

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144538.7 + phase: 1 /pseudo
(632 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
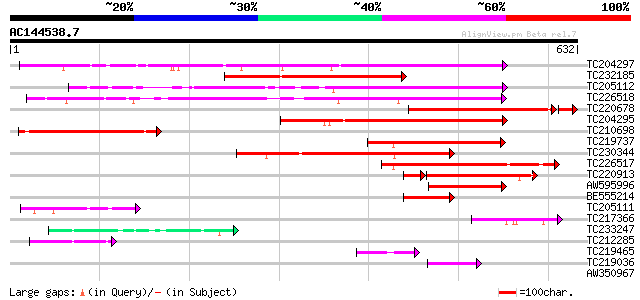
Score E
Sequences producing significant alignments: (bits) Value
TC204297 similar to UP|Q9LJE5 (Q9LJE5) Gb|AAD10646.1 (AT3g13460/... 398 e-111
TC232185 similar to GB|BAC75841.1|30060380|AP003799 RNA-binding ... 338 4e-93
TC205112 similar to UP|Q93ZP1 (Q93ZP1) AT5g61020/maf19_20, parti... 321 7e-88
TC226518 similar to UP|YTH2_HUMAN (Q9Y5A9) YTH domain protein 2 ... 306 2e-83
TC220678 similar to UP|Q9LDZ8 (Q9LDZ8) Gb|AAF35955.1, partial (27%) 297 4e-82
TC204295 similar to UP|Q9LJE5 (Q9LJE5) Gb|AAD10646.1 (AT3g13460/... 288 4e-78
TC210698 similar to UP|Q9LDZ8 (Q9LDZ8) Gb|AAF35955.1, partial (7%) 271 8e-73
TC219737 similar to UP|Q9LNG4 (Q9LNG4) F21D18.17, partial (23%) 233 2e-61
TC230344 similar to UP|Q9LNG4 (Q9LNG4) F21D18.17, partial (20%) 216 3e-56
TC226517 similar to UP|YTH2_HUMAN (Q9Y5A9) YTH domain protein 2 ... 190 2e-48
TC220913 similar to UP|YTH2_HUMAN (Q9Y5A9) YTH domain protein 2 ... 129 6e-34
AW595996 116 3e-26
BE555214 104 1e-22
TC205111 similar to UP|Q6IH72 (Q6IH72) HDC03105, partial (19%) 80 4e-15
TC217366 weakly similar to UP|Q9LUT8 (Q9LUT8) Arabidopsis thalia... 56 6e-08
TC233247 weakly similar to UP|Q9LUT8 (Q9LUT8) Arabidopsis thalia... 55 7e-08
TC212285 similar to UP|Q9LNG4 (Q9LNG4) F21D18.17, partial (13%) 46 4e-05
TC219465 similar to GB|AAH53863.1|31808095|BC053863 YT521 protei... 45 1e-04
TC219036 44 3e-04
AW350967 similar to GP|15912287|gb| AT5g61020/maf19_20 {Arabidop... 41 0.001
>TC204297 similar to UP|Q9LJE5 (Q9LJE5) Gb|AAD10646.1 (AT3g13460/MRP15_10),
partial (48%)
Length = 2995
Score = 398 bits (1023), Expect = e-111
Identities = 243/606 (40%), Positives = 340/606 (56%), Gaps = 63/606 (10%)
Frame = +2
Query: 12 IPESSKFSNSSQ-GTTTIGHSRD--ITSQSGSFGSGADQPLYPPNVYAPQ---AQAFYYG 65
IPE +K + +Q G+ G++ + I S S + P Y P + A+YYG
Sbjct: 296 IPEPTKKATGNQYGSVDSGNAANGQIQSYDRSVTPVLQDFIDPTMCYLPNGYPSTAYYYG 475
Query: 66 GFDNGNGEWDEYPSYVNNGGIEIGSPGVYNENQSLVFHSGYGFNPQMPYGPYSPVTSPLP 125
G+D EWDEY YVN+ G+E+ S GVY +N SL++H GYG+ PYGPYSP SP+P
Sbjct: 476 GYDGTGNEWDEYSRYVNSEGVEMTS-GVYGDNGSLLYHHGYGY---APYGPYSPAGSPVP 643
Query: 126 SVGGDAQLYSPQQFPYTPPYYNQLVPPPNLPYLNSPTPVSQPELTNLLGIDQQ---VES- 181
++G D QLY PQ + Y PPY+ L P + P+ +P + Q E++ + DQ+ VE+
Sbjct: 644 TMGNDGQLYGPQHYQY-PPYFQPLTPT-SAPFTPTPAVLPQGEVSTSVAADQKPLPVEAA 817
Query: 182 ----------------NFFGP------RASYPSVGSFARGSFPVAPGSFSFHESQQGFDG 219
N P +S+ S S R + P + + + + G+DG
Sbjct: 818 NGNSNGVSNGGNAKGNNAAAPIKQANQNSSFSSKASNERVAMPGRGPTSGYQDPRFGYDG 997
Query: 220 SRSGGLWSDSSKPSERQRSFMPLSPSVPQQPIGSVRS-------FGPSAG-MASHQQRSL 271
RS W D+ S+ Q P+S + I + F P++ M H R +
Sbjct: 998 VRSPIPWLDAPLFSDGQPR--PVSSTTITSSISGGNNTASRNPTFRPNSQFMGLHHPRPM 1171
Query: 272 YGFGSSSNPYGRGYLSNPGSSFGGSAISGLS---------ANDRSFLSLENS-RRHGRET 321
G++ + R Y S +G + SG+ N R++L++++ + GR
Sbjct: 1172PAMGATHSFINRMYPSKLYGQYGNTVRSGMGYGTHGYDSRTNGRAWLAVDSKYKTRGRSG 1351
Query: 322 ASFCRCNGTLDILSEQNRGPRASKLKNHISSENNAI-------------DGSKNNASTAK 368
F N D L+E NRGPRA KN + D K+ ST
Sbjct: 1352GYFGYGNENADGLNELNRGPRAKGGKNQKGFAPTILAVKGQTLPATLGTDEEKDKTSTI- 1528
Query: 369 FQDESLNRPDFATDFKDAKFFVIKSYSEDNVHKSIKYGVWASTPNGNRKLDAAYCQAKEK 428
+ + N+ DF ++ DAKFFVIKSYSED++HKSIKY VWAST NGN+KLDAAY +A++K
Sbjct: 1529LECDQYNKADFPEEYTDAKFFVIKSYSEDDIHKSIKYNVWASTQNGNKKLDAAYQEAQQK 1708
Query: 429 QDACRIFLFFSVNASAQFCGVAEMVGPVNFDKSVDFWQQDKWSGQFPVKWHIIKDVPNSQ 488
+FLFFSVN S QF G+AEM+GPV+F+KSV++WQQDKW+G FP+KWHI+KDVPN+
Sbjct: 1709PGGTPVFLFFSVNTSGQFVGLAEMIGPVDFNKSVEYWQQDKWNGCFPLKWHIVKDVPNNL 1888
Query: 489 FRHIVLENNDNKPVTNSRDTQEVKLQQGIEMLTIFKNYETDTSILDDFDFYEDRQKAMQE 548
RHI L+NN+NKPVTNSRDTQEV L+ G++++ IFK Y + T ILDDF FYE RQK + E
Sbjct: 1889LRHITLDNNENKPVTNSRDTQEVMLEPGLKLIKIFKEYTSKTCILDDFGFYEARQKTILE 2068
Query: 549 RKARQQ 554
+KA+QQ
Sbjct: 2069KKAKQQ 2086
>TC232185 similar to GB|BAC75841.1|30060380|AP003799 RNA-binding protein-like
contains ESTs
AU101986(S3844),AU068423(C30241),AU068424(C30241),
D41369(S3844) {Oryza sativa (japonica cultivar-group);}
, partial (15%)
Length = 610
Score = 338 bits (867), Expect = 4e-93
Identities = 169/203 (83%), Positives = 181/203 (88%)
Frame = +3
Query: 240 MPLSPSVPQQPIGSVRSFGPSAGMASHQQRSLYGFGSSSNPYGRGYLSNPGSSFGGSAIS 299
MPLSPSV QP+GS+ SFGPS GMASHQQ+SLYGFGS+SN YGRGYL N GSSFGG++IS
Sbjct: 6 MPLSPSVSPQPMGSLGSFGPSVGMASHQQQSLYGFGSASNSYGRGYLPNQGSSFGGTSIS 185
Query: 300 GLSANDRSFLSLENSRRHGRETASFCRCNGTLDILSEQNRGPRASKLKNHISSENNAIDG 359
L NDRSF SLENSRR GR TAS C CNGTLDILSEQNRGPRASKLKN IS+ENN++D
Sbjct: 186 NL--NDRSFASLENSRRQGRPTASLCNCNGTLDILSEQNRGPRASKLKNQISTENNSVDS 359
Query: 360 SKNNASTAKFQDESLNRPDFATDFKDAKFFVIKSYSEDNVHKSIKYGVWASTPNGNRKLD 419
SKN+ASTAKFQ+ESLNR DFATD+KDAKFFVIKSYSEDNVHKSIKYGVWASTPNGNRKLD
Sbjct: 360 SKNSASTAKFQNESLNRSDFATDYKDAKFFVIKSYSEDNVHKSIKYGVWASTPNGNRKLD 539
Query: 420 AAYCQAKEKQDACRIFLFFSVNA 442
AY QA EKQDAC IFLFFSVNA
Sbjct: 540 DAYRQAMEKQDACPIFLFFSVNA 608
>TC205112 similar to UP|Q93ZP1 (Q93ZP1) AT5g61020/maf19_20, partial (38%)
Length = 1757
Score = 321 bits (822), Expect = 7e-88
Identities = 202/506 (39%), Positives = 266/506 (51%), Gaps = 17/506 (3%)
Frame = +1
Query: 66 GFDNGNGEWDEYPSYVN-NGGIEIGSPGVYNENQSLVFHSGYGFNPQMPYGPYSPVTSPL 124
G+D G G+W+ Y Y+N +GG+ + GVY ++ S ++H GYG+ P YG Y+P S
Sbjct: 46 GYD-GQGDWNAYSRYMNLDGGM---AQGVYGDSCSYMYHQGYGYTP---YGTYAPPNSSS 204
Query: 125 PSVGGDAQLYSPQQFPYTPPYYNQLVPPPNLPYLNSPTPVSQPELTNLLGIDQQVESNFF 184
P + D Q Y QQ Y P Y L S +
Sbjct: 205 PMIQQDGQHYGLQQ--YHLPAYASL-------------------------------SGYQ 285
Query: 185 GPRASYPSVGSFARGSFPVAPGSFSFHESQQGFDGSRSGGLWSDSSKPSERQRSFMPLSP 244
GPR+++ + PV P S +Q G++ G S + L
Sbjct: 286 GPRSTHGT-------QLPV-PSDVSLVSDRQSKHGAKVGLSSSVVPVKDFTSQRNQRLPQ 441
Query: 245 SVPQQPIGSVRSFGPSAGMASHQQRSLYGFGSSSNPYGRGYLSNPGSSFGGSAISGLSAN 304
+PQ S + S +Y + YG + P S FG + G +
Sbjct: 442 PLPQYVSMSGSRHPSGLDLVSGFMNGMYPSNRMYSQYGNTF--RPDSRFGSA---GYGSR 606
Query: 305 DRSFLSLENSRRHGRETASFCRCNGTLDILSEQNRGPRASKLKNHISSENNAIDG----- 359
SF S N +G C ++D SE N+GPRA+K SS+N I
Sbjct: 607 MGSFDSKFNGTGYG------CGLKKSMDGFSELNKGPRAAK-----SSDNKNIKSLGPVT 753
Query: 360 -----------SKNNASTAKFQDESLNRPDFATDFKDAKFFVIKSYSEDNVHKSIKYGVW 408
S N E N DFA ++ DAKFFVIKSYSED++HKSIKY W
Sbjct: 754 LLLKGQNLPVKSDNKEVPPVPDKEQYNGKDFAENYSDAKFFVIKSYSEDDIHKSIKYSAW 933
Query: 409 ASTPNGNRKLDAAYCQAKEKQDACRIFLFFSVNASAQFCGVAEMVGPVNFDKSVDFWQQD 468
ASTPNGN+KLDAAY +AKEK C IFL FSVN S QF G+AEM+GPV+F K+VD+WQQD
Sbjct: 934 ASTPNGNKKLDAAYQEAKEKPGGCPIFLLFSVNTSGQFVGLAEMLGPVDFGKTVDYWQQD 1113
Query: 469 KWSGQFPVKWHIIKDVPNSQFRHIVLENNDNKPVTNSRDTQEVKLQQGIEMLTIFKNYET 528
+W+G F VKWH+IKD+PNS RHI LENN+NKPVTNSRDTQEVK ++G+++ IFK + +
Sbjct: 1114RWTGCFSVKWHVIKDIPNSVLRHITLENNENKPVTNSRDTQEVKFEKGVQIAKIFKEHSS 1293
Query: 529 DTSILDDFDFYEDRQKAMQERKARQQ 554
T ILDDF FYE R+KA QE+K+++Q
Sbjct: 1294QTCILDDFGFYEAREKATQEKKSKEQ 1371
>TC226518 similar to UP|YTH2_HUMAN (Q9Y5A9) YTH domain protein 2
(High-glucose-regulated protein 8) (NY-REN-2 antigen)
(CLL-associated antigen KW-14), partial (20%)
Length = 2656
Score = 306 bits (784), Expect = 2e-83
Identities = 213/561 (37%), Positives = 281/561 (49%), Gaps = 26/561 (4%)
Frame = +3
Query: 19 SNS--SQGTTTIGHSRDITSQSGSFGSGADQPLYPPNVYAPQAQA----------FYYGG 66
SNS S+G ++ SR S G SG+ + + + Y Q +YY G
Sbjct: 462 SNSVPSKGASSPSDSRSCVSSIGD-ASGSVKEVDVDHEYLSTDQGVPYPAGGYYGYYYPG 638
Query: 67 FDNGNGEWDEYPSYVNNGGIEIGSPGVYNENQSLVFHSGYGFNPQMP--YGPYSPVTSPL 124
+ GE D YV +++ P + +N S V+ GF P Y P SP
Sbjct: 639 YGGFYGESDNQGYYVGADAVDLQYPVMQADNGSYVYLVP-GFQTGYPSYYHPLSPAGIEG 815
Query: 125 PSVGGDAQLYSP----QQFPYTPPYYNQLVPPPNLPYLNSPTPVSQPELTNLLGIDQQVE 180
VG + +Y P QQ +P YY P +S EL Q
Sbjct: 816 QYVGHN--VYPPGSIYQQPIGSPGYY--------------PASLSYGEL-------QPTT 926
Query: 181 SNFFGPRASYPSVGSFARGSFPVAPGSFSFHESQQGFDGSRSGGLWSDSSKPSERQRSFM 240
++ P + P S SQ G S +S + S +
Sbjct: 927 YSWDSPLIKQDGLQGHGYNELAGKPNGRSNLSSQSHTSGVVSKSAPPPNSAEGKGLTSLL 1106
Query: 241 PLSPSVPQQPIGSVRSFGPSAGMASHQQR-SLYGFGSSSNPYGRGYLSNPGSSFGGSAIS 299
+S + + + P + + S + Y G S Y L+
Sbjct: 1107EVSSTHVKHNQPKQTNKAPVSVLHSPVAKFPAYNQGKSGFLYPNNLLN------------ 1250
Query: 300 GLSANDRSFLSLENSRRHGRETASFCRCNGTLDILSEQNRGPRASKLKNHISSENNAIDG 359
+ AN + ++S E ++ N D L+EQN+GPR + K + S N++ G
Sbjct: 1251-VKANTKGWVSTEKLKQR----------NKVNDSLNEQNQGPRTANAKGALMSGGNSVRG 1397
Query: 360 SKNNAS---TAKFQDESLNRPDFATDFKDAKFFVIKSYSEDNVHKSIKYGVWASTPNGNR 416
S S T K + + N PDF T + A FFVIKSYSED++HKSIKY VWASTPNGN+
Sbjct: 1398SAPGGSGNVTNKIRTDQYNLPDFPTKYDHALFFVIKSYSEDDIHKSIKYNVWASTPNGNK 1577
Query: 417 KLDAAYCQAKEKQDA----CRIFLFFSVNASAQFCGVAEMVGPVNFDKSVDFWQQDKWSG 472
+LD A+ A+++ + C +FLFFSVNAS QFCGVAEM G V+F+KS+DFWQQDKW+G
Sbjct: 1578RLDGAFQDAQKRMEEKGCKCPVFLFFSVNASGQFCGVAEMTGRVDFNKSMDFWQQDKWNG 1757
Query: 473 QFPVKWHIIKDVPNSQFRHIVLENNDNKPVTNSRDTQEVKLQQGIEMLTIFKNYETDTSI 532
FPVKWHIIKDVPN Q RHI+LENND+KPVT+SRDTQEV QG+EML IFKNY TSI
Sbjct: 1758YFPVKWHIIKDVPNPQLRHIILENNDHKPVTSSRDTQEVSFPQGVEMLNIFKNYVARTSI 1937
Query: 533 LDDFDFYEDRQKAMQERKARQ 553
LDDF+FYE RQK MQE+K RQ
Sbjct: 1938LDDFEFYESRQKVMQEKKTRQ 2000
>TC220678 similar to UP|Q9LDZ8 (Q9LDZ8) Gb|AAF35955.1, partial (27%)
Length = 966
Score = 297 bits (760), Expect(2) = 4e-82
Identities = 143/165 (86%), Positives = 157/165 (94%)
Frame = +3
Query: 445 QFCGVAEMVGPVNFDKSVDFWQQDKWSGQFPVKWHIIKDVPNSQFRHIVLENNDNKPVTN 504
QFCGVAEMVGPVNFDKSVDFWQQDKWSGQFPVKWHIIKDVPNSQFRHIVLENNDNKPVTN
Sbjct: 3 QFCGVAEMVGPVNFDKSVDFWQQDKWSGQFPVKWHIIKDVPNSQFRHIVLENNDNKPVTN 182
Query: 505 SRDTQEVKLQQGIEMLTIFKNYETDTSILDDFDFYEDRQKAMQERKARQQSSIMPTGLVG 564
SRDTQEVKL QG+EMLTIFKNYETD SILDDFDFYEDRQKAMQERKARQQSS++ TGLVG
Sbjct: 183 SRDTQEVKLTQGVEMLTIFKNYETDVSILDDFDFYEDRQKAMQERKARQQSSMVATGLVG 362
Query: 565 GSEHRSSSDSTGDFIKQVSKNFSLVVRLDDNNNEVIAANRDKLAS 609
+EHRSS+++TGDF+KQ+SK+F+LVVRLD+NN EV+ A+RD L S
Sbjct: 363 ENEHRSSANTTGDFMKQMSKSFALVVRLDENNKEVV-ADRDSLVS 494
Score = 26.9 bits (58), Expect(2) = 4e-82
Identities = 13/21 (61%), Positives = 14/21 (65%)
Frame = +1
Query: 612 PTGNVFKPGEGILVTASSMQT 632
P GNV KP +G TASS QT
Sbjct: 508 PIGNVVKPDDGQSGTASSTQT 570
>TC204295 similar to UP|Q9LJE5 (Q9LJE5) Gb|AAD10646.1 (AT3g13460/MRP15_10),
partial (34%)
Length = 1745
Score = 288 bits (738), Expect = 4e-78
Identities = 144/266 (54%), Positives = 190/266 (71%), Gaps = 14/266 (5%)
Frame = +3
Query: 303 ANDRSFLSLENS-RRHGRETASFCRCNGTLDILSEQNRGPRASKLKNH---------ISS 352
AN R++L++++ + GR F N +D L+E NRGPRA KN +
Sbjct: 30 ANGRAWLAVDSKYKTRGRSGGYFGYGNENVDGLNELNRGPRAKGGKNQKGFAPTILAVKG 209
Query: 353 ENN----AIDGSKNNASTAKFQDESLNRPDFATDFKDAKFFVIKSYSEDNVHKSIKYGVW 408
+N D K+ ST +D+ N+ DF ++ DAKFFVIKSYSED++HKSIKY VW
Sbjct: 210 QNLPASLGTDEEKDKTSTVPDRDQ-YNKADFPEEYTDAKFFVIKSYSEDDIHKSIKYNVW 386
Query: 409 ASTPNGNRKLDAAYCQAKEKQDACRIFLFFSVNASAQFCGVAEMVGPVNFDKSVDFWQQD 468
AST NGN+KLDAAY +A++K C +FLFFSVN S QF G+AEM+GPV+F+KSV++WQQD
Sbjct: 387 ASTQNGNKKLDAAYHEAQQKPGGCPVFLFFSVNTSGQFVGLAEMIGPVDFNKSVEYWQQD 566
Query: 469 KWSGQFPVKWHIIKDVPNSQFRHIVLENNDNKPVTNSRDTQEVKLQQGIEMLTIFKNYET 528
KW+G FP+KWH++KDVPN+ RHI L+NN+NKPVTNSRDTQEV L+ G++++ IFK Y +
Sbjct: 567 KWNGCFPLKWHVVKDVPNNLLRHITLDNNENKPVTNSRDTQEVMLEPGLKLIKIFKEYTS 746
Query: 529 DTSILDDFDFYEDRQKAMQERKARQQ 554
T ILDDF FYE RQK + E+KA+QQ
Sbjct: 747 KTCILDDFGFYEARQKTILEKKAKQQ 824
>TC210698 similar to UP|Q9LDZ8 (Q9LDZ8) Gb|AAF35955.1, partial (7%)
Length = 676
Score = 271 bits (692), Expect = 8e-73
Identities = 130/161 (80%), Positives = 141/161 (86%), Gaps = 1/161 (0%)
Frame = +2
Query: 10 PSIPESSKFSNSSQGTTTIGHSRDITSQSGSFGSGADQPLYPPNVYAPQAQAFYYGGFDN 69
P+ P++ K QGTTTIGHSR+ TSQSGS GS D PLYPPNVYAPQAQAFYY GFDN
Sbjct: 137 PAEPDNLK----EQGTTTIGHSRETTSQSGSLGSVGDCPLYPPNVYAPQAQAFYYRGFDN 304
Query: 70 GNGEWDEYPSYVNNGGIEIGSPGVYNENQSLVFHSGYGFNPQMPYGPYSPVTSPLPSVGG 129
GNGEWDEY SYVN+ G++IGSPGVYNEN SL+FHSGYGFNPQMPYGPYSPVT+PLPSVGG
Sbjct: 305 GNGEWDEYSSYVNSEGLDIGSPGVYNENPSLIFHSGYGFNPQMPYGPYSPVTTPLPSVGG 484
Query: 130 DAQLYSPQQFPYT-PPYYNQLVPPPNLPYLNSPTPVSQPEL 169
D QLYSPQQFPYT PPYY+QLV P +LPYLNSPTPVSQPEL
Sbjct: 485 DTQLYSPQQFPYTGPPYYHQLV-PXSLPYLNSPTPVSQPEL 604
>TC219737 similar to UP|Q9LNG4 (Q9LNG4) F21D18.17, partial (23%)
Length = 1333
Score = 233 bits (594), Expect = 2e-61
Identities = 109/157 (69%), Positives = 128/157 (81%), Gaps = 4/157 (2%)
Frame = +1
Query: 400 HKSIKYGVWASTPNGNRKLDAAYCQAK----EKQDACRIFLFFSVNASAQFCGVAEMVGP 455
HKSIKY VW+STP+GN+KL+ AY AK EK + C IFL FSVNAS QFCGVAEMVG
Sbjct: 1 HKSIKYNVWSSTPHGNKKLENAYEDAKKIAAEKSEVCPIFLLFSVNASGQFCGVAEMVGT 180
Query: 456 VNFDKSVDFWQQDKWSGQFPVKWHIIKDVPNSQFRHIVLENNDNKPVTNSRDTQEVKLQQ 515
V+F K++DFWQQDKWSG FPVKWHIIKDVPN FRHI+LENN+NKPVTNSRD QE+ +
Sbjct: 181 VDFSKNMDFWQQDKWSGSFPVKWHIIKDVPNPNFRHIILENNENKPVTNSRDAQEIMYLK 360
Query: 516 GIEMLTIFKNYETDTSILDDFDFYEDRQKAMQERKAR 552
G+EML IFKN+ TS+LDDF +YE+RQK MQ+ KA+
Sbjct: 361 GLEMLKIFKNHTLKTSLLDDFMYYENRQKIMQDEKAK 471
>TC230344 similar to UP|Q9LNG4 (Q9LNG4) F21D18.17, partial (20%)
Length = 828
Score = 216 bits (549), Expect = 3e-56
Identities = 122/257 (47%), Positives = 156/257 (60%), Gaps = 13/257 (5%)
Frame = +1
Query: 253 SVRSFGPSAGMASHQQRSLYGFGSSSNPYGRG-------YLSNPGSSFGGSAISGLSAND 305
S R F A HQ RS+ + G L S G + G +AN
Sbjct: 64 SGRGFLNMASSPVHQARSIDASTHPVDTISNGNVLSHHNQLKIASSLSSGFSDYGSNANG 243
Query: 306 RSFLSLENSRRHGRETASFCRCNGTLDILSEQNRGPRASKLKNH--ISSENNAIDGSKNN 363
+S ++ + H + S NG+ D+L EQNRGPR S K+ ++ + A G N
Sbjct: 244 QSVVAKLRPKVHIGKGLS--EVNGSSDVLGEQNRGPRISNYKSKFPLAVKAYANKGDGNT 417
Query: 364 ASTAKFQDESLNRPDFATDFKDAKFFVIKSYSEDNVHKSIKYGVWASTPNGNRKLDAAYC 423
+ NR DF ++++AKFFVIKSYSED+VHKSIKY VW+STP+GN+KL +A+
Sbjct: 418 QENIIISTDQYNREDFPVNYENAKFFVIKSYSEDDVHKSIKYNVWSSTPHGNKKLQSAHE 597
Query: 424 QAKE----KQDACRIFLFFSVNASAQFCGVAEMVGPVNFDKSVDFWQQDKWSGQFPVKWH 479
AK K +C IFLFFSVNAS QFCGVAEM+GPV+F+K +DFWQQDKWSG FPVKW+
Sbjct: 598 DAKRIASGKFGSCPIFLFFSVNASGQFCGVAEMIGPVDFNKDMDFWQQDKWSGSFPVKWY 777
Query: 480 IIKDVPNSQFRHIVLEN 496
IIKDV N+ FRHI+LEN
Sbjct: 778 IIKDVSNANFRHIILEN 828
>TC226517 similar to UP|YTH2_HUMAN (Q9Y5A9) YTH domain protein 2
(High-glucose-regulated protein 8) (NY-REN-2 antigen)
(CLL-associated antigen KW-14), partial (8%)
Length = 740
Score = 190 bits (482), Expect = 2e-48
Identities = 105/202 (51%), Positives = 127/202 (61%), Gaps = 4/202 (1%)
Frame = +3
Query: 415 NRKLDAAYCQAK----EKQDACRIFLFFSVNASAQFCGVAEMVGPVNFDKSVDFWQQDKW 470
N++L A+ EK C +FLFFSVNAS QF EM G V+F+KS+DF QDKW
Sbjct: 27 NKRLXGAFQDXXKRMXEKGCKCPVFLFFSVNASGQFAEXXEMTGRVDFNKSMDFXXQDKW 206
Query: 471 SGQFPVKWHIIKDVPNSQFRHIVLENNDNKPVTNSRDTQEVKLQQGIEMLTIFKNYETDT 530
+G F VKWHIIKDVPN Q RHI+LENND+KPVTNSRDTQEV QG+EML IFKNY T
Sbjct: 207 NGYFSVKWHIIKDVPNPQLRHIILENNDHKPVTNSRDTQEVSFPQGVEMLNIFKNYVART 386
Query: 531 SILDDFDFYEDRQKAMQERKARQQSSIMPTGLVGGSEHRSSSDSTGDFIKQVSKNFSLVV 590
SILDDF+FYE RQK +QE+K RQ MP V E +++ + D + VV
Sbjct: 387 SILDDFEFYESRQKVLQEKKTRQS---MPHTSVQQIEELTTTLGSVDLSSVKNMEDPKVV 557
Query: 591 RLDDNNNEVIAANRDKLASDVP 612
+ N+ IA D + D P
Sbjct: 558 ---ERVND*IACGADAIPGDPP 614
>TC220913 similar to UP|YTH2_HUMAN (Q9Y5A9) YTH domain protein 2
(High-glucose-regulated protein 8) (NY-REN-2 antigen)
(CLL-associated antigen KW-14), partial (5%)
Length = 898
Score = 129 bits (323), Expect(2) = 6e-34
Identities = 65/138 (47%), Positives = 90/138 (65%), Gaps = 14/138 (10%)
Frame = +2
Query: 465 WQQDKWSGQFPVKWHIIKDVPNSQFRHIVLENNDNKPVTNSRDTQEVKLQQGIEMLTIFK 524
W+ DK++G FP+KWHIIKDVPN+QF HI+L +N+NKPVT +RDTQE+ L++G+EML IF+
Sbjct: 80 WKLDKYNGFFPIKWHIIKDVPNNQFVHIILPSNENKPVTYTRDTQEIGLKEGLEMLNIFR 259
Query: 525 NYETDTSILDDFDFYEDRQKAMQERKARQQSSIMPTGLVGGS--------------EHRS 570
+Y TS+LDDFDFYE R+K + +++ + P V G+ E +S
Sbjct: 260 SYTAKTSLLDDFDFYERREKLFRSQRSTKHKHARPEQEVYGNDNYQNTVKAREKKMEMQS 439
Query: 571 SSDSTGDFIKQVSKNFSL 588
S G IK +KN SL
Sbjct: 440 SGTKQGTLIKH-TKNLSL 490
Score = 34.3 bits (77), Expect(2) = 6e-34
Identities = 15/24 (62%), Positives = 19/24 (78%)
Frame = +3
Query: 440 VNASAQFCGVAEMVGPVNFDKSVD 463
VNAS QF GVAEM+GPV+F ++
Sbjct: 3 VNASRQFVGVAEMLGPVDFKNDMN 74
>AW595996
Length = 328
Score = 116 bits (290), Expect = 3e-26
Identities = 55/87 (63%), Positives = 66/87 (75%)
Frame = -1
Query: 467 QDKWSGQFPVKWHIIKDVPNSQFRHIVLENNDNKPVTNSRDTQEVKLQQGIEMLTIFKNY 526
QDKW+G F V WHII DV N + RHI+LEN+D+K VT+SR+TQEV QG+ ML IFK Y
Sbjct: 328 QDKWNGYFHVTWHII*DVTNLRLRHIILENSDHKRVTSSRNTQEVSFPQGVTMLNIFK*Y 149
Query: 527 ETDTSILDDFDFYEDRQKAMQERKARQ 553
TSILDD++FY RQK MQE+K RQ
Sbjct: 148 VARTSILDDYEFYASRQKVMQEKKTRQ 68
>BE555214
Length = 412
Score = 104 bits (259), Expect = 1e-22
Identities = 45/56 (80%), Positives = 50/56 (88%)
Frame = +1
Query: 440 VNASAQFCGVAEMVGPVNFDKSVDFWQQDKWSGQFPVKWHIIKDVPNSQFRHIVLE 495
VNAS QFCGVAEM G V+F+KS+DFWQQDKW+G FPVKWHIIKDVPN Q RHI+LE
Sbjct: 193 VNASGQFCGVAEMTGRVDFNKSMDFWQQDKWNGYFPVKWHIIKDVPNPQLRHIILE 360
Score = 32.0 bits (71), Expect = 0.86
Identities = 17/38 (44%), Positives = 20/38 (51%)
Frame = +3
Query: 474 FPVKWHIIKDVPNSQFRHIVLENNDNKPVTNSRDTQEV 511
FP + K S ND+KPVT+SRDTQEV
Sbjct: 294 FPCQMAHYKGCSKSSTTAYNTRENDHKPVTSSRDTQEV 407
>TC205111 similar to UP|Q6IH72 (Q6IH72) HDC03105, partial (19%)
Length = 626
Score = 79.7 bits (195), Expect = 4e-15
Identities = 50/147 (34%), Positives = 74/147 (50%), Gaps = 13/147 (8%)
Frame = +2
Query: 13 PESSKFSNSSQGTT------TIGHSRDITSQSGSFGSGADQ----PLYPPNVYAPQA--Q 60
P S K ++ Q T IG + +F +GA + P P + + P
Sbjct: 74 PSSDKTADLLQNLTLDSESKAIGVAEPAKKNGPAFSNGAAKGRAKPFNPNSCFVPNGYPS 253
Query: 61 AFYYGGFDNGNGEWDEYPSYVN-NGGIEIGSPGVYNENQSLVFHSGYGFNPQMPYGPYSP 119
A+YYGG+D G G+W+ Y Y+N +GG+ + GVY ++ S ++H GYG+ P YG Y+P
Sbjct: 254 AYYYGGYD-GQGDWNAYSRYMNLDGGM---AQGVYGDSCSYMYHQGYGYTP---YGTYAP 412
Query: 120 VTSPLPSVGGDAQLYSPQQFPYTPPYY 146
S P + D Q Y QQ+ Y YY
Sbjct: 413 PNSSSPMIQQDGQHYGLQQYQYPWSYY 493
>TC217366 weakly similar to UP|Q9LUT8 (Q9LUT8) Arabidopsis thaliana genomic
DNA, chromosome 3, P1 clone: MGD8, partial (3%)
Length = 944
Score = 55.8 bits (133), Expect = 6e-08
Identities = 41/127 (32%), Positives = 63/127 (49%), Gaps = 25/127 (19%)
Frame = +2
Query: 515 QGIEMLTIFKNYETDTSILDDFDFYEDRQKAMQERKAR------QQSSIMPT-------G 561
+G+EML IFKN+ TS+LDDF +YE+RQK MQE KA+ + ++PT
Sbjct: 8 KGLEMLKIFKNHNLKTSLLDDFMYYENRQKIMQEEKAKLLIRSFKNPLVLPTLEPPRKLN 187
Query: 562 LV-----GGSEHRSSSDSTGDFIKQVSKNFSLVVRLD-------DNNNEVIAANRDKLAS 609
V G E + D D +KQ+S +V + D E + +++ +AS
Sbjct: 188 FVIDIPPVGDEKNAKMDDEVDSLKQISSAGHIVSSSEITSTTSVDEKAEKGSVDKEDIAS 367
Query: 610 DVPTGNV 616
+ G+V
Sbjct: 368 VLKIGSV 388
>TC233247 weakly similar to UP|Q9LUT8 (Q9LUT8) Arabidopsis thaliana genomic
DNA, chromosome 3, P1 clone: MGD8, partial (4%)
Length = 676
Score = 55.5 bits (132), Expect = 7e-08
Identities = 65/230 (28%), Positives = 91/230 (39%), Gaps = 18/230 (7%)
Frame = +3
Query: 44 GADQPLYPPNVYAPQAQ--AFYYGGFDNGNGEWDEYPSYVNNGGIEIGSPGVYNENQSLV 101
GA + NVY P A +Y GF+ GEW+++ G I G NE+ V
Sbjct: 27 GAPEYFCYQNVYYPAATNYGYYCTGFETP-GEWEDHHRIFGVDGPNIQFMGAQNESLPYV 203
Query: 102 FHS-GYGFNPQMPYGPYSPVTSPLPSVGGDAQLYSPQQFPYTPPYYNQLVPPPNLPYLNS 160
+++ GY +P PY PY P +G D L Q + YT P Y V P
Sbjct: 204 YYNYGYAQSPYNPYNPYIPGA----MIGADGSLGGGQHY-YTLPNYQSPVSAPGY----- 353
Query: 161 PTPVSQPELTNLLGIDQQVESNFFGPRASYPSV-GSFARGSFPVAPGSFSFHESQQGFDG 219
P QP+ +FFG AS G + F A G+FS + S F
Sbjct: 354 -IPPVQPD-----NFSDSSADSFFGASASVSKPDGRGLKPKFNSASGNFSRNSSI--FLS 509
Query: 220 SRSGGLWSDSSKP-------------SERQRSFMPL-SPSVPQQPIGSVR 255
+++ L S +P S SF+ L SP+V Q + +R
Sbjct: 510 NQTSSLARASERPRANDGRKQGLTHASVSGSSFLNLASPAVHQSAVAKLR 659
>TC212285 similar to UP|Q9LNG4 (Q9LNG4) F21D18.17, partial (13%)
Length = 579
Score = 46.2 bits (108), Expect = 4e-05
Identities = 32/102 (31%), Positives = 50/102 (48%), Gaps = 5/102 (4%)
Frame = +2
Query: 23 QGTTTIGH--SRDITSQSGSFGSGADQPLYPPNVYAPQAQ--AFYYGGFDNGNGEWDEYP 78
+GT H S +I F GA + + N+Y P A +Y GF++ GEW+++
Sbjct: 242 EGTDLNSHLTSPNIQQFQAMFNDGAPEFVADQNLYYPAATNYGYYCTGFESP-GEWEDHH 418
Query: 79 SYVNNGGIEIGSPGVYNENQSLVFHS-GYGFNPQMPYGPYSP 119
G +I G NE+ ++++ YGF Q PY PY+P
Sbjct: 419 RIFGVDGPDIQYTGAQNESFPYIYYTPSYGF-AQSPYNPYNP 541
>TC219465 similar to GB|AAH53863.1|31808095|BC053863 YT521 protein {Homo
sapiens;} , partial (3%)
Length = 563
Score = 45.1 bits (105), Expect = 1e-04
Identities = 24/70 (34%), Positives = 37/70 (52%)
Frame = +3
Query: 387 KFFVIKSYSEDNVHKSIKYGVWASTPNGNRKLDAAYCQAKEKQDACRIFLFFSVNASAQF 446
K+F+IKS + N+H SI+ G+WA+ L A ++ + L FSVN S F
Sbjct: 372 KYFIIKSLNHQNIHLSIEKGIWATQIMNEPILQEA------XHNSGSVILIFSVNMSGSF 533
Query: 447 CGVAEMVGPV 456
G A+M+ +
Sbjct: 534 QGYAQMMSSI 563
>TC219036
Length = 1141
Score = 43.5 bits (101), Expect = 3e-04
Identities = 23/63 (36%), Positives = 32/63 (50%), Gaps = 2/63 (3%)
Frame = +3
Query: 466 QQDKWSGQFPVKWHIIKDVPNSQFRHIVLENNDNKPVTNSRDTQEVKLQQGIEMLTIF-- 523
Q + W F VKW + D+P + H+ ND KPV SRD QE+ G+ + +
Sbjct: 3 QSNPWGRSFKVKWMCLNDLPFHKTLHLKNPLNDYKPVKISRDCQELSPDIGLALCELLDG 182
Query: 524 KNY 526
KNY
Sbjct: 183 KNY 191
>AW350967 similar to GP|15912287|gb| AT5g61020/maf19_20 {Arabidopsis
thaliana}, partial (6%)
Length = 244
Score = 41.2 bits (95), Expect = 0.001
Identities = 18/44 (40%), Positives = 28/44 (62%)
Frame = -2
Query: 361 KNNASTAKFQDESLNRPDFATDFKDAKFFVIKSYSEDNVHKSIK 404
+ + S+A + + + DF ++ D FVIKSYSED++HKS K
Sbjct: 144 EKDKSSAILECDQYTKADFPEEYTDXXXFVIKSYSEDDIHKSSK 13
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.313 0.132 0.392
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 27,404,785
Number of Sequences: 63676
Number of extensions: 430442
Number of successful extensions: 3345
Number of sequences better than 10.0: 170
Number of HSP's better than 10.0 without gapping: 2372
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2844
length of query: 632
length of database: 12,639,632
effective HSP length: 103
effective length of query: 529
effective length of database: 6,081,004
effective search space: 3216851116
effective search space used: 3216851116
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 62 (28.5 bits)
Medicago: description of AC144538.7


