

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
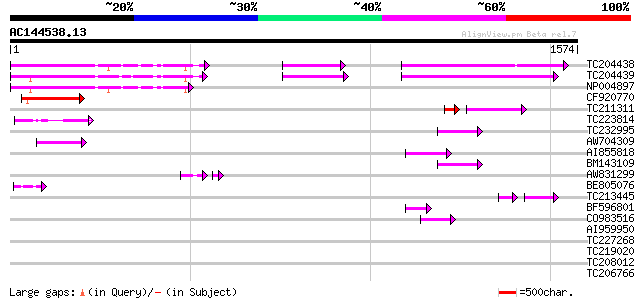
Query= AC144538.13 + phase: 0 /pseudo
(1574 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, co... 221 1e-57
TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete 213 5e-55
NP004897 gag-protease polyprotein 213 7e-55
CF920770 153 5e-37
TC211311 weakly similar to UP|O24587 (O24587) Pol protein, parti... 78 4e-19
TC223814 78 3e-14
TC232995 76 1e-13
AW704309 weakly similar to GP|6642775|gb| gag-pol polyprotein {V... 72 2e-12
AI855818 weakly similar to GP|21741393|e OSJNBb0051N19.6 {Oryza ... 58 3e-08
BM143109 54 7e-07
AW831299 47 2e-06
BE805076 weakly similar to GP|9294121|dbj copia-like retrotransp... 52 3e-06
TC213445 42 2e-05
BF596801 weakly similar to GP|29423270|g gag-pol polyprotein {Gl... 47 9e-05
CO983516 44 4e-04
AI959950 43 0.001
TC227268 33 0.99
TC219020 similar to UP|SRB7_SCHPO (O94376) RNA polymerase II med... 32 1.7
TC208012 similar to UP|Q761Z7 (Q761Z7) BRI1-KD interacting prote... 32 1.7
TC206766 similar to PIR|T47576|T47576 FKBP12 interacting protein... 32 1.7
>TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, complete
Length = 4734
Score = 221 bits (564), Expect = 1e-57
Identities = 170/589 (28%), Positives = 276/589 (45%), Gaps = 38/589 (6%)
Frame = +1
Query: 3 EGGSSNRPPLFEGTDYYYWKGKMELFLQS*DNHMWTVVENGNYIPYDEDL-----NE-KK 56
EGG NRPP+ +GT+Y YWK +M FL+S D+ W V G P D NE K
Sbjct: 16 EGGPVNRPPILDGTNYEYWKARMVAFLKSLDSRTWKAVIKGWEHPKMLDTEGKPTNELKP 195
Query: 57 KEDWSEEEGKRMLLNFKAKLFLTMALSREEYDRVQECKNAKEIWDTLKVHHEGTSHVKET 116
+EDW++EE + L N KA L + + + + C AK+ W+ LK HEGTS VK +
Sbjct: 196 EEDWTKEEDELALGNSKALNALFNGVDKNIFRLINTCTVAKDAWEILKTTHEGTSKVKMS 375
Query: 117 RIDIGVRKFELFEMKETETIDEMYGRFTIIMNELRSLEKDFTIHERVRKILRCLPRSWRH 176
R+ + KFE +MKE E I + + I N +L + T + VRKILR LP+ +
Sbjct: 376 RLQLLATKFENLKMKEEECIHDFHMNILEIANACTALGERMTDEKLVRKILRSLPKRFDM 555
Query: 177 IVTAITEAKDLKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQELEMNFD 236
VTAI EA+D+ +R+++LIGSL+ E+ L D++ KSK +A +N E E +++ D
Sbjct: 556 KVTAIEEAQDICNMRVDELIGSLQTFELGL-SDRTEKKSKNLAFVSNDEGEEDEYDLDTD 732
Query: 237 NQQQELDEEDHQDQIILLTRKLQRMIQRRDQNK----RNFP---------ARKENAKTEL 283
E + ++LL ++ +++ R D+ + RN P +K + K
Sbjct: 733 --------EGLTNAVVLLGKQFNKVLNRMDRRQKPHVRNIPFDIRKGSEYQKKSDEKPSH 888
Query: 284 DKSQVTCYGCYKLGHYKNECPLNKRKSSNLQTNQKSFMTTWDEPEEQTNEDEQADLCLMA 343
K + C+GC GH K ECP + +K Q S + D EQ ++ ++ L
Sbjct: 889 SKG-IQCHGCEGYGHIKAECPTHLKK----QRKGLSVCRSDDTESEQESDSDRDVNALTG 1053
Query: 344 KSDNEEVNLDPCSSCEKTEHLFDNLFYSSQMLEQECERLRNENLELRKEKEILEKENKDL 403
+ ++ E + D S E FD L S + L + E++ + +L+K LE E
Sbjct: 1054RFESAEDSSDTDS-----EITFDELAISYRELCIKSEKILQQEAQLKKVIANLEAE---- 1206
Query: 404 IKNIQHSESKEVSENKNNSQKENIILKENVLKLKNDISNFVKSTETFQKIMGSQVSVFDK 463
+ + +E+SE K E L + + I K ++ +++ +V ++
Sbjct: 1207----KEAHEEEISELKG----EVGFLNSKLENMTKSIKMLNKGSDMLDEVLQLGKNVGNQ 1362
Query: 464 ACIGFKTNQKQKLYENFFIPKKN-------------------KNTCERYKCTYCEKYGHL 504
+GF + F+P KN K+ ++++C YC KYGH+
Sbjct: 1363RGLGFNHKSAGRTTMTEFVPAKNSTGATMSQHRSRHHGTQQKKSKRKKWRCHYCGKYGHI 1542
Query: 505 EPFCFRKRKRLASQKFKNNYTNFHKKETLNNTNHQGPKKIWVPKALLTS 553
+PFC+ + + H +++ G K +WVPK + S
Sbjct: 1543KPFCY--------------HLHGHPHHGTQSSS-SGRKMMWVPKHKIVS 1644
Score = 105 bits (263), Expect = 1e-22
Identities = 141/466 (30%), Positives = 211/466 (45%), Gaps = 2/466 (0%)
Frame = +2
Query: 1087 KKIKFGLLYLILKLNLSLVQDGYLETN*MKKEK*HATRPDW*LKVTINKKALTTMKPMLL 1146
K +KFG +L + + L G T MKK * TRPD LK T+ K T MK L
Sbjct: 3278 KGMKFGS*FLDPRELM*LAPSGSSRTKPMKKVL*PETRPDLLLKATLRLKV*TLMKLSPL 3457
Query: 1147 WLDLKP*EFFLHMHLISASNFFKWT*KVLS*MVF*MKRFMFINLLVLKI*KNQITCSN*P 1206
LDL P + +L S S+ +W *+ *M *MK+ M+ + L I QI +
Sbjct: 3458 LLDLSPSDCYLV*LASSNSSCTRWM*RARF*MDT*MKKPMWSSQRDL*IQLIQIMYTGSR 3637
Query: 1207 KLYMD*NKLQEPGMKD*AFF*LKMGFHEEKLTPHFLEKLTKMIY*SYKCTSMISFLELQM 1266
+L MD*+KLQE GMK * L G E+LT L * ++ M LE
Sbjct: 3638 RLSMD*SKLQELGMKG*QSSLLSKGIGREELTRLSLSNKMLKT***HRYMLMTLCLEGCR 3817
Query: 1267 KKCVMIFQNSWKVNLK*V*WGNLNSFWDYKSNNTKMVFSFVKKNISKIFLKNIK*MKPKL 1326
+C I N +NL+*V + FWD K + K +S K ++ + +++ P +
Sbjct: 3818 MRCFDILSNRCNLNLR*VLLES*LIFWDSK*SRWKTPYSSHKASMQRTLSRSLGWKMPAI 3997
Query: 1327 *LLLCTLHQILTKMKMEKIFLKKNIEE**VLFYI*LPVDLTLFLQLVYVLDFKLLQKSHT 1386
L L KMK+ + +K E * +YI* DLT +Q V+V D K + + T
Sbjct: 3998 KEHLHLLT*SCQKMKLAPVLIKVCTEA*LGAYYI*QLADLTSPMQ*VFVQDIKPILR*VT 4177
Query: 1387 TPQLKEFSDTL*ALQVSVYGIKREHILI-LW-HIVMLTTPEIK*KEKVQVAHVNYLEKL* 1444
+ +EF + +* V++ G+ + I W IVML E++ EK + V+ E +
Sbjct: 4178 *IK*REF*N-M*MAPVTM-GLCTVIVQIQCWLGIVMLIGLEVQMTEKALLVDVSIWEPIL 4351
Query: 1445 SAGVVENKAL*HYPPQKLNIFPLQTAALRYYGLEIN*KIFL*GTQIFKYYAIILVL*IYQ 1504
G ++ + Y K +I + A +G *+ + VL I+
Sbjct: 4352 FHGSARSRTVCPYLLLKQSILQQEAAVHN*FG*SRC*RSTMSNKMS*HCTVTT*VLLIFL 4531
Query: 1505 KILFNIPDQNTSKLSITLFVTM*VKRKLNLFLLIQTIN*LIFLQNH 1550
KILFN + +T L IT+ + + + + +L * IF Q H
Sbjct: 4532 KILFNTAEPSTLTLDITILEILLMIKLSHWSMLTLRNK*QIFSQRH 4669
Score = 78.2 bits (191), Expect = 3e-14
Identities = 61/175 (34%), Positives = 93/175 (52%)
Frame = +3
Query: 758 TSHRFIWTYLNRKSWWQKIWLCNR**FFQIHLGFIFET*K*FF*SFSNLLQKSSK*ERNE 817
TSH Y + K W +K+ LC *F QI+LG +++ *S + K+SK +R
Sbjct: 2265 TSHGLDGAYAS*KPWRKKVCLCCCG*FLQIYLGQLYQREIRHL*SIQGVESKTSKRKRLC 2444
Query: 818 HYRSKK*SWRRI*KCLF*TILQ**WNIS*FFLS*NSSTKWSC*KKK*DFARNGKNYFK*I 877
H +++* W+R+*K IL * + S* S ++TKW *K+K DFAR+ +
Sbjct: 2445 HQENQE*PWQRV*KQQVY*ILHI*RHHS*VLCSHYTTTKWHS*KEKQDFARSC*GHASCQ 2624
Query: 878 KC*EIFLGRSHKYILLHFKQSFNKKSFKQNPV*TLERKKARHFLFSYFWMLLLYF 932
+ LG SH++ +LH +QS K + V* LER++A + W +L+F
Sbjct: 2625 RTSL*SLG*SHEHSMLHPQQSHT*KRDSNHTV*NLEREEANCQALPHLWKSMLHF 2789
>TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete
Length = 4731
Score = 213 bits (542), Expect = 5e-55
Identities = 163/583 (27%), Positives = 272/583 (45%), Gaps = 37/583 (6%)
Frame = +1
Query: 3 EGGSSNRPPLFEGTDYYYWKGKMELFLQS*DNHMWTVVENGNYIPYDEDLNEKK------ 56
EGG NRPP+ +G++Y YWK +M FL+S D+ W V G P D K
Sbjct: 16 EGGPVNRPPILDGSNYEYWKARMVAFLKSLDSRTWKAVIKGWEHPKMLDTEGKPTDELKP 195
Query: 57 KEDWSEEEGKRMLLNFKAKLFLTMALSREEYDRVQECKNAKEIWDTLKVHHEGTSHVKET 116
+EDW++EE + L N KA L + + + + C AK+ W+ LK+ HEGTS VK +
Sbjct: 196 EEDWTKEEDELALGNSKALNALFNGVDKNIFRLINTCTVAKDAWEILKITHEGTSKVKMS 375
Query: 117 RIDIGVRKFELFEMKETETIDEMYGRFTIIMNELRSLEKDFTIHERVRKILRCLPRSWRH 176
R+ + KFE +MKE E I + + I N +L + T + VRKILR LP+ +
Sbjct: 376 RLQLLATKFENLKMKEEECIHDFHMNILEIANACTALGERITDEKLVRKILRSLPKRFDM 555
Query: 177 IVTAITEAKDLKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQELEMNFD 236
VTAI EA+D+ +R+++LIGSL+ E+ L D++ KSK +A +N E E +++ D
Sbjct: 556 KVTAIEEAQDICNMRVDELIGSLQTFELGL-SDRAEKKSKNLAFVSNDEGEEDEYDLDTD 732
Query: 237 NQQQELDEEDHQDQIILLTRKLQRMIQRRDQNKR----NFP-----ARKENAKTELDKSQ 287
E + ++LL ++ +++ R D+ ++ N P K ++++ S
Sbjct: 733 --------EGLTNAVVLLGKQFNKVLNRMDKRQKPHVQNIPFDIRKGSKYQKRSDVKPSH 888
Query: 288 ---VTCYGCYKLGHYKNECPLNKRKSSNLQTNQKSFMTTWDEPEEQTNEDEQADLCLMAK 344
+ C+GC GH ECP ++L+ ++K + E + D D + A
Sbjct: 889 SKGIQCHGCEGYGHIIAECP------THLKKHRKGLSVCQSDTESEQESDSDRD--VNAL 1044
Query: 345 SDNEEVNLDPCSSCEKTEHLFDNLFYSSQMLEQECERLRNENLELRKEKEILEKENKDLI 404
+ E D SS +E FD L S + L + E++ + +L+K LE E
Sbjct: 1045TGIFETAED--SSDTDSEITFDELAASYRKLCIKSEKILQQEAQLKKVIADLEAE----- 1203
Query: 405 KNIQHSESKEVSENKNNSQKENIILKENVLKLKNDISNFVKSTETFQKIMGSQVSVFDKA 464
+ + +E+SE K E L + + I K ++T +++ + ++
Sbjct: 1204---KEAHKEEISELKG----EVGFLNSKLENMTKSIKMLNKGSDTLDEVLLLGKNAGNQR 1362
Query: 465 CIGFKTNQKQKLYENFFIPKKN-------------------KNTCERYKCTYCEKYGHLE 505
+GF + F+P KN K+ ++++C YC KYGH++
Sbjct: 1363GLGFNPKSAGRTTMTEFVPAKNRTGATMSQHRSRHHGMQQKKSKRKKWRCHYCGKYGHIK 1542
Query: 506 PFCFRKRKRLASQKFKNNYTNFHKKETLNNTNHQGPKKIWVPK 548
PFC+ + + H ++N + K +WVPK
Sbjct: 1543PFCY--------------HLHGHPHHGTQSSNSR-KKMMWVPK 1626
Score = 108 bits (270), Expect = 2e-23
Identities = 129/436 (29%), Positives = 194/436 (43%)
Frame = +2
Query: 1087 KKIKFGLLYLILKLNLSLVQDGYLETN*MKKEK*HATRPDW*LKVTINKKALTTMKPMLL 1146
K +K G +L L+ + L G T MKK * TRPDW LK T+ K T M+ +
Sbjct: 3275 KGMKSGS*FLGLRELM*LAPSGSSRTKPMKKVS*PETRPDWLLKATLRLKV*TLMRLLPQ 3454
Query: 1147 WLDLKP*EFFLHMHLISASNFFKWT*KVLS*MVF*MKRFMFINLLVLKI*KNQITCSN*P 1206
LDL P +++L + S S+ +W *+ *M *MK+ M+ + L+ QI +
Sbjct: 3455 LLDLSPSDYYLV*LVSSNSSCTRWM*RAHF*MDT*MKKSMWSSQRDLQTRLIQIMYTGSR 3634
Query: 1207 KLYMD*NKLQEPGMKD*AFF*LKMGFHEEKLTPHFLEKLTKMIY*SYKCTSMISFLELQM 1266
+L MD*+KLQE GMK * L G E+LT L * ++ M LE
Sbjct: 3635 RLSMD*SKLQELGMKG*QSSLLSKGIGREELTRPSLSNKMLKT**LHRYMLMTLCLEGCR 3814
Query: 1267 KKCVMIFQNSWKVNLK*V*WGNLNSFWDYKSNNTKMVFSFVKKNISKIFLKNIK*MKPKL 1326
+C I N +NL+*V + FWD+K + + +S K + + +++ P +
Sbjct: 3815 MRCFDILFNRCNLNLR*VLLES*LIFWDFK*SRWRTPYSSHKAGMQRTLSRSLGWRMPVI 3994
Query: 1327 *LLLCTLHQILTKMKMEKIFLKKNIEE**VLFYI*LPVDLTLFLQLVYVLDFKLLQKSHT 1386
L L +MK + +K E ** +YI* D T +Q V+V D K + + T
Sbjct: 3995 KGHLHLLT*SCQRMKQAPVLIKVCTEA**GAYYI*QLADPTSPMQ*VFVQDIKPIPR*VT 4174
Query: 1387 TPQLKEFSDTL*ALQVSVYGIKREHILILWHIVMLTTPEIK*KEKVQVAHVNYLEKL*SA 1446
+ +EF + AL I IVML E++ EK + + E
Sbjct: 4175 *LK*REF*NM*MALVTMGLCTVIVQIQCWLGIVMLIGLEVQMTEKALLVDASIWETTLFH 4354
Query: 1447 GVVENKAL*HYPPQKLNIFPLQTAALRYYGLEIN*KIFL*GTQIFKYYAIILVL*IYQKI 1506
G ++ + Y QK +I + A +G *+ + VL I+ KI
Sbjct: 4355 GSARSRTVCPYLQQKPSILQQEAAVHS*FG*SRC*RSTMSNKMS*HCTVTT*VLLIFLKI 4534
Query: 1507 LFNIPDQNTSKLSITL 1522
LFN + +T L IT+
Sbjct: 4535 LFNTAEPSTLTLDITI 4582
Score = 82.4 bits (202), Expect = 1e-15
Identities = 64/183 (34%), Positives = 98/183 (52%)
Frame = +3
Query: 758 TSHRFIWTYLNRKSWWQKIWLCNR**FFQIHLGFIFET*K*FF*SFSNLLQKSSK*ERNE 817
TSH F Y KSW +++ LC *F QI+LG +++ *S + K+SK ER
Sbjct: 2262 TSHGFDGAYAG*KSWRKEVCLCCCG*FLQIYLGKLYQREIRNL*SIQRVESKTSKRERLC 2441
Query: 818 HYRSKK*SWRRI*KCLF*TILQ**WNIS*FFLS*NSSTKWSC*KKK*DFARNGKNYFK*I 877
H +++* W+RI*K IL * + S* S ++T+W *++K DFAR +
Sbjct: 2442 HQENQE*PWQRI*KQQVH*ILHI*RHHS*VLCSHYTTTEWDS*EEKQDFARGCSGHASCQ 2621
Query: 878 KC*EIFLGRSHKYILLHFKQSFNKKSFKQNPV*TLERKKARHFLFSYFWMLLLYFKQQR* 937
+ LG SH++ +LH +QS +K +PV* LER++A + W +L+ + R
Sbjct: 2622 RTSL*SLG*SHEHSMLHPQQSHTEKRDSNHPV*NLEREEAICQALPHLWKSMLHLGR*RA 2801
Query: 938 PRK 940
+K
Sbjct: 2802 KKK 2810
>NP004897 gag-protease polyprotein
Length = 1923
Score = 213 bits (541), Expect = 7e-55
Identities = 157/544 (28%), Positives = 256/544 (46%), Gaps = 37/544 (6%)
Frame = +1
Query: 3 EGGSSNRPPLFEGTDYYYWKGKMELFLQS*DNHMWTVVENGNYIPYDEDLNEKK------ 56
EGG NRPP+ +GT+Y YWK +M FL+S D+ W V P D K
Sbjct: 16 EGGPVNRPPILDGTNYEYWKARMVAFLKSLDSRTWKAVIKDWEHPKMLDTEGKPTDGLKP 195
Query: 57 KEDWSEEEGKRMLLNFKAKLFLTMALSREEYDRVQECKNAKEIWDTLKVHHEGTSHVKET 116
+EDW++EE + L N KA L + + + + C AK+ W+ LK HEGTS VK +
Sbjct: 196 EEDWTKEEDELALGNSKALNALFNGVDKNIFRLINTCTVAKDAWEILKTTHEGTSKVKMS 375
Query: 117 RIDIGVRKFELFEMKETETIDEMYGRFTIIMNELRSLEKDFTIHERVRKILRCLPRSWRH 176
R+ + KFE +MKE E I + + I N +L + T + VRKILR LP+ +
Sbjct: 376 RLQLLATKFENLKMKEEECIHDFHMNILEIANACTALGERMTDEKLVRKILRSLPKRFDM 555
Query: 177 IVTAITEAKDLKKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQELEMNFD 236
VTAI EA+D+ LR+++LIGSL+ E+ L D++ KSK +A +N E E +++ D
Sbjct: 556 KVTAIEEAQDICNLRVDELIGSLQTFELGL-SDRTEKKSKNLAFVSNDEGEEDEYDLDTD 732
Query: 237 NQQQELDEEDHQDQIILLTRKLQRMIQRRDQNK----RNFP--------ARKENAKTELD 284
E + ++LL ++ +++ R D+ + RN P +K + +
Sbjct: 733 --------EGLTNAVVLLGKQFNKVLNRMDRRQKPHVRNIPFDIRKGSEYQKRSDEKPSH 888
Query: 285 KSQVTCYGCYKLGHYKNECPLNKRKSSNLQTNQKSFMTTWDEPEEQTNEDEQADLCLMAK 344
C+GC GH K ECP + +K Q S + D EQ ++ ++ L +
Sbjct: 889 SKGFQCHGCEGYGHIKAECPTHLKK----QRKGLSVCRSDDTESEQESDSDRDVNALTGR 1056
Query: 345 SDNEEVNLDPCSSCEKTEHLFDNLFYSSQMLEQECERLRNENLELRKEKEILEKENKDLI 404
++ E + D S E FD L S + L + E++ + +L+K LE E
Sbjct: 1057FESAEDSSDTDS-----EITFDELATSYRELCIKSEKILQQEAQLKKVIANLEAE----- 1206
Query: 405 KNIQHSESKEVSENKNNSQKENIILKENVLKLKNDISNFVKSTETFQKIMGSQVSVFDKA 464
+ + +E+SE K E L + + I K ++ +++ +V ++
Sbjct: 1207---KEAHEEEISELKG----EVGFLNSKLENMTKSIKMLNKGSDMLDEVLQLGKNVGNQR 1365
Query: 465 CIGFKTNQKQKLYENFFIPKK-------------------NKNTCERYKCTYCEKYGHLE 505
+GF ++ F+P K K+ ++++C YC KYGH++
Sbjct: 1366GLGFNHKSAGRITMTEFVPAKISTGATMSQHRSRHHGTQQKKSKRKKWRCHYCGKYGHIK 1545
Query: 506 PFCF 509
PFC+
Sbjct: 1546PFCY 1557
>CF920770
Length = 581
Score = 153 bits (387), Expect = 5e-37
Identities = 81/187 (43%), Positives = 113/187 (60%), Gaps = 13/187 (6%)
Frame = -2
Query: 33 DNHMWTVVENGNYIPY------------DEDLN-EKKKEDWSEEEGKRMLLNFKAKLFLT 79
D ++W +E G YIP E + EK ++ WSEE+ KR+ N KAK +T
Sbjct: 574 DLNIWEAIEIGPYIPTTVERVSIDGSSSSESITIEKPRDRWSEEDRKRVQYNLKAKNIIT 395
Query: 80 MALSREEYDRVQECKNAKEIWDTLKVHHEGTSHVKETRIDIGVRKFELFEMKETETIDEM 139
AL +EY RV CK+AKE+WDTL++ HEGT+ VK +RI+ ++ELF M E I M
Sbjct: 394 SALGMDEYFRVSNCKSAKEMWDTLRLTHEGTTDVKRSRINALTHEYELFRMNTNENIQSM 215
Query: 140 YGRFTIIMNELRSLEKDFTIHERVRKILRCLPRSWRHIVTAITEAKDLKKLRLEDLIGSL 199
RFT I+N L +L K+F + + K+LRCL R W+ VTAI+E++DL + L L G L
Sbjct: 214 QKRFTHIVNHLAALGKEFQNEDLINKVLRCLSREWQPKVTAISESRDLSNMSLATLFGKL 35
Query: 200 KAHEVLL 206
+ HE+ L
Sbjct: 34 QEHEMEL 14
>TC211311 weakly similar to UP|O24587 (O24587) Pol protein, partial (15%)
Length = 1213
Score = 78.2 bits (191), Expect(2) = 4e-19
Identities = 64/168 (38%), Positives = 89/168 (52%)
Frame = +1
Query: 1267 KKCVMIFQNSWKVNLK*V*WGNLNSFWDYKSNNTKMVFSFVKKNISKIFLKNIK*MKPKL 1326
K C F + +++LK V* + +S D+KS M F +K+NI K + MKP L
Sbjct: 538 KGCARSFLS**RMDLKRV*KVS*SSS*DFKSFRKFMGFLSIKRNIQSPI*KGSEWMKPNL 717
Query: 1327 *LLLCTLHQILTKMKMEKIFLKKNIEE**VLFYI*LPVDLTLFLQLVYVLDFKLLQKSHT 1386
LC + Q LT+M+ I K++I * +L++I*L VD L L +V DF L+QK
Sbjct: 718 WQPLCIVPQSLTRMRKVIILHKRSIVV*LILYHI*LLVDQILCLSFAFVQDFSLIQKFLM 897
Query: 1387 TPQLKEFSDTL*ALQVSVYGIKREHILILWHIVMLTTPEIK*KEKVQV 1434
QLK D L L + VYG+++ LI IVM IK*K + V
Sbjct: 898 LLQLKGS*DILLELLIIVYGLRKGLSLIF*DIVMFILLVIK*KGRALV 1041
Score = 36.6 bits (83), Expect(2) = 4e-19
Identities = 21/40 (52%), Positives = 26/40 (64%)
Frame = +3
Query: 1208 LYMD*NKLQEPGMKD*AFF*LKMGFHEEKLTPHFLEKLTK 1247
++M *NKL E GMK * F* +M EE TPH+ E+L K
Sbjct: 360 VFMV*NKL*ELGMKG*VHF*FQMDSPEE*RTPHYSERLKK 479
>TC223814
Length = 607
Score = 78.2 bits (191), Expect = 3e-14
Identities = 64/220 (29%), Positives = 94/220 (42%), Gaps = 1/220 (0%)
Frame = +1
Query: 13 FEGTDYYYWKGKMELFLQS*DNHMWTVVENGNYIPYDEDLNEKKKEDWSEEEGKRMLLNF 72
F+G +Y WK ++ + MW VVENGNYIP DE E + W+++
Sbjct: 1 FKGQNYDNWKQRIIELFDACHIDMWDVVENGNYIPIDEKGGENPRSAWTKDH-------- 156
Query: 73 KAKLFLTMALSREEYDRVQECKNAKEIWDTLKVHHEGTSHVKETRIDIGVRKFELFEMKE 132
+ K FL Y + + + + D KV +
Sbjct: 157 QTKYFL--------YSKARNTRPSNNCLDNSKVQN------------------------- 237
Query: 133 TETIDEMYGRFTIIMNELRSLEKDFTIHE-RVRKILRCLPRSWRHIVTAITEAKDLKKLR 191
IMN LRSL K H+ + KIL+ L R V A+ ++KDLK L
Sbjct: 238 -------------IMNNLRSLSKT*DNHDDHITKILQSLLIQ*RP*VIALCDSKDLKSLP 378
Query: 192 LEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQEL 231
+E+ G+L+ HE+ L ED+ K K IA K Q++L + L
Sbjct: 379 VEEFDGTLQVHELELMEDEGQRKGKFIASKV-QKALKRSL 495
>TC232995
Length = 1009
Score = 75.9 bits (185), Expect = 1e-13
Identities = 51/125 (40%), Positives = 66/125 (52%)
Frame = +3
Query: 1189 NLLVLKI*KNQITCSN*PKLYMD*NKLQEPGMKD*AFF*LKMGFHEEKLTPHFLEKLTKM 1248
N LVLK NQ N +L+M *NK GM D* F LK E K PH+ + + M
Sbjct: 9 NPLVLKFLINQTMFINYKRLFMV*NKPLGHGMND*VIFFLKKNSPEVKWIPHYS*RESIM 188
Query: 1249 IY*SYKCTSMISFLELQMKKCVMIFQNSWKVNLK*V*WGNLNSFWDYKSNNTKMVFSFVK 1308
I+ +K MI FL+ M C F KVNLK *W N ++FWDYKS+ V+S +
Sbjct: 189 IFCWFKYMLMI*FLDPLMIHCARSFPLICKVNLKCQ*WEN*STFWDYKSSKLNKVYSSIN 368
Query: 1309 KNISK 1313
N ++
Sbjct: 369 PNTAR 383
>AW704309 weakly similar to GP|6642775|gb| gag-pol polyprotein {Vitis
vinifera}, partial (23%)
Length = 423
Score = 72.0 bits (175), Expect = 2e-12
Identities = 42/140 (30%), Positives = 76/140 (54%), Gaps = 2/140 (1%)
Frame = -1
Query: 74 AKLFLTMALSREEYDRVQECKNAKEIWDTLKVHHEGTSHVKETRIDIGVRKFELFEMKET 133
A+ L +S+ + R+ K+ K IWD LK + G ++ ++ R+FEL MKE+
Sbjct: 423 ARSCLFTGVSQMIFIRIMTLKSPKAIWDCLKEEYAGDDRIRSMQVLNLRREFEL*RMKES 244
Query: 134 ETIDEMYGRFTIIMNELRSLEKDFTIHERVRKILRCLPRSWRHIVTAITEAKDLKKLRLE 193
ETI E + I+N+++ L DF V K L +P + + ++ KDL K+ L
Sbjct: 243 ETIKEYSNKLLGIVNKIKLLGSDFADSRIVEKNLVTVPERYEAFIASLENTKDLSKITLA 64
Query: 194 DLIGSLKAHEV--LLQEDKS 211
+++ +L+A E L+++D++
Sbjct: 63 EVLHALQAQEQRRLMRQDRA 4
>AI855818 weakly similar to GP|21741393|e OSJNBb0051N19.6 {Oryza sativa
(japonica cultivar-group)}, partial (10%)
Length = 463
Score = 58.2 bits (139), Expect = 3e-08
Identities = 54/129 (41%), Positives = 64/129 (48%)
Frame = -1
Query: 1098 LKLNLSLVQDGYLETN*MKKEK*HATRPDW*LKVTINKKALTTMKPMLLWLDLKP*EFFL 1157
LK+ LS Q+G+LE N*M R D * K I K+ T K ML D K E F
Sbjct: 397 LKIILS*EQNGFLEIN*MNMA*LLEIRLD**QKGIIKKRE*TMKKHMLQLQD*KSLECF* 218
Query: 1158 HMHLISASNFFKWT*KVLS*MVF*MKRFMFINLLVLKI*KNQITCSN*PKLYMD*NKLQE 1217
HM+ N KW KVL *MV K++M N LK NQ+ N +L+M *NK
Sbjct: 217 HMYP**ILNSIKWMLKVLF*MV*FKKKYMLNNPQALKSRINQLMFINCKRLFMV*NKPLG 38
Query: 1218 PGMKD*AFF 1226
GM *A F
Sbjct: 37 RGMNV*AIF 11
>BM143109
Length = 415
Score = 53.5 bits (127), Expect = 7e-07
Identities = 51/127 (40%), Positives = 65/127 (51%), Gaps = 1/127 (0%)
Frame = +2
Query: 1188 INLLVLKI*KNQITCSN*PKLYMD*NKLQEPGMKD*A-FF*LKMGFHEEKLTPHFLEKLT 1246
INLL K K+ I N* + YMD*NK GM * FF* ++ F + +L FL K
Sbjct: 2 INLL*GKTQKSLIMSLN*KRFYMD*NKPLGLGMNF*VNFF*TRV-FQKVRLILTFLFKRN 178
Query: 1247 KMIY*SYKCTSMISFLELQMKKCVMIFQNSWKVNLK*V*WGNLNSFWDYKSNNTKMVFSF 1306
MIY Y+ MI FL M F K+NLK * + F DYKS+ +M +
Sbjct: 179 *MIYS*YRYMLMILFLVQLMILFAKNFLKICKMNLKCQ*CVS*TFFLDYKSSKQRMEYLS 358
Query: 1307 VKKNISK 1313
V +NI+K
Sbjct: 359 VNQNIAK 379
>AW831299
Length = 334
Score = 47.0 bits (110), Expect(2) = 2e-06
Identities = 29/80 (36%), Positives = 39/80 (48%), Gaps = 4/80 (5%)
Frame = +3
Query: 474 QKLYENFFIPKKNKNTCERYKCTYCEKYGHLEPFCFRKRKRLASQKFKNNYTNFHK---- 529
QK+++NF + N+ C YC + GH C+ FK NY+N
Sbjct: 9 QKVHKNFSTSTQKCNS-NSITCFYCGRRGHGISTCY----------FKKNYSNIKMIWVP 155
Query: 530 KETLNNTNHQGPKKIWVPKA 549
K + NTN QGP KIWVPK+
Sbjct: 156 KGSSVNTNMQGPNKIWVPKS 215
Score = 24.3 bits (51), Expect(2) = 2e-06
Identities = 14/30 (46%), Positives = 16/30 (52%)
Frame = +1
Query: 563 EKALVLGQWMLKTYDWR*TKLSLLHQKGRR 592
EK +V MLKTYDWR K K +R
Sbjct: 241 EKEVVHR*RMLKTYDWRCIKFYTHISKEKR 330
>BE805076 weakly similar to GP|9294121|dbj copia-like retrotransposable
element {Arabidopsis thaliana}, partial (2%)
Length = 388
Score = 51.6 bits (122), Expect = 3e-06
Identities = 31/91 (34%), Positives = 47/91 (51%), Gaps = 1/91 (1%)
Frame = +1
Query: 11 PLFEGTDYYYWKGKMELFLQS*DNHMWTVVENGNYIPYDED-LNEKKKEDWSEEEGKRML 69
P+F G +Y +W+ KME +L S D +W +VE G IP D LN ++++ + + K
Sbjct: 133 PIFNGENYDFWRVKMETYLSSQD--LWDIVEEGFTIPADTSALNASQEKELKKNKQK--- 297
Query: 70 LNFKAKLFLTMALSREEYDRVQECKNAKEIW 100
N K L A + + R+ K AKE W
Sbjct: 298 -NSKTLFTLQQAETDPIFPRIMGAKTAKEAW 387
>TC213445
Length = 705
Score = 42.0 bits (97), Expect(2) = 2e-05
Identities = 35/95 (36%), Positives = 48/95 (49%)
Frame = +2
Query: 1430 EKVQVAHVNYLEKL*SAGVVENKAL*HYPPQKLNIFPLQTAALRYYGLEIN*KIFL*GTQ 1489
EKV V V L L* G+V++K + Y QK NIF L+ + +G + N +*
Sbjct: 410 EKVLVTLVILLVLL*YHGIVKSKIVLSYQLQKQNIFLLEVIMHKSFG*DNNFLTMV*NLI 589
Query: 1490 IFKYYAIILVL*IYQKILFNIPDQNTSKLSITLFV 1524
I+ I V IY KI+F +Q+ K IT FV
Sbjct: 590 IYLSDVTIQVQLIYPKIIFCTLEQSILK*GITSFV 694
Score = 25.8 bits (55), Expect(2) = 2e-05
Identities = 22/53 (41%), Positives = 26/53 (48%)
Frame = +3
Query: 1357 LFYI*LPVDLTLFLQLVYVLDFKLLQKSHTTPQLKEFSDTL*ALQVSVYGIKR 1409
LF+I DL L V V D K + K+ T LKE D AL + YGI R
Sbjct: 207 LFFIYQQADLI*CLVFVCV*DIKQIPKNLT*VSLKE**DICWAL*I*GYGILR 365
>BF596801 weakly similar to GP|29423270|g gag-pol polyprotein {Glycine max},
partial (7%)
Length = 336
Score = 46.6 bits (109), Expect = 9e-05
Identities = 33/73 (45%), Positives = 38/73 (51%)
Frame = +1
Query: 1098 LKLNLSLVQDGYLETN*MKKEK*HATRPDW*LKVTINKKALTTMKPMLLWLDLKP*EFFL 1157
LK+ L L Q+G+LE N*M +P * K I K+ T K MLL D KP E F
Sbjct: 112 LKIILLLEQNGFLEIN*MNMV*LLEIKPG**RKDIIKKRE*TMKKHMLLLQD*KPLECFW 291
Query: 1158 HMHLISASNFFKW 1170
HMH NF KW
Sbjct: 292 HMHP**TLNFIKW 330
>CO983516
Length = 724
Score = 44.3 bits (103), Expect = 4e-04
Identities = 37/98 (37%), Positives = 53/98 (53%)
Frame = +3
Query: 1141 MKPMLLWLDLKP*EFFLHMHLISASNFFKWT*KVLS*MVF*MKRFMFINLLVLKI*KNQI 1200
++ +L LDL P + +L S S+ +W *+ *M *MK+ M+ + L I QI
Sbjct: 357 IRSSILLLDLSPSDCYLV*LASSNSSCTRWM*RARF*MDT*MKKSMWSSQRDL*IQLIQI 536
Query: 1201 TCSN*PKLYMD*NKLQEPGMKD*AFF*LKMGFHEEKLT 1238
+ +L MD*+KLQE GMK * + L G E+LT
Sbjct: 537 MYTGSRRLSMD*SKLQELGMKG*QSYLLSKGIGREELT 650
>AI959950
Length = 466
Score = 42.7 bits (99), Expect = 0.001
Identities = 34/89 (38%), Positives = 46/89 (51%)
Frame = -2
Query: 1107 DGYLETN*MKKEK*HATRPDW*LKVTINKKALTTMKPMLLWLDLKP*EFFLHMHLISASN 1166
+GY TN* + + T+ D LKVT N+K TT KP+ L K + H+ I +
Sbjct: 303 NGYFVTN*TRMVRL*DTKQD*LLKVTHNRKV*TTQKPLHLLHV*K*YASYFHLQPIVI*S 124
Query: 1167 FFKWT*KVLS*MVF*MKRFMFINLLVLKI 1195
KW *KV *M ++FM N L LK+
Sbjct: 123 CIKWM*KVHF*MA*SKRKFMLNNRLDLKM 37
>TC227268
Length = 894
Score = 33.1 bits (74), Expect = 0.99
Identities = 56/277 (20%), Positives = 113/277 (40%), Gaps = 5/277 (1%)
Frame = +1
Query: 188 KKLRLEDLIGSLKAHEVLLQEDKSSNKSKMIALKTNQESLNQELEMNFDNQQQELDEEDH 247
K R+ + + V +Q + K K K + +S + ++ +N Q EL++
Sbjct: 148 KSKRINRKLPTASFDSVPVQANPEFEKPKRNTRKISNQSSDPHVQ---ENPQSELEK--- 309
Query: 248 QDQIILLTRKLQRMIQRRDQNKRNFPARKENAKTELDKSQVTCYGCYKLGHYKNECPLNK 307
+ R L+++ +N E K L+K VT C + +
Sbjct: 310 ------IKRNLRKVYNPVVENAVPSEVESEMPKDHLEKVTVT--SCLAVSE-------QE 444
Query: 308 RKSSNLQTNQKSFMTTWDEPEEQTNEDEQADLCLMAKSDNEEVNLDPCSSCEKTEHLFDN 367
SSN + +++ +T P+ +T S ++EV+ P S E
Sbjct: 445 VISSNEKIKKEAILTVSSVPDIETTP---------RLSVSKEVSDTPSSYQVTVESKPLT 597
Query: 368 LFYSSQMLEQECERLRNENLELRKEKEILEKENKDLIK-NIQHSESKEVSENKNNSQKEN 426
+ + ++NE ++L + I + EN L ++ H E + SEN+ +QK +
Sbjct: 598 EITTKDKNISVSDEVKNEPIDL--PEPICKDENSHLTNGDLSHKEDQIGSENQKPNQKAS 771
Query: 427 IILK----ENVLKLKNDISNFVKSTETFQKIMGSQVS 459
I+ K EN ++ + +++ +TE+ + + +Q S
Sbjct: 772 IVAKQERAENGIQNSPTLPSYMAATESAKAKLRAQGS 882
>TC219020 similar to UP|SRB7_SCHPO (O94376) RNA polymerase II mediator
complex protein srb7, partial (14%)
Length = 1446
Score = 32.3 bits (72), Expect = 1.7
Identities = 36/154 (23%), Positives = 71/154 (45%), Gaps = 3/154 (1%)
Frame = +2
Query: 328 EEQTNEDEQADLCLMAKSDNEEVNLDPCSSCEKTEHLFDNLFYSSQMLEQECERLRNENL 387
E T + EQ + K +N++ + S EK + L + + + + + L+NE
Sbjct: 488 ESLTRDSEQKLQEAIEKFNNKDSEVQ--SLLEKIKILEEQIAKAGE----QSTSLKNEFE 649
Query: 388 ELRKEKEILEKENKDLIKNIQHSESKEVSENKNNS--QKENIILKENVLKLKNDISNFVK 445
E + LE EN+DL + I +ESK N NI LK + +L+ +++ +
Sbjct: 650 ESLSKLTSLESENEDLKRQILDAESKSSQSFSENELLVGTNIQLKTKIDELEESLNHALS 829
Query: 446 STE-TFQKIMGSQVSVFDKACIGFKTNQKQKLYE 478
E Q+++ + S+ + + K+++ Q+ E
Sbjct: 830 EKEAAAQELVSHKNSITELNDLQSKSSEIQRANE 931
>TC208012 similar to UP|Q761Z7 (Q761Z7) BRI1-KD interacting protein 117
(Fragment), partial (38%)
Length = 993
Score = 32.3 bits (72), Expect = 1.7
Identities = 23/102 (22%), Positives = 40/102 (38%)
Frame = +1
Query: 266 DQNKRNFPARKENAKTELDKSQVTCYGCYKLGHYKNECPLNKRKSSNLQTNQKSFMTTWD 325
DQ K+ + K D ++ CY C K GH +C K + L ++ + +
Sbjct: 67 DQGKKPGSGPSDAPKVPSDAPKI-CYKCKKAGHLSRDC---KEQPDGL-LHRNAIGEAEE 231
Query: 326 EPEEQTNEDEQADLCLMAKSDNEEVNLDPCSSCEKTEHLFDN 367
P+ + QAD M + D E+ + ++L N
Sbjct: 232 NPKSTAIDTSQADRVAMEEDDINEIGEEEKEKLNDVDYLTGN 357
>TC206766 similar to PIR|T47576|T47576 FKBP12 interacting protein (FIP37) -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(74%)
Length = 1952
Score = 32.3 bits (72), Expect = 1.7
Identities = 47/227 (20%), Positives = 99/227 (42%), Gaps = 18/227 (7%)
Frame = +3
Query: 219 ALKTNQESLNQELE----------MNFDNQQQELDEEDHQDQIILLTRKLQRMIQRR--- 265
ALK+++ESL ++LE + F ++QE+ E + + + K M RR
Sbjct: 540 ALKSSEESLREQLEKAKKKEAAFIVTFAKREQEIAELKSAVRDLKVQLKPPSMQSRRLLL 719
Query: 266 DQNKRNFPARKENAKTELDKSQVTCYGCYKLGHYKNECPLNKRKSSNLQTNQKSFMTTWD 325
D R +N E DK + + + K + +T Q+ +
Sbjct: 720 DPAVHEEFTRLKNLVEEKDKKVKELQDNIAAVSFTPQSKMGKMLMAKCRTLQEEN----E 887
Query: 326 EPEEQTNEDEQADLCL---MAKSDNEEVNLDPCSSCEKTEHLFDNLFYSSQMLEQECERL 382
E Q +E + +L + + KS N ++ + E L +++ S++M+ E+L
Sbjct: 888 EIGNQASEGKMHELGMKLALQKSQNSQLRSQFEGLQKHMEGLTNDVERSNEMVLMLQEKL 1067
Query: 383 RNENLELRKEKEILEKENKD--LIKNIQHSESKEVSENKNNSQKENI 427
+ + E+++ K L+++N + +K+ ++ E K+ Q++N+
Sbjct: 1068EDRDREIQRLKHELQQKNLEDQRLKHKLQQKNLEDQRLKHELQQKNL 1208
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.355 0.157 0.539
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 66,260,795
Number of Sequences: 63676
Number of extensions: 925365
Number of successful extensions: 11624
Number of sequences better than 10.0: 51
Number of HSP's better than 10.0 without gapping: 6487
Number of HSP's successfully gapped in prelim test: 483
Number of HSP's that attempted gapping in prelim test: 4822
Number of HSP's gapped (non-prelim): 7490
length of query: 1574
length of database: 12,639,632
effective HSP length: 110
effective length of query: 1464
effective length of database: 5,635,272
effective search space: 8250038208
effective search space used: 8250038208
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.6 bits)
S2: 65 (29.6 bits)
Medicago: description of AC144538.13


