

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144538.1 - phase: 0 /pseudo
(942 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
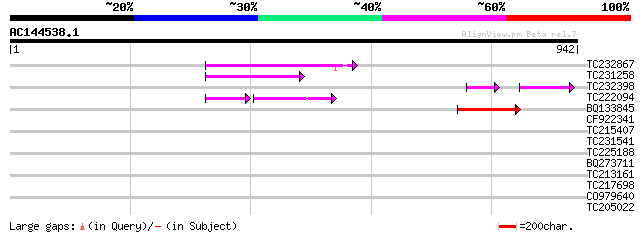
Sequences producing significant alignments: (bits) Value
TC232867 similar to UP|Q9MA68 (Q9MA68) F2J6.13 protein, partial ... 121 1e-27
TC231258 similar to UP|Q886T6 (Q886T6) Sensory box histidine kin... 119 8e-27
TC232398 75 2e-22
TC222094 61 8e-21
BQ133845 90 5e-18
CF922341 36 0.070
TC215407 weakly similar to GB|CAD29448.2|28804246|MMU440757 IL-1... 31 2.2
TC231541 similar to UP|Q8LCH9 (Q8LCH9) Thioredoxin-like protein,... 26 2.8
TC225188 similar to UP|Q9FYB7 (Q9FYB7) Splicing factor-like prot... 30 3.8
BQ273711 30 3.8
TC213161 30 5.0
TC217698 similar to UP|Q89GY6 (Q89GY6) Blr6209 protein, partial ... 30 5.0
CO979640 30 6.5
TC205022 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, pa... 30 6.5
>TC232867 similar to UP|Q9MA68 (Q9MA68) F2J6.13 protein, partial (3%)
Length = 1144
Score = 121 bits (304), Expect = 1e-27
Identities = 86/264 (32%), Positives = 134/264 (50%), Gaps = 11/264 (4%)
Frame = -3
Query: 326 FSGSPTEDPNLHISSFLRLSGTIK---ENQEAVRLHLFPFSLRDRASAWFHSLEVGSITS 382
F G P EDP H+++++ + TI+ +A+RL L FSL A W HS + S+ S
Sbjct: 782 FHGLPNEDPYAHLATYIEICNTIRLAGVPADAIRLSLLSFSLSGEAKRWLHSFKGNSLKS 603
Query: 383 WDDMRRAFLARFSPPSKTAKLRDQITRFNQKDGESLYEA*ERFKEMLRLCPHHGLEKWLI 442
WD++ FL ++ P SKTA+ + I+ F+Q ESL EA ERF+ +LR P HG + +
Sbjct: 602 WDEVVEKFLKKYFPESKTAEGKAAISSFHQFPDESLSEALERFRGLLRKTPTHGFSEPIQ 423
Query: 443 VHTFYNGLSYTTKMSVDAAAGGALMNKNYTEAYALIEDMAQNHYQWTNERAVTTPTPSKK 502
++ F + L +K VDA+ GG + K EA LIE MA + +RA S
Sbjct: 422 LNIFIDELRPESKQLVDASVGGKIKMKTPDEAMDLIESMAASDIAILRDRAHIPTKKSLL 243
Query: 503 EAGIYE--VSEYNHLAAKVEALTQKIEKL--NVNAAQPSPAS----PTCEVCGITGHTGV 554
E + +++ L+ ++E LT+ + KL +++AQ S +S C + G +G
Sbjct: 242 ELTSQDTLLAQNKLLSKQLETLTKTLSKLPTQLHSAQTSHSSILQVTGCTIFGEAHESGC 63
Query: 555 DCQLGSAANIEQLNYAQYNQGMRP 578
N EQ+ + G RP
Sbjct: 62 -----CIPNEEQIAHEVNYMGNRP 6
>TC231258 similar to UP|Q886T6 (Q886T6) Sensory box histidine kinase/response
regulator, partial (3%)
Length = 1374
Score = 119 bits (297), Expect = 8e-27
Identities = 67/167 (40%), Positives = 93/167 (55%), Gaps = 3/167 (1%)
Frame = +2
Query: 326 FSGSPTEDPNLHISSFLRLSGTIKE---NQEAVRLHLFPFSLRDRASAWFHSLEVGSITS 382
F G EDP+ H+ F + T+K ++ + L FP SL A W + L SI +
Sbjct: 554 FHGLVGEDPHKHLKEFHIVCSTMKPPDVQEDHIFLKAFPHSLEGVAKDWLYYLAPRSIFN 733
Query: 383 WDDMRRAFLARFSPPSKTAKLRDQITRFNQKDGESLYEA*ERFKEMLRLCPHHGLEKWLI 442
WDD++R FL +F P S+T +R I+ Q ESLYE ERFK+ CPHH + K L+
Sbjct: 734 WDDLKRVFLEKFFPASRTTTIRKDISGIRQLSRESLYEYWERFKKSCASCPHHQISKQLL 913
Query: 443 VHTFYNGLSYTTKMSVDAAAGGALMNKNYTEAYALIEDMAQNHYQWT 489
+ FY LS + +DAA+GGAL + TEA LI+ MA N Q++
Sbjct: 914 L*YFYEELSNMKRSMIDAASGGALGDMTPTEARNLIKKMASNSQQFS 1054
>TC232398
Length = 1054
Score = 74.7 bits (182), Expect(2) = 2e-22
Identities = 36/92 (39%), Positives = 55/92 (59%)
Frame = +1
Query: 847 KRMGIGELKPTEMILKLADRSTIPLAGYIEDIPVKIEGIYIPTDFVVVDIEEDHDVPIIL 906
+R+G +++PT M L+LAD S G +EDI VK+ + DFV++DIEED ++ +IL
Sbjct: 172 RRIGNQKIEPTRMTLQLADHSITRSFGVVEDILVKVHQLIFLVDFVIMDIEEDAEIRLIL 351
Query: 907 GRPFLATAGAIIDVQSGRIVFQASDAMIGFEL 938
G PF+ TA ++D+ G + D F L
Sbjct: 352 GWPFMVTAKCVVDMGKGNLEMSVEDQKATFNL 447
Score = 50.1 bits (118), Expect(2) = 2e-22
Identities = 24/56 (42%), Positives = 34/56 (59%)
Frame = +2
Query: 759 IKIPFAEALSRMPLYAKFLREIFSKKKAIDHKETIALTRESSAIVKKQPQKLRDPG 814
I IPF E + +MPLY KFL++I KK + ETI + A+++K P K +D G
Sbjct: 8 ITIPFGERIQQMPLYKKFLKDILIKKGKYINSETIVVGEYCRALIQKLPPKFKDLG 175
>TC222094
Length = 984
Score = 60.8 bits (146), Expect(2) = 8e-21
Identities = 43/139 (30%), Positives = 69/139 (48%), Gaps = 2/139 (1%)
Frame = -3
Query: 406 QITRFNQKDGESLYEA*ERFKEMLRLCPHHGLEKWLIVHTFYNGLSYTTKMSVDAAAGGA 465
+I+ F+Q ESL EA +RF +L P HG + + ++ F +G+ +K +DA+AGG
Sbjct: 472 EISSFHQHPHESLSEALDRFHGLLWKTPTHGFSEPVQLNIFIDGMQPHSKQLLDASAGGK 293
Query: 466 LMNKNYTEAYALIEDMAQNHYQWTNERAVTTPTPSKKEAGIYE--VSEYNHLAAKVEALT 523
+ K EA LIE+MA N Y ++ P+P ++ + E V + ++
Sbjct: 292 IKLKTPEEAIELIENMAANDYVILRDQ---EPSPQEESTRLEELLVQFMQETRSHQKSTD 122
Query: 524 QKIEKLNVNAAQPSPASPT 542
I L V A+ P PT
Sbjct: 121 AAIRNLEVQLAKLVPERPT 65
Score = 58.9 bits (141), Expect(2) = 8e-21
Identities = 27/78 (34%), Positives = 45/78 (57%), Gaps = 3/78 (3%)
Frame = -2
Query: 326 FSGSPTEDPNLHISSFLRLSGTIK---ENQEAVRLHLFPFSLRDRASAWFHSLEVGSITS 382
F G PTEDP H+++++ + T+K ++A+ L LF FSL A W S + ++ +
Sbjct: 722 FYGLPTEDPYAHLATYIDICNTVKIVGVPEDAIHLDLFCFSLAGEARTWLRSFKGNNLRT 543
Query: 383 WDDMRRAFLARFSPPSKT 400
W++ FL ++ P SKT
Sbjct: 542 WNEXXEKFLKKYFPESKT 489
>BQ133845
Length = 389
Score = 89.7 bits (221), Expect = 5e-18
Identities = 46/106 (43%), Positives = 69/106 (64%), Gaps = 1/106 (0%)
Frame = +2
Query: 744 EAQFAKFLNILKRICIKIPFAEALSRMPLYAKFLREIFSKKKAIDHKETIALTRESSAIV 803
E AKFL+I K++ I +PF EAL +MPLYA FL+++ +KK H + I + SA++
Sbjct: 71 EQHLAKFLDIFKKLEITLPFEEALQQMPLYANFLKDMLTKKNWYIHSDKIVVEGNCSAVI 250
Query: 804 KK-QPQKLRDPGSFAIPCVIGKETVDKALCHLGASVSLLPLSLFKR 848
++ P DPG +PC IG+ V KAL LGAS++L+PLS+ ++
Sbjct: 251 QRILPP*HTDPGFVTMPCSIGEVAVGKALIDLGASINLMPLSMCRQ 388
>CF922341
Length = 675
Score = 36.2 bits (82), Expect = 0.070
Identities = 18/64 (28%), Positives = 33/64 (51%)
Frame = +1
Query: 331 TEDPNLHISSFLRLSGTIKENQEAVRLHLFPFSLRDRASAWFHSLEVGSITSWDDMRRAF 390
T P H+ + + G +++E + +H F SL A W+ +LE + SW D+ AF
Sbjct: 448 TTCPKNHLKMYCQKMGAYAKDEELL-IHSFQESLTGVAVTWYTNLEPSRVHSWKDLMVAF 624
Query: 391 LARF 394
+ ++
Sbjct: 625 VRQY 636
>TC215407 weakly similar to GB|CAD29448.2|28804246|MMU440757 IL-1
receptor-associated kinase M {Mus musculus;} , partial
(6%)
Length = 1177
Score = 31.2 bits (69), Expect = 2.2
Identities = 30/123 (24%), Positives = 52/123 (41%), Gaps = 15/123 (12%)
Frame = +2
Query: 597 GYTNNQRVAQKSSLEILLENCMMNQNKQLQELKNQTGSLNDSLSKLNT-KVDSI----AT 651
G +NN + S+ +L C +NQ + L EL ++ L L+K+ KV I +
Sbjct: 401 GKSNNNYLPDDGSVCLLDRACQLNQTRNLMELIDE--RLGPDLNKMEVEKVVKIGLLCSN 574
Query: 652 HTKMLETQISQVAQQVAISSQTPGVFPGQTETNPKAHVNAI----------SLGGNKLEE 701
+ L +S+V + + P V P + N A+ SL GN+ +
Sbjct: 575 ASPTLRPTMSEVVNMLEGHADIPDVIPEPSTYNDDLRFKALRNLHQYQSKQSLSGNQSQS 754
Query: 702 TIT 704
++T
Sbjct: 755 SMT 763
>TC231541 similar to UP|Q8LCH9 (Q8LCH9) Thioredoxin-like protein, partial
(83%)
Length = 536
Score = 25.8 bits (55), Expect(2) = 2.8
Identities = 11/32 (34%), Positives = 19/32 (59%)
Frame = -3
Query: 907 GRPFLATAGAIIDVQSGRIVFQASDAMIGFEL 938
G P+L TAG + D+ R++ ++S + F L
Sbjct: 192 GAPYLMTAGVLEDLMCARMLVKSSSLVALFRL 97
Score = 23.5 bits (49), Expect(2) = 2.8
Identities = 14/35 (40%), Positives = 21/35 (60%)
Frame = -2
Query: 854 LKPTEMILKLADRSTIPLAGYIEDIPVKIEGIYIP 888
++P ++ L STI L Y+ IPVK+EG +P
Sbjct: 412 VQPQTIMEPLFPTSTIHLI-YLFTIPVKLEGGSVP 311
>TC225188 similar to UP|Q9FYB7 (Q9FYB7) Splicing factor-like protein, partial
(62%)
Length = 1131
Score = 30.4 bits (67), Expect = 3.8
Identities = 13/28 (46%), Positives = 14/28 (49%)
Frame = +1
Query: 536 PSPASPTCEVCGITGHTGVDCQLGSAAN 563
P P S C CGI GH DC+ G N
Sbjct: 343 PPPGSGRCFNCGIDGHWARDCKAGDWKN 426
>BQ273711
Length = 409
Score = 30.4 bits (67), Expect = 3.8
Identities = 10/29 (34%), Positives = 16/29 (54%)
Frame = -2
Query: 17 WWVSLLPTLEQDGAVVTWALFRREFLDRY 45
WW + P+ G V+ W F+ +FL+ Y
Sbjct: 87 WWKFVKPSFVAPGGVIPWNAFKDKFLENY 1
>TC213161
Length = 502
Score = 30.0 bits (66), Expect = 5.0
Identities = 17/62 (27%), Positives = 25/62 (39%), Gaps = 3/62 (4%)
Frame = +1
Query: 433 PHHGLEKWLIVHTFYNGLSYTTKMSVDAAAGGALMNKN---YTEAYALIEDMAQNHYQWT 489
PH+ L WL V +N ++T A G N N E +++D+ W
Sbjct: 109 PHNNLSSWLDVSEDHNQKQFSTLSPTITACGSFSNNGNGRFLREESGVVDDVISPDLAWV 288
Query: 490 NE 491
NE
Sbjct: 289 NE 294
>TC217698 similar to UP|Q89GY6 (Q89GY6) Blr6209 protein, partial (5%)
Length = 865
Score = 30.0 bits (66), Expect = 5.0
Identities = 18/46 (39%), Positives = 22/46 (47%), Gaps = 5/46 (10%)
Frame = +2
Query: 566 QLNYAQYNQGMRPNQNF-----YKNPQGSYGQTAPPGYTNNQRVAQ 606
Q NY Q +Q P QN+ PQ +YGQ A P Y Q+ Q
Sbjct: 44 QXNYGQASQNYPPQQNYGMTSQNYTPQQNYGQ-ASPNYPPQQKYGQ 178
>CO979640
Length = 722
Score = 29.6 bits (65), Expect = 6.5
Identities = 23/79 (29%), Positives = 35/79 (44%), Gaps = 4/79 (5%)
Frame = +1
Query: 799 SSAIVKKQPQKLRD---PGSFAIPCVIGKETVDKALCHLGASVSLLPLSLFK-RMGIGEL 854
S+ V+KQ QKL P F + T+ CH+ +S++ LPL++ R+ I L
Sbjct: 343 STTAVEKQNQKLHSLHCPSMFCMWSATMSATISGENCHIMSSLAFLPLAVASVRLSITCL 522
Query: 855 KPTEMILKLADRSTIPLAG 873
T + PL G
Sbjct: 523 IRTRSLSSEGSAQRCPLNG 579
>TC205022 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, partial (69%)
Length = 1623
Score = 29.6 bits (65), Expect = 6.5
Identities = 12/28 (42%), Positives = 14/28 (49%)
Frame = +2
Query: 536 PSPASPTCEVCGITGHTGVDCQLGSAAN 563
P P S C CG+ GH DC+ G N
Sbjct: 293 PPPGSGRCFNCGLDGHWARDCKAGDWKN 376
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.332 0.142 0.431
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 38,773,410
Number of Sequences: 63676
Number of extensions: 529594
Number of successful extensions: 3149
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 3098
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3145
length of query: 942
length of database: 12,639,632
effective HSP length: 106
effective length of query: 836
effective length of database: 5,889,976
effective search space: 4924019936
effective search space used: 4924019936
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.9 bits)
S2: 63 (28.9 bits)
Medicago: description of AC144538.1


