
Score = 36.2 bits (82), Expect = 0.008
Identities = 21/51 (41%), Positives = 30/51 (58%)
Frame = +3
Query: 63 PQPPRQPSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKGNYVQTRPPPPRP 113
P+PPR+P+ + PR A +TR P P + ++FLP+ PPPPRP
Sbjct: 210 PKPPRRPTSSNPPR-FARQTRSSPSPTQ-SQFLPNSPPLPPPPLPPPPPRP 356
>TC227244 similar to UP|Q941A9 (Q941A9) At1g26300/F28B23_4, partial (67%)
Length = 1408
Score = 35.4 bits (80), Expect = 0.013
Identities = 22/55 (40%), Positives = 28/55 (50%)
Frame = +1
Query: 63 PQPPRQPSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKGNYVQTRPPPPRPEKLP 117
P+PPR+P+ + PR P TR P P + FLP+ PPPP P LP
Sbjct: 244 PKPPRRPTSSNPPRSARP-TRSSPSPTP-SPFLPN------SPPLPPPPPPLPLP 384
>TC210023 weakly similar to UP|Q868B4 (Q868B4) C. elegans GRL-25 protein
(Corresponding sequence ZK643.8), partial (12%)
Length = 819
Score = 33.9 bits (76), Expect = 0.037
Identities = 29/99 (29%), Positives = 42/99 (42%), Gaps = 1/99 (1%)
Frame = -3
Query: 61 QRPQPPRQPSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKGNYVQTRPPPPRPEKLPAGF 120
QRPQPP F Q P Q AP+ P+ +K+ + PPP+P P
Sbjct: 247 QRPQPPLLIKFPQPPPQPAPQPPQPPLLIKFPQ---------------PPPQPAPQP--- 122
Query: 121 RADLSCVFHQGAPGHDVERCYAL-RNAVQDLVEAKILSF 158
LS F Q P ++ Y + +++ LVE + F
Sbjct: 121 --PLSTAFPQ--PPMEISDLYVIPSSSLHILVELNPVPF 17
>TC220389
Length = 829
Score = 33.1 bits (74), Expect = 0.064
Identities = 27/78 (34%), Positives = 37/78 (46%), Gaps = 10/78 (12%)
Frame = +1
Query: 58 QFQQRPQPPRQPSFTQQPRQQ------APRTRFDPIPMKYAEF--LPSLLKGNYV--QTR 107
Q+ +R +QP F +Q Q A + R D IP AE+ LP + V +
Sbjct: 358 QYLERSASAQQPGFGEQVMHQLYHVKNATQLRMDDIPKLLAEYKQLPI*M---LV**K*S 528
Query: 108 PPPPRPEKLPAGFRADLS 125
PPPP P+K P F +LS
Sbjct: 529 PPPPSPQKNPKEFHGELS 582
>TC233834 similar to UP|Q9FGZ0 (Q9FGZ0) Arabidopsis thaliana genomic DNA,
chromosome 5, TAC clone:K6M13, partial (18%)
Length = 432
Score = 32.3 bits (72), Expect = 0.11
Identities = 22/77 (28%), Positives = 35/77 (44%)
Frame = +1
Query: 5 NLSAHMQTLAQTHKMDQLEQELHELRKEMTALRAKVEELTNRVSSLMVTKDLPQFQQRPQ 64
++SA + A H + Q H LR + + + + + + LP+ RP
Sbjct: 112 SVSATLPAAAPNHHLLQH----HPLRSIQSTVAIRQLPSRGPLLPALARRHLPRLPPRP- 276
Query: 65 PPRQPSFTQQPRQQAPR 81
PPR+PS Q P +APR
Sbjct: 277 PPRKPSLRQPPLSRAPR 327
>TC224979 similar to UP|Q09083 (Q09083) Hydroxyproline-rich glycoprotein
precursor, partial (32%)
Length = 693
Score = 32.3 bits (72), Expect = 0.11
Identities = 18/53 (33%), Positives = 23/53 (42%)
Frame = +1
Query: 63 PQPPRQPSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKGNYVQTRPPPPRPEK 115
P PP++P + P +P P P Y P K Y PPPP P+K
Sbjct: 325 PPPPKKPYYYHSPPPPSPPP---PKPYYYHSPPPPPKKKPYYYHSPPPPPPKK 474
>CF922465
Length = 538
Score = 32.0 bits (71), Expect = 0.14
Identities = 12/42 (28%), Positives = 23/42 (54%)
Frame = -2
Query: 119 GFRADLSCVFHQGAPGHDVERCYALRNAVQDLVEAKILSFTD 160
G+ ++ +C H+ + GH +E C + VQ ++A L F +
Sbjct: 465 GYDSNATCACHEESLGHSIEHCMTSKRKVQGPIDAGWLKFEE 340
>TC230571 similar to UP|Q41645 (Q41645) Extensin (Fragment), partial (7%)
Length = 667
Score = 32.0 bits (71), Expect = 0.14
Identities = 26/69 (37%), Positives = 30/69 (42%), Gaps = 4/69 (5%)
Frame = +2
Query: 53 TKDLPQFQQRPQPPRQPSFTQQPRQ----QAPRTRFDPIPMKYAEFLPSLLKGNYVQTRP 108
T LP QR PP P +PR PR R P P Y P+ L+ TRP
Sbjct: 239 TPPLPP*PQRFHPPPLPGHPPRPRAFPRPLPPRPRRRPRPPLY----PARLR-----TRP 391
Query: 109 PPPRPEKLP 117
PRP +LP
Sbjct: 392 RLPRPHQLP 418
>TC209193 weakly similar to UP|O64640 (O64640) Expressed protein
(At2g45600/F17K2.13), partial (34%)
Length = 1145
Score = 31.6 bits (70), Expect = 0.19
Identities = 27/76 (35%), Positives = 32/76 (41%), Gaps = 5/76 (6%)
Frame = +1
Query: 46 RVSSLMVTKDLPQFQQRPQP--PRQPS---FTQQPRQQAPRTRFDPIPMKYAEFLPSLLK 100
R SSL P Q P P P+ PS P PR R P+P LP LL+
Sbjct: 28 RTSSLKRYPSQPNHQHLPPPLPPKPPSPLCRQTPPHNILPRRRLHPLP-PLLPHLPPLLR 204
Query: 101 GNYVQTRPPPPRPEKL 116
+ RP PPR +L
Sbjct: 205 ----RPRPLPPRHHRL 240
>TC229926 similar to UP|Q93VB2 (Q93VB2) AT3g62770/F26K9_200, partial (46%)
Length = 829
Score = 31.6 bits (70), Expect = 0.19
Identities = 22/59 (37%), Positives = 28/59 (47%)
Frame = +2
Query: 57 PQFQQRPQPPRQPSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKGNYVQTRPPPPRPEK 115
P+ ++R PR P P QQ P+P+ LP L G Q RPPPPRP +
Sbjct: 407 PRLRRRRLLPRPPL----PSQQGHDLGRPPVPLHRRALLP--LGG---QGRPPPPRPNR 556
>TC208857 weakly similar to UP|Q84KG8 (Q84KG8) Fatty acid delta-6 desaturase,
partial (21%)
Length = 648
Score = 31.2 bits (69), Expect = 0.24
Identities = 22/64 (34%), Positives = 26/64 (40%)
Frame = +2
Query: 54 KDLPQFQQRPQPPRQPSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKGNYVQTRPPPPRP 113
+DL + P PPR+P T PR R R P P PS L + PP P P
Sbjct: 125 QDLRRLLLAPPPPRRPPPTPHPR----RERTPPTPSS-----PSTLPTPPPSSSPPSPPP 277
Query: 114 EKLP 117
P
Sbjct: 278 TVFP 289
>TC230592 similar to UP|Q902U4 (Q902U4) Rev protein, partial (16%)
Length = 1282
Score = 31.2 bits (69), Expect = 0.24
Identities = 19/61 (31%), Positives = 29/61 (47%)
Frame = +1
Query: 53 TKDLPQFQQRPQPPRQPSFTQQPRQQAPRTRFDPIPMKYAEFLPSLLKGNYVQTRPPPPR 112
+ D + QQ+P PP+ SF+ P Q + F +P + SL Q+ PPP +
Sbjct: 4 SSDEEEDQQQPSPPKTTSFSSLP--QPKSSLFQSLPQPKSSPFSSLF-----QSLPPPKQ 162
Query: 113 P 113
P
Sbjct: 163 P 165
>TC216574 similar to UP|RK4_SPIOL (O49937) 50S ribosomal protein L4,
chloroplast precursor (R-protein L4), partial (74%)
Length = 1364
Score = 30.8 bits (68), Expect = 0.32
Identities = 24/88 (27%), Positives = 36/88 (40%), Gaps = 5/88 (5%)
Frame = +2
Query: 57 PQFQQRPQPPRQPSFTQ-QPRQQAPRTRFDPIPMKYAEFLPSLLKGNYVQTRPPPPRPEK 115
P R +P R+P + +P +QAPR R P P + L ++RPP +P
Sbjct: 293 PLRPSRHRPSRRPPRRRHRPPEQAPRHRLHPHPRRGPRRRTEALPAEKNRSRPPRLQPHA 472
Query: 116 LP----AGFRADLSCVFHQGAPGHDVER 139
P R + + HQ P + R
Sbjct: 473 PPPRRRRHLRPEAPRLVHQDQPQGEAPR 556
>BQ253705 weakly similar to GP|27817931|db P0453E05.6 {Oryza sativa (japonica
cultivar-group)}, partial (2%)
Length = 403
Score = 30.8 bits (68), Expect = 0.32
Identities = 30/116 (25%), Positives = 46/116 (38%), Gaps = 4/116 (3%)
Frame = +2
Query: 5 NLSAHMQTLAQTHKMDQLEQELHELRKEMTALRAKVEELTNRVSSLMVTKDLPQFQQRPQ 64
NL ++ A L++E+ E+ + R EEL + S++ + + P
Sbjct: 47 NLMRSLKKTASDLGSGNLKREVSEISERPLHSRCNSEELVDCSDSVLSGSVRSRAPRVPN 226
Query: 65 PPRQPSFTQQPRQ----QAPRTRFDPIPMKYAEFLPSLLKGNYVQTRPPPPRPEKL 116
PP +PS T P + R P P + LP PPPP P+KL
Sbjct: 227 PPPKPS-TSSPSSGSSGETERRIPPPPPPPPMKALPP----------PPPPPPKKL 361
>AW831539
Length = 397
Score = 30.8 bits (68), Expect = 0.32
Identities = 23/64 (35%), Positives = 30/64 (45%), Gaps = 7/64 (10%)
Frame = +1
Query: 55 DLPQFQQRPQPPRQ----PSFTQQPR---QQAPRTRFDPIPMKYAEFLPSLLKGNYVQTR 107
D PQ Q Q PRQ P QPR QQ P R + ++ + LL+G+ Q +
Sbjct: 118 DAPQ-QHDAQTPRQSHRQPHVPPQPRCLRQQPPFLRRQGVRVQLQQHSKQLLRGHCQQQQ 294
Query: 108 PPPP 111
PPP
Sbjct: 295 APPP 306
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.318 0.132 0.388
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,250,237
Number of Sequences: 63676
Number of extensions: 119089
Number of successful extensions: 1500
Number of sequences better than 10.0: 144
Number of HSP's better than 10.0 without gapping: 1332
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1448
length of query: 167
length of database: 12,639,632
effective HSP length: 90
effective length of query: 77
effective length of database: 6,908,792
effective search space: 531976984
effective search space used: 531976984
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 55 (25.8 bits)
Medicago: description of AC144591.1


