

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
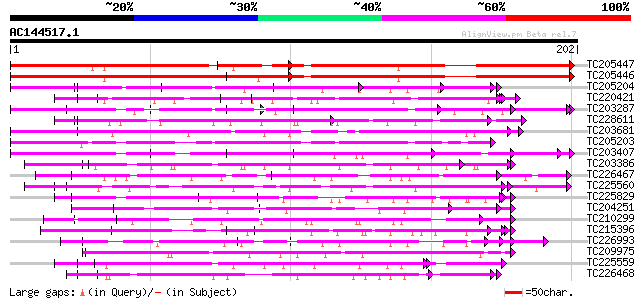
Query= AC144517.1 + phase: 0
(202 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC205447 weakly similar to UP|Q9SXQ2 (Q9SXQ2) Polyprotein, parti... 112 1e-25
TC205446 weakly similar to UP|Q8LCN5 (Q8LCN5) Arabinogalactan-pr... 110 3e-25
TC205204 homologue to UP|Q43558 (Q43558) Proline rich protein pr... 94 3e-20
TC220421 weakly similar to UP|GP1_CHLRE (Q9FPQ6) Vegetative cell... 91 5e-19
TC203287 weakly similar to UP|Q41645 (Q41645) Extensin (Fragment... 87 4e-18
TC228611 weakly similar to GB|CAA72487.1|1890317|ATP26A peroxida... 87 5e-18
TC203681 similar to UP|Q9XIV1 (Q9XIV1) Cucumis sativus mRNA expr... 87 5e-18
TC205203 similar to UP|Q39763 (Q39763) Proline-rich cell wall pr... 86 9e-18
TC203407 weakly similar to UP|Q9ZT16 (Q9ZT16) Arabinogalactan-pr... 84 4e-17
TC203386 similar to UP|Q9XIV1 (Q9XIV1) Cucumis sativus mRNA expr... 83 1e-16
TC226467 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 pr... 82 2e-16
TC225560 similar to UP|Q9FUL8 (Q9FUL8) Arabinogalactan protein, ... 81 4e-16
TC225829 weakly similar to UP|Q9C5Q6 (Q9C5Q6) Fasciclin-like ara... 80 6e-16
TC204251 weakly similar to UP|O36027 (O36027) Wiskott-Aldrich Sy... 80 6e-16
TC210299 homologue to UP|EFT2_SOYBN (P46280) Elongation factor T... 80 6e-16
TC215396 weakly similar to UP|Q8H9F4 (Q8H9F4) 41 kD chloroplast ... 80 8e-16
TC226993 homologue to UP|GMD1_ARATH (Q9SNY3) GDP-mannose 4,6 deh... 79 1e-15
TC209975 weakly similar to UP|Q7T9Y7 (Q7T9Y7) Odv-e66, partial (9%) 79 1e-15
TC225559 similar to UP|Q9XIV1 (Q9XIV1) Cucumis sativus mRNA expr... 79 1e-15
TC226468 weakly similar to UP|O65450 (O65450) Glycine-rich prote... 77 4e-15
>TC205447 weakly similar to UP|Q9SXQ2 (Q9SXQ2) Polyprotein, partial (3%)
Length = 790
Score = 112 bits (280), Expect = 1e-25
Identities = 65/130 (50%), Positives = 85/130 (65%), Gaps = 3/130 (2%)
Frame = +2
Query: 75 SPAS-PVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSP 133
SPAS P +P TP++ P ++PSPAI+PS A SP +S P PV ++PSPSP++++SP
Sbjct: 104 SPASSPALSPKRTPVAT---PQKSPSPAISPS--AVSPSSSPPAPVVNTPSPSPNSVDSP 268
Query: 134 PSPP--PHASPAAAPAITPSSISTPPTKAPSSISTPPTKAPSTPTNGATMNGFNVAGYAA 191
PSPP P SPA P++TPS+IS PPT AP+ NGA +N F VAG AA
Sbjct: 269 PSPPQSPSVSPAGVPSVTPSAISA-----------PPTGAPAPSQNGAALNRFTVAGSAA 415
Query: 192 AVVFIAALIL 201
VVF AAL++
Sbjct: 416 VVVFAAALLM 445
Score = 92.4 bits (228), Expect = 1e-19
Identities = 56/113 (49%), Positives = 72/113 (63%), Gaps = 12/113 (10%)
Frame = +2
Query: 1 MASYMVVLMLVASLLVSSTFARSLSSSP--------VATP----SPAGSPLAISPAVSSP 48
MA VVL LVA+LLV+S A+S +SSP VATP SPA SP A+SP+ S P
Sbjct: 38 MACTTVVLTLVAALLVTSVVAQSPASSPALSPKRTPVATPQKSPSPAISPSAVSPSSSPP 217
Query: 49 VPLMNAPSPSPFAVNYPTSPPPASFGSPASPVAAPAVTPLSISTPPSQAPSPA 101
P++N PSPSP +V+ P SPP + SP P+VTP +IS PP+ AP+P+
Sbjct: 218 APVVNTPSPSPNSVDSPPSPPQS---PSVSPAGVPSVTPSAISAPPTGAPAPS 367
>TC205446 weakly similar to UP|Q8LCN5 (Q8LCN5) Arabinogalactan-protein,
partial (39%)
Length = 668
Score = 110 bits (276), Expect = 3e-25
Identities = 60/128 (46%), Positives = 85/128 (65%), Gaps = 4/128 (3%)
Frame = +1
Query: 78 SPVAAPAVTP--LSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSPPS 135
SP ++PA++P + P ++PSPAI+PS A SP +S P P ++PSPSP++++SPPS
Sbjct: 124 SPASSPALSPKRTPVLATPQKSPSPAISPS--AVSPSSSPPAPTVNAPSPSPTSVDSPPS 297
Query: 136 PP--PHASPAAAPAITPSSISTPPTKAPSSISTPPTKAPSTPTNGATMNGFNVAGYAAAV 193
PP P SPA P++TPS+IS+ PP++AP+ NGA +N F VAG AA V
Sbjct: 298 PPLSPSDSPAGVPSVTPSAISS-----------PPSEAPAPSQNGAALNRFTVAGSAAVV 444
Query: 194 VFIAALIL 201
VF AAL++
Sbjct: 445 VFAAALLM 468
Score = 93.2 bits (230), Expect = 7e-20
Identities = 54/114 (47%), Positives = 72/114 (62%), Gaps = 13/114 (11%)
Frame = +1
Query: 1 MASYMVVLMLVASLLVSSTFARSLSSSPVATP-------------SPAGSPLAISPAVSS 47
MAS V+ LVA+L V+S A+S +SSP +P SPA SP A+SP+ S
Sbjct: 58 MASSTVLFTLVAALFVTSVVAQSPASSPALSPKRTPVLATPQKSPSPAISPSAVSPSSSP 237
Query: 48 PVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVAAPAVTPLSISTPPSQAPSPA 101
P P +NAPSPSP +V+ P SPP + SPA P+VTP +IS+PPS+AP+P+
Sbjct: 238 PAPTVNAPSPSPTSVDSPPSPPLSPSDSPA---GVPSVTPSAISSPPSEAPAPS 390
>TC205204 homologue to UP|Q43558 (Q43558) Proline rich protein precursor,
partial (44%)
Length = 996
Score = 94.4 bits (233), Expect = 3e-20
Identities = 61/173 (35%), Positives = 91/173 (52%)
Frame = +1
Query: 1 MASYMVVLMLVASLLVSSTFARSLSSSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPF 60
M +++V+L+ + +V+ A+ S+SP A P+P SP P SSP P+ ++P P+
Sbjct: 115 METHLVLLLALIFTVVAGVGAQGPSTSP-APPTPQSSP---PPIQSSPPPVPSSPPPA-- 276
Query: 61 AVNYPTSPPPASFGSPASPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVT 120
SPPPAS PA P+++P S+PP +P PA P SP +SP P +
Sbjct: 277 -----QSPPPASTPPPA-PLSSPPPASPPPSSPPPASPPPASPPPA---SPPPASPPPAS 429
Query: 121 SSPSPSPSAINSPPSPPPHASPAAAPAITPSSISTPPTKAPSSISTPPTKAPS 173
P+ P A P SPPP + P A TP + PP P+ +S+PP +P+
Sbjct: 430 PPPASPPPATPPPASPPPFSPPPA----TPPPATPPPALTPTPLSSPPATSPA 576
Score = 69.3 bits (168), Expect = 1e-12
Identities = 53/133 (39%), Positives = 69/133 (51%), Gaps = 1/133 (0%)
Frame = +1
Query: 24 LSSSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVAAP 83
LSS P A+P P+ SP +SP P P+ P P SPPPAS PASP P
Sbjct: 316 LSSPPPASPPPS------SPPPASPPPASPPPASPP-----PASPPPAS-PPPASP---P 450
Query: 84 AVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSPPSPPPHASPA 143
TP S PP +P PA P A P A +P P++S P+ SP+ P+P +PA
Sbjct: 451 PATPPPASPPPF-SPPPATPP--PATPPPALTPTPLSSPPATSPA-----PAPAKVVAPA 606
Query: 144 AAPAITPS-SIST 155
+P++ P S+ST
Sbjct: 607 LSPSLAPGPSLST 645
Score = 66.6 bits (161), Expect = 7e-12
Identities = 44/105 (41%), Positives = 58/105 (54%), Gaps = 3/105 (2%)
Frame = +1
Query: 25 SSSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVAA-- 82
SS P A+P PA P A SP +SP P P+ P A P SPPP S PA+P A
Sbjct: 349 SSPPPASPPPASPPPA-SPPPASPPPASPPPASPPPATPPPASPPPFS-PPPATPPPATP 522
Query: 83 -PAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPS 126
PA+TP +S+PP+ +P+PA A + + +P P S+ SPS
Sbjct: 523 PPALTPTPLSSPPATSPAPAPAKVVAPALSPSLAPGPSLSTISPS 657
Score = 65.9 bits (159), Expect = 1e-11
Identities = 40/121 (33%), Positives = 53/121 (43%)
Frame = +1
Query: 55 PSPSPFAVNYPTSPPPASFGSPASPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVAS 114
PS SP +SPPP P P+ S PP+Q+P PA P
Sbjct: 184 PSTSPAPPTPQSSPPPIQSSPP----------PVPSSPPPAQSPPPASTPP--------- 306
Query: 115 SPVPVTSSPSPSPSAINSPPSPPPHASPAAAPAITPSSISTPPTKAPSSISTPPTKAPST 174
P P++S P SP + PP+ PP PA+ P +P S PP P + P T P++
Sbjct: 307 -PAPLSSPPPASPPPSSPPPASPP---PASPPPASPPPASPPPASPPPASPPPATPPPAS 474
Query: 175 P 175
P
Sbjct: 475 P 477
Score = 60.1 bits (144), Expect = 7e-10
Identities = 37/96 (38%), Positives = 51/96 (52%), Gaps = 15/96 (15%)
Frame = +1
Query: 95 SQAPSPAIAPSTSANS--PVASSPVPVTSSP----------SPSPSAINSPP--SPPPHA 140
+Q PS + AP T +S P+ SSP PV SSP +P P+ ++SPP SPPP +
Sbjct: 175 AQGPSTSPAPPTPQSSPPPIQSSPPPVPSSPPPAQSPPPASTPPPAPLSSPPPASPPPSS 354
Query: 141 -SPAAAPAITPSSISTPPTKAPSSISTPPTKAPSTP 175
PA+ P +P S PP P + P + P+TP
Sbjct: 355 PPPASPPPASPPPASPPPASPPPASPPPASPPPATP 462
>TC220421 weakly similar to UP|GP1_CHLRE (Q9FPQ6) Vegetative cell wall
protein gp1 precursor (Hydroxyproline-rich glycoprotein
1), partial (13%)
Length = 1409
Score = 90.5 bits (223), Expect = 5e-19
Identities = 62/164 (37%), Positives = 92/164 (55%), Gaps = 6/164 (3%)
Frame = +3
Query: 25 SSSPVATPSPAGSPLAISPAVSSPVP-LMNAPSPSPFAVNYPTSPPPASFGSPASPVA-- 81
+S P TP+P+ P + P+ + P P ++PS SP A PTSP P+ SP+ P A
Sbjct: 210 TSPPHRTPAPS-PPSTMPPSATPPSPSATSSPSASPTAP--PTSPSPSPPSSPSPPSAPS 380
Query: 82 --APAVTPLSISTPPSQAPSPAIAPST-SANSPVASSPVPVTSSPSPSPSAINSPPSPPP 138
+ A +P S S P S +PSP ST SANSP + SP TS SPSP++ + PP+P
Sbjct: 381 PSSTAPSPPSASPPTSPSPSPPSPASTASANSPASGSPTSRTSPTSPSPTSTSKPPAPS- 557
Query: 139 HASPAAAPAITPSSISTPPTKAPSSISTPPTKAPSTPTNGATMN 182
+ +P+ S+ P+ +P+ IS P+ A STPT+ +++
Sbjct: 558 --------SSSPA*PSSKPSPSPTPISPDPSPATSTPTSHTSIS 665
Score = 82.0 bits (201), Expect = 2e-16
Identities = 51/144 (35%), Positives = 77/144 (53%)
Frame = +3
Query: 32 PSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVAAPAVTPLSIS 91
P P + P S P P+PSP + P++ PP+ + +SP A+P P S S
Sbjct: 162 PHPLWTQNKQQPLNPSTSPPHRTPAPSPPSTMPPSATPPSP-SATSSPSASPTAPPTSPS 338
Query: 92 TPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSPPSPPPHASPAAAPAITPS 151
P +PSP APS P +++P P ++SP SPS SPPSP AS + + +P+
Sbjct: 339 PSPPSSPSPPSAPS-----PSSTAPSPPSASPPTSPSP--SPPSPASTASANSPASGSPT 497
Query: 152 SISTPPTKAPSSISTPPTKAPSTP 175
S ++P + +P+S S PP + S+P
Sbjct: 498 SRTSPTSPSPTSTSKPPAPSSSSP 569
Score = 71.2 bits (173), Expect = 3e-13
Identities = 58/166 (34%), Positives = 84/166 (49%), Gaps = 6/166 (3%)
Frame = +3
Query: 17 SSTFARSLSSSPVATP-SPAGSPLAISPAVSSPVPLMNAPSP---SPFAVNYPTSPPPAS 72
S + S S+SP A P SP+ SP + S+P P APSP SP P+ P PAS
Sbjct: 279 SPSATSSPSASPTAPPTSPSPSPPSSPSPPSAPSPSSTAPSPPSASPPTSPSPSPPSPAS 458
Query: 73 FGSPASPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINS 132
S SP + +P S ++P S +P+ P +P +SSP +S PSPSP+ I+
Sbjct: 459 TASANSPASG---SPTSRTSPTSPSPTSTSKPP----APSSSSPA*PSSKPSPSPTPISP 617
Query: 133 PPSPPPHASPAAAPAITPSSISTPPTKAPSSISTPPT--KAPSTPT 176
PSP ++P + +I+P + S P ST + +P TP+
Sbjct: 618 DPSPAT-STPTSHTSISPITASKATFLLP*LCSTASSF*ISPLTPS 752
Score = 62.8 bits (151), Expect = 1e-10
Identities = 41/97 (42%), Positives = 56/97 (57%), Gaps = 1/97 (1%)
Frame = +3
Query: 87 PLSIST-PPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSPPSPPPHASPAAA 145
PL+ ST PP + P+P+ PST S SP TSSPS SP+A + PSP P +SP+
Sbjct: 195 PLNPSTSPPHRTPAPS-PPSTMPPSATPPSP-SATSSPSASPTAPPTSPSPSPPSSPS-- 362
Query: 146 PAITPSSISTPPTKAPSSISTPPTKAPSTPTNGATMN 182
P PS ST P+ +S T P+ +P +P + A+ N
Sbjct: 363 PPSAPSPSSTAPSPPSASPPTSPSPSPPSPASTASAN 473
Score = 58.9 bits (141), Expect = 1e-09
Identities = 40/105 (38%), Positives = 59/105 (56%), Gaps = 3/105 (2%)
Frame = +3
Query: 76 PASPVAAPAV-TPLSISTPPSQAPSPAIAPSTSA-NSPVASSPVPVTSSPSPSPSAINSP 133
P +P +P TP +PPS P A PS SA +SP AS P T SPSPSP + SP
Sbjct: 195 PLNPSTSPPHRTPAP--SPPSTMPPSATPPSPSATSSPSASPTAPPT-SPSPSPPSSPSP 365
Query: 134 PS-PPPHASPAAAPAITPSSISTPPTKAPSSISTPPTKAPSTPTN 177
PS P P ++ + P+ +P + +P +P+S ++ + A +PT+
Sbjct: 366 PSAPSPSSTAPSPPSASPPTSPSPSPPSPASTASANSPASGSPTS 500
Score = 38.1 bits (87), Expect = 0.003
Identities = 40/131 (30%), Positives = 60/131 (45%), Gaps = 5/131 (3%)
Frame = +3
Query: 25 SSSPVATPSPAG-SPLAIS--PAVSSPVPL--MNAPSPSPFAVNYPTSPPPASFGSPASP 79
S SP + SP SP + S PA SS P + PSPSP P SP P+ + ++P
Sbjct: 483 SGSPTSRTSPTSPSPTSTSKPPAPSSSSPA*PSSKPSPSP----TPISPDPSP--ATSTP 644
Query: 80 VAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSPPSPPPH 139
+ +++P++ S P S+ SP+ SP PS + +S SP P
Sbjct: 645 TSHTSISPITASKATFLLP*LCSTASSF*ISPLTPSPEKYL---LPSVT*FHSKTSPSP- 812
Query: 140 ASPAAAPAITP 150
+P+ P + P
Sbjct: 813 PTPSPVPFLIP 845
Score = 29.6 bits (65), Expect = 0.97
Identities = 24/78 (30%), Positives = 36/78 (45%)
Frame = +2
Query: 10 LVASLLVSSTFARSLSSSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPP 69
+V LL+S T +SLS +PSP+ SP SS L + + ++N+PTS
Sbjct: 71 VVILLLLSPTIVKSLSLPHTPSPSPSPSP-------SSSSTLDPKQATALESLNFPTS*D 229
Query: 70 PASFGSPASPVAAPAVTP 87
P + S + A P
Sbjct: 230 PCAKPSFHNATLCDAAKP 283
Score = 26.9 bits (58), Expect = 6.3
Identities = 14/30 (46%), Positives = 17/30 (56%)
Frame = +2
Query: 112 VASSPVPVTSSPSPSPSAINSPPSPPPHAS 141
V S +P T SPSPSPS +S P A+
Sbjct: 104 VKSLSLPHTPSPSPSPSPSSSSTLDPKQAT 193
>TC203287 weakly similar to UP|Q41645 (Q41645) Extensin (Fragment), partial
(14%)
Length = 967
Score = 87.4 bits (215), Expect = 4e-18
Identities = 57/155 (36%), Positives = 83/155 (52%), Gaps = 1/155 (0%)
Frame = +3
Query: 1 MASYMVVLMLVASLLVSSTFARSLSSSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPF 60
M S + L+L+ LL SS A++ ++P P+ SP P S+P AP+P+P
Sbjct: 57 MGSGALQLLLILGLLASSCLAQAPGAAPTQPPTTTPSP---PPPRSAP-----APAPTPP 212
Query: 61 AVNYPTSPPPASFGSPASPVAAPAVTPLSISTP-PSQAPSPAIAPSTSANSPVASSPVPV 119
A P +PPPA+ +P + PA TP TP P+ AP+PA +P+TS +S P P
Sbjct: 213 ATPPPATPPPAA-ATPPPATSPPATTP----TPTPTAAPTPASSPATSPVPSASSPPAPG 377
Query: 120 TSSPSPSPSAINSPPSPPPHASPAAAPAITPSSIS 154
T+ P+P P+ PP PP A A+ I S++S
Sbjct: 378 TAGPAPGPAGGAEPPPPPSAAFSASKAFIAGSALS 482
Score = 68.9 bits (167), Expect = 1e-12
Identities = 46/150 (30%), Positives = 71/150 (46%)
Frame = +3
Query: 51 LMNAPSPSPFAVNYPTSPPPASFGSPASPVAAPAVTPLSISTPPSQAPSPAIAPSTSANS 110
L AP +P + PP + SP P +APA P +TPP P PA A A S
Sbjct: 114 LAQAPGAAP-------TQPPTTTPSPPPPRSAPAPAPTPPATPPPATPPPAAATPPPATS 272
Query: 111 PVASSPVPVTSSPSPSPSAINSPPSPPPHASPAAAPAITPSSISTPPTKAPSSISTPPTK 170
P A++P +P+P+A +P P +SPA +P + SS P T P+ +
Sbjct: 273 PPATTP-------TPTPTA-----APTPASSPATSPVPSASSPPAPGTAGPAPGPAGGAE 416
Query: 171 APSTPTNGATMNGFNVAGYAAAVVFIAALI 200
P P+ + + +AG A + +F+A ++
Sbjct: 417 PPPPPSAAFSASKAFIAGSALSGIFVAMVL 506
Score = 60.5 bits (145), Expect = 5e-10
Identities = 43/130 (33%), Positives = 59/130 (45%), Gaps = 6/130 (4%)
Frame = +3
Query: 78 SPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSPPSPP 137
+P AAP P + +PP +PA AP+ A P A+ P P ++P P+ S P
Sbjct: 123 APGAAPTQPPTTTPSPPPPRSAPAPAPTPPATPPPATPP-PAAATPPPATS--------P 275
Query: 138 PHASPAAAPAITPSSISTPPTKAPSSISTPPTKAPST------PTNGATMNGFNVAGYAA 191
P +P P P+ S+P T S S+PP AP T P GA A ++A
Sbjct: 276 PATTPTPTPTAAPTPASSPATSPVPSASSPP--APGTAGPAPGPAGGAEPPPPPSAAFSA 449
Query: 192 AVVFIAALIL 201
+ FIA L
Sbjct: 450 SKAFIAGSAL 479
Score = 52.4 bits (124), Expect = 1e-07
Identities = 32/82 (39%), Positives = 46/82 (56%), Gaps = 3/82 (3%)
Frame = +3
Query: 102 IAPSTSANSPVASSPVPVTSSPSPSPSAINSPPSPPPHASPAAAPAITPSSISTPP--TK 159
+A S A +P A+ P T++PSP P P+P P A+P PA P + +TPP T
Sbjct: 99 LASSCLAQAPGAAPTQPPTTTPSPPPPRSAPAPAPTPPATP--PPATPPPAAATPPPATS 272
Query: 160 APSSISTP-PTKAPSTPTNGAT 180
P++ TP PT AP+ ++ AT
Sbjct: 273 PPATTPTPTPTAAPTPASSPAT 338
Score = 41.2 bits (95), Expect = 3e-04
Identities = 25/74 (33%), Positives = 39/74 (51%), Gaps = 3/74 (4%)
Frame = +3
Query: 21 ARSLSSSPVATPSPAGSPLAISP---AVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPA 77
A + + +P A P+PA SP A SP A S P P P+P P P PP A+F +
Sbjct: 279 ATTPTPTPTAAPTPASSP-ATSPVPSASSPPAPGTAGPAPGPAGGAEPPPPPSAAFSASK 455
Query: 78 SPVAAPAVTPLSIS 91
+ +A A++ + ++
Sbjct: 456 AFIAGSALSGIFVA 497
>TC228611 weakly similar to GB|CAA72487.1|1890317|ATP26A peroxidase ATP26a
{Arabidopsis thaliana;} , partial (49%)
Length = 731
Score = 87.0 bits (214), Expect = 5e-18
Identities = 71/185 (38%), Positives = 93/185 (49%), Gaps = 25/185 (13%)
Frame = +3
Query: 25 SSSPVATPSP---------AGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPA---- 71
S SP P P A SP A++P SSP PSP P A TSP PA
Sbjct: 159 SRSPAPPPPPPRSASSSTTASSPTAVTPPSSSPP----RPSPGPSATPTSTSPSPATPST 326
Query: 72 SFGSPASPVAAPAVTPL---SISTPPSQAPSP-AIAPSTSANSPVASS------PVPVTS 121
S +P P+++PA TP + S PP SP + APS+ ++S A++ P P TS
Sbjct: 327 SSSAPRPPLSSPAPTPSPAPTFSPPPPATSSPCSAAPSSPSSSAAATAEPPSPPPYPPTS 506
Query: 122 SPSPSPSAINSPPSPPPHASPA--AAPAITPSSISTPPTKAPSSISTPPTKAPSTPTNGA 179
P P PS + SP S P ASP+ ++P++ P+ + APSS T PT P PT A
Sbjct: 507 PPPPCPS-LKSPNSSPNAASPSKNSSPSVGPT--PSDSLTAPSSSRTSPTTLP-PPTTLA 674
Query: 180 TMNGF 184
T+ GF
Sbjct: 675 TLRGF 689
Score = 68.9 bits (167), Expect = 1e-12
Identities = 54/157 (34%), Positives = 74/157 (46%), Gaps = 8/157 (5%)
Frame = +3
Query: 24 LSSSPVATPSPAGSP--LAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVA 81
L S+P SP S L S A S P + SP+P PPP S S + +
Sbjct: 63 LFSAPPTRDSPWTSTRTLVRSSARS*GTPSQASRSPAP------PPPPPRSASSSTTASS 224
Query: 82 APAVTPLSISTPPSQAPSPAIAPSTSANSPVA---SSPVPVTSSPSPSPSAINSP---PS 135
AVTP S S+PP +P P+ P++++ SP SS P SP+P+ +P P
Sbjct: 225 PTAVTPPS-SSPPRPSPGPSATPTSTSPSPATPSTSSSAPRPPLSSPAPTPSPAPTFSPP 401
Query: 136 PPPHASPAAAPAITPSSISTPPTKAPSSISTPPTKAP 172
PP +SP +A +PSS + + PS PPT P
Sbjct: 402 PPATSSPCSAAPSSPSSSAAATAEPPSPPPYPPTSPP 512
Score = 53.9 bits (128), Expect = 5e-08
Identities = 42/111 (37%), Positives = 53/111 (46%), Gaps = 11/111 (9%)
Frame = +3
Query: 17 SSTFARSLSSSPVATPSPAGSPLAISPAVSSPVPLM-NAPSPSPFAVNYPTSPPPASFGS 75
SS+ R SSP TPSPA + PA SSP ++PS S A P SPPP S
Sbjct: 327 SSSAPRPPLSSPAPTPSPAPTFSPPPPATSSPCSAAPSSPSSSAAATAEPPSPPPYPPTS 506
Query: 76 PA-------SPVAAPAVTPLSISTPPSQAPSPA---IAPSTSANSPVASSP 116
P SP ++P S ++ PS P+P+ APS+S SP P
Sbjct: 507 PPPPCPSLKSPNSSPNAASPSKNSSPSVGPTPSDSLTAPSSSRTSPTTLPP 659
>TC203681 similar to UP|Q9XIV1 (Q9XIV1) Cucumis sativus mRNA expressed in
cucumber hypocotyls, complete cds (Arabinogalactan
protein), partial (40%)
Length = 913
Score = 87.0 bits (214), Expect = 5e-18
Identities = 63/190 (33%), Positives = 94/190 (49%), Gaps = 7/190 (3%)
Frame = +3
Query: 1 MASYMVVLMLVASLLVSSTFARSLSSSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPF 60
+ S ++ ++VA++ S A +S P AT +P SP SP P+ + S SP
Sbjct: 27 LLSLAIIAIVVAAVGGQSPAASPTTSPPAATTTPFASPATAPSKPKSPAPVASPTSSSPP 206
Query: 61 AV--NYPTSPPPASFGSPASPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVP 118
A N T+ PPAS + ASP + A P ++TPP+ P A P A +P A +PV
Sbjct: 207 ASSPNAATATPPASSPTVASP-PSKAAAPAPVATPPAATPPAATPP---AATPPAVTPV- 371
Query: 119 VTSSPSPSPSAINSPPSPPPHASPAAAPAITPSSISTPPTKAP-----SSISTPPTKAPS 173
SSP P+P ++SPP+P P +SP A TP+ + P + P S T +K +
Sbjct: 372 --SSP-PAPVPVSSPPAPVPVSSPPALAPTTPAPVVAPSAEVPAPAPKSKKKTKKSKKHT 542
Query: 174 TPTNGATMNG 183
P ++ G
Sbjct: 543 APAPSPSLLG 572
Score = 70.1 bits (170), Expect = 6e-13
Identities = 58/168 (34%), Positives = 82/168 (48%), Gaps = 13/168 (7%)
Frame = +3
Query: 25 SSSPVATPSPA-GSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVAAP 83
SS P ++P+ A +P A SP V+SP AP+P P + PPA+ A+P P
Sbjct: 195 SSPPASSPNAATATPPASSPTVASPPSKAAAPAP---VATPPAATPPAATPPAATP---P 356
Query: 84 AVTPLS-------ISTPPSQAP---SPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSP 133
AVTP+S +S+PP+ P PA+AP+T A PV + P P+P+P +
Sbjct: 357 AVTPVSSPPAPVPVSSPPAPVPVSSPPALAPTTPA--PVVA---PSAEVPAPAPKSKKKT 521
Query: 134 PSPPPHASPAAAPAITPSSISTPPTKAPSSI--STPPTKAPSTPTNGA 179
H +PA +P++ PP AP S S P A S +GA
Sbjct: 522 KKSKKHTAPAPSPSLL--GPPAPPVGAPGSSQDSMSPGPAVSEDESGA 659
>TC205203 similar to UP|Q39763 (Q39763) Proline-rich cell wall protein,
partial (48%)
Length = 1076
Score = 86.3 bits (212), Expect = 9e-18
Identities = 65/175 (37%), Positives = 97/175 (55%), Gaps = 2/175 (1%)
Frame = +3
Query: 1 MASYMVVLMLVASLLVSSTFARSL--SSSPVATPSPAGSPLAISPAVSSPVPLMNAPSPS 58
M +++V L+ + LV+ A++ S+SP A P+P SP P SSP P+ ++P PS
Sbjct: 201 METHLVFLLGLIFTLVAGVGAQAQGPSTSP-APPTPQASP---PPVQSSPPPVPSSPPPS 368
Query: 59 PFAVNYPTSPPPASFGSPASPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVP 118
SPPPAS P SP+++P + S+PP +P PA P +SP +SP P
Sbjct: 369 Q-------SPPPAS-TPPPSPLSSPPPSSPPPSSPPPSSPPPASPP---PSSPPPASP-P 512
Query: 119 VTSSPSPSPSAINSPPSPPPHASPAAAPAITPSSISTPPTKAPSSISTPPTKAPS 173
+S P SP + PP+ PP A+P PA+ P S++ PT P +S+PP +P+
Sbjct: 513 PSSPPPSSPPPFSPPPATPPPATP--PPAVPPPSLT--PTVTP--LSSPPVSSPA 659
Score = 34.7 bits (78), Expect = 0.030
Identities = 26/89 (29%), Positives = 41/89 (45%), Gaps = 1/89 (1%)
Frame = +2
Query: 89 SISTPPSQAPSPAIAPSTSANSPVASSPVPV-TSSPSPSPSAINSPPSPPPHASPAAAPA 147
S S P + PS S + PV S+ + TS+ S S ++ +SP P +S +
Sbjct: 260 SPSPRPIHLSCASHTPSISTSRPVFSATCSLFTSTLSVSTTSFHSPSIPTQLSSSFLSST 439
Query: 148 ITPSSISTPPTKAPSSISTPPTKAPSTPT 176
+ PSS + P + S STP + P+
Sbjct: 440 LLPSSFFSTPRLSSSFFSTPRLSSSFFPS 526
>TC203407 weakly similar to UP|Q9ZT16 (Q9ZT16) Arabinogalactan-protein
(AtAGP4) (AT5g10430/F12B17_220), partial (50%)
Length = 966
Score = 84.0 bits (206), Expect = 4e-17
Identities = 46/147 (31%), Positives = 71/147 (48%)
Frame = +2
Query: 51 LMNAPSPSPFAVNYPTSPPPASFGSPASPVAAPAVTPLSISTPPSQAPSPAIAPSTSANS 110
L AP +P S PP + SP P +APA P + +TPP P PA P +A
Sbjct: 185 LAQAPGAAP-------SQPPTTTPSPPPPRSAPAPAPTTPATPPPATPPPAATPPPAATP 343
Query: 111 PVASSPVPVTSSPSPSPSAINSPPSPPPHASPAAAPAITPSSISTPPTKAPSSISTPPTK 170
P A++P P + P+ +P+ +SP + PP SP P+ T + P+ PS + PP
Sbjct: 344 PPAATPTPAPAPPTAAPTPASSPAASPPSPSPTVTPSPTSPNTPPGPSPGPSGSAEPPPP 523
Query: 171 APSTPTNGATMNGFNVAGYAAAVVFIA 197
+ + + A + +AG A+ +A
Sbjct: 524 SAAFSASKAFIATSALAGTFVAMALVA 604
Score = 80.1 bits (196), Expect = 6e-16
Identities = 56/157 (35%), Positives = 79/157 (49%), Gaps = 5/157 (3%)
Frame = +2
Query: 1 MASYMVVLMLVASLLVSSTFARSLSSSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPF 60
M S V L L+ LL SS A++ ++P P+ SP P S+P P P+ P
Sbjct: 128 MGSGAVQLFLILGLLASSCLAQAPGAAPSQPPTTTPSP---PPPRSAPAPAPTTPATPP- 295
Query: 61 AVNYPTSPPPASFGSPASPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVAS-SPVPV 119
P +PPPA+ PA+ PA TP PP+ AP+PA +P+ S SP + +P P
Sbjct: 296 ----PATPPPAATPPPAA-TPPPAATPTPAPAPPTAAPTPASSPAASPPSPSPTVTPSPT 460
Query: 120 TSS----PSPSPSAINSPPSPPPHASPAAAPAITPSS 152
+ + PSP PS P PPP A+ +A+ A +S
Sbjct: 461 SPNTPPGPSPGPSGSAEP--PPPSAAFSASKAFIATS 565
Score = 72.4 bits (176), Expect = 1e-13
Identities = 47/128 (36%), Positives = 69/128 (53%), Gaps = 4/128 (3%)
Frame = +2
Query: 78 SPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPS--PSPSAINSPPS 135
+P AAP+ P + +PP +PA AP+T A P A+ P T P+ P P+A +P
Sbjct: 194 APGAAPSQPPTTTPSPPPPRSAPAPAPTTPATPPPATPPPAATPPPAATPPPAATPTPAP 373
Query: 136 PPPHA--SPAAAPAITPSSISTPPTKAPSSISTPPTKAPSTPTNGATMNGFNVAGYAAAV 193
PP A +PA++PA +P S S T +P+S +TPP PS +G+ A ++A+
Sbjct: 374 APPTAAPTPASSPAASPPSPSPTVTPSPTSPNTPP--GPSPGPSGSAEPPPPSAAFSASK 547
Query: 194 VFIAALIL 201
FIA L
Sbjct: 548 AFIATSAL 571
Score = 53.5 bits (127), Expect = 6e-08
Identities = 31/79 (39%), Positives = 41/79 (51%)
Frame = +2
Query: 102 IAPSTSANSPVASSPVPVTSSPSPSPSAINSPPSPPPHASPAAAPAITPSSISTPPTKAP 161
+A S A +P A+ P T++PSP P S P+P P +PA P TP +TPP A
Sbjct: 170 LASSCLAQAPGAAPSQPPTTTPSPPPP--RSAPAPAP-TTPATPPPATPPPAATPPPAAT 340
Query: 162 SSISTPPTKAPSTPTNGAT 180
+ PT AP+ PT T
Sbjct: 341 PPPAATPTPAPAPPTAAPT 397
>TC203386 similar to UP|Q9XIV1 (Q9XIV1) Cucumis sativus mRNA expressed in
cucumber hypocotyls, complete cds (Arabinogalactan
protein), partial (40%)
Length = 649
Score = 82.8 bits (203), Expect = 1e-16
Identities = 53/143 (37%), Positives = 78/143 (54%), Gaps = 9/143 (6%)
Frame = +1
Query: 29 VATPSPAGSPLAISPAVSSPVPLMNAPSP-SPFAVNYPTSPPPASFGSPASPVAAPAVTP 87
V SPA +P S +SP + P P SP V PTS PPAS SP + A P V+
Sbjct: 61 VGGQSPAAAPTTSS---ASPATAPSKPKPKSPAPVASPTSSPPAS--SPNAATATPPVSS 225
Query: 88 LSISTPPSQAPSPAIAPSTSANSPVASSP--------VPVTSSPSPSPSAINSPPSPPPH 139
++++PPS+A +PA A + +P A++P PV+S P+P P ++SPP+P P
Sbjct: 226 PTVASPPSKAAAPAPATTPPVATPPAATPPAATPPAVTPVSSPPAPVP--VSSPPAPVPV 399
Query: 140 ASPAAAPAITPSSISTPPTKAPS 162
+SP A TP+ + P + P+
Sbjct: 400 SSPPALAPTTPAPVVAPSAEVPA 468
Score = 75.9 bits (185), Expect = 1e-14
Identities = 60/199 (30%), Positives = 100/199 (50%), Gaps = 24/199 (12%)
Frame = +1
Query: 6 VVLMLVASLLVSSTFARSLSSSPV-ATPSPAGSPLAISPAVSSPV--PLMNAPSPSPFAV 62
++ + + ++V++ +S +++P ++ SPA +P P +PV P + P+ SP
Sbjct: 22 LLCLAIIGIVVAAVGGQSPAAAPTTSSASPATAPSKPKPKSPAPVASPTSSPPASSP--- 192
Query: 63 NYPTSPPPASFGSPASPVA-----APAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPV 117
N T+ PP S + ASP + APA TP ++TPP+ P A P+ + PV+S P
Sbjct: 193 NAATATPPVSSPTVASPPSKAAAPAPATTP-PVATPPAATPPAATPPAVT---PVSSPPA 360
Query: 118 PVTSSPSPSPSAINSPP-----SPPPHASPAA-APAITPSSIS--------TPPTKAPSS 163
PV S P+P ++SPP +P P +P+A PA P S T P +P+
Sbjct: 361 PVPVSSPPAPVPVSSPPALAPTTPAPVVAPSAEVPAPAPKSKKKTKKSKKHTAPAPSPAL 540
Query: 164 IS--TPPTKAPSTPTNGAT 180
+ PP AP + + ++
Sbjct: 541 LGPPAPPVGAPGSSQDASS 597
Score = 73.9 bits (180), Expect = 4e-14
Identities = 58/171 (33%), Positives = 83/171 (47%), Gaps = 18/171 (10%)
Frame = +1
Query: 27 SPVATPSPAGSPLAISPAVSSPVPLMNAPS----PSPFAVNYPTSPPPASFGSPASPVAA 82
SP SP SP A SP ++ P +++P+ PS A P + PP + A+P AA
Sbjct: 142 SPAPVASPTSSPPASSPNAATATPPVSSPTVASPPSKAAAPAPATTPPVATPPAATPPAA 321
Query: 83 --PAVTPLS-------ISTPPSQAP---SPAIAPSTSANSPVASSPVPVTSSPSPSPSAI 130
PAVTP+S +S+PP+ P PA+AP+T A PV + P P+P+P +
Sbjct: 322 TPPAVTPVSSPPAPVPVSSPPAPVPVSSPPALAPTTPA--PVVA---PSAEVPAPAPKSK 486
Query: 131 NSPPSPPPHASPAAAPAITPSSISTPPTKAPSSI--STPPTKAPSTPTNGA 179
H +PA +PA+ PP AP S ++ P A S +GA
Sbjct: 487 KKTKKSKKHTAPAPSPALL--GPPAPPVGAPGSSQDASSPGPAVSEDESGA 633
>TC226467 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 protein
precursor, partial (6%)
Length = 1017
Score = 81.6 bits (200), Expect = 2e-16
Identities = 68/216 (31%), Positives = 94/216 (43%), Gaps = 25/216 (11%)
Frame = +3
Query: 10 LVASLLVSSTFARSLSSSPVATPSPAG-SPLA--ISPAVSSPVPLMNAPSPSPFAVN--- 63
+VAS +ST + + P PA +P A ++PA S P P P P V+
Sbjct: 147 VVASPPTTSTPPTTSQPPAIVAPKPAPVTPPAPKVAPASSPKAPPPQTPQPQPPKVSPVV 326
Query: 64 YPTSPPPASFG--SPASPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTS 121
PTSPPP S P+ P + P TPP+ AP+ A A + A +P PV
Sbjct: 327 TPTSPPPIPPPPVSTPPPLPPPKIAPTPAKTPPAPAPAKATPAPAPAPTKPAPTPAPVPP 506
Query: 122 SPSPSPSAINSPPSPPP--------------HASPAAAPAITPSSISTPPTKAPSSISTP 167
P+ +P+ + P+P P H +PA AP I S PP S T
Sbjct: 507 PPTLAPTPVVEVPAPAPSSHKHRRHRHRHRRHQAPAPAPTIVRKSPPAPPDSTAES-DTT 683
Query: 168 PTKAPSTPTNGATMN---GFNVAGYAAAVVFIAALI 200
P +PS N A+ N G N+ +A A + IA L+
Sbjct: 684 PAPSPSLNLNAASSNHQQGRNI--WATAALAIAVLL 785
Score = 70.9 bits (172), Expect = 4e-13
Identities = 57/161 (35%), Positives = 74/161 (45%), Gaps = 8/161 (4%)
Frame = +3
Query: 23 SLSSSPVATP--SPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASPV 80
S + +P P SP +P A SP SS PL P+ A TS PP + PA +
Sbjct: 39 SNAQAPATAPANSPPTTPAATSP--SSGTPLAVTLPPTVVASPPTTSTPPTTSQPPA--I 206
Query: 81 AAPAVTPLSISTPPSQAPSPAIAPSTSANSPVA---SSPVPVTSSPSPSPSAINSPPSPP 137
AP P+ TPP+ P +AP A+SP A +P P SP + + PP PP
Sbjct: 207 VAPKPAPV---TPPA----PKVAP---ASSPKAPPPQTPQPQPPKVSPVVTPTSPPPIPP 356
Query: 138 PHAS---PAAAPAITPSSISTPPTKAPSSISTPPTKAPSTP 175
P S P P I P+ TPP AP+ + P AP+ P
Sbjct: 357 PPVSTPPPLPPPKIAPTPAKTPPAPAPAKATPAPAPAPTKP 479
Score = 47.0 bits (110), Expect = 6e-06
Identities = 32/91 (35%), Positives = 45/91 (49%), Gaps = 9/91 (9%)
Frame = +3
Query: 94 PSQAPSPAIAPSTS------ANSPVASSPVPVTSSPSPSPSAINSPP---SPPPHASPAA 144
PS A +PA AP+ S A SP + +P+ VT P+ + SPP +PP + P A
Sbjct: 36 PSNAQAPATAPANSPPTTPAATSPSSGTPLAVTLPPT----VVASPPTTSTPPTTSQPPA 203
Query: 145 APAITPSSISTPPTKAPSSISTPPTKAPSTP 175
A P+ + TPP + S+P P TP
Sbjct: 204 IVAPKPAPV-TPPAPKVAPASSPKAPPPQTP 293
>TC225560 similar to UP|Q9FUL8 (Q9FUL8) Arabinogalactan protein, partial
(48%)
Length = 1030
Score = 80.9 bits (198), Expect = 4e-16
Identities = 61/178 (34%), Positives = 94/178 (52%), Gaps = 4/178 (2%)
Frame = +2
Query: 6 VVLMLVASLLVSSTFARSLSSSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYP 65
V+ + ++V+ +S +++P TP+ +P A +PA + P N+P+P V P
Sbjct: 86 VLSLAFICIVVAGVGGQSPAAAPSNTPA---TPAAATPAQAPSTP--NSPAP----VASP 238
Query: 66 TSPPPASFGSPAS--PVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSP 123
S PPAS PA+ P ++PA +P +T P+ AP+PA P ++ P +PVPV+S P
Sbjct: 239 KSSPPASSPKPATATPASSPAASP---TTTPA-APAPATKPPAASPPP---APVPVSSPP 397
Query: 124 SPSPSAINSPPSPPPHASPA--AAPAITPSSISTPPTKAPSSISTPPTKAPSTPTNGA 179
+P P ++SPP+P P A+P APA PS K + +P P P GA
Sbjct: 398 APVP--VSSPPAPVPVAAPTTPVAPAPAPSKHKKKGKKHGAPAPSPSLLGPPAPPTGA 565
Score = 74.3 bits (181), Expect = 3e-14
Identities = 54/166 (32%), Positives = 76/166 (45%), Gaps = 5/166 (3%)
Frame = +2
Query: 17 SSTFARSLSSSPVATPSPAGSPLAI-----SPAVSSPVPLMNAPSPSPFAVNYPTSPPPA 71
S+T A +++P PS SP + SP SSP P P+ SP A PT+ P A
Sbjct: 155 SNTPATPAAATPAQAPSTPNSPAPVASPKSSPPASSPKPATATPASSPAAS--PTTTPAA 328
Query: 72 SFGSPASPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAIN 131
+ P A+P P+ +S+PP AP P +P A PVA+ PV +P+PS
Sbjct: 329 PAPATKPPAASPPPAPVPVSSPP--APVPVSSPP--APVPVAAPTTPVAPAPAPS----K 484
Query: 132 SPPSPPPHASPAAAPAITPSSISTPPTKAPSSISTPPTKAPSTPTN 177
H +PA +P++ PPT AP + P+T N
Sbjct: 485 HKKKGKKHGAPAPSPSLL--GPPAPPTGAPGPSEDASSPGPATAAN 616
Score = 70.1 bits (170), Expect = 6e-13
Identities = 58/199 (29%), Positives = 88/199 (44%), Gaps = 19/199 (9%)
Frame = +2
Query: 21 ARSLSSSPVA--TPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPAS 78
A + + +P +P+P SP + SP SSP P P+ SP A PT+ P A +
Sbjct: 179 AATPAQAPSTPNSPAPVASPKS-SPPASSPKPATATPASSPAA--SPTTTPAAPAPATKP 349
Query: 79 PVAAPAVTPLSISTPPSQAP---SPAIAPSTSANSPVASSPVPVT--------SSPSPSP 127
P A+P P+ +S+PP+ P PA P + +PVA +P P +P+PSP
Sbjct: 350 PAASPPPAPVPVSSPPAPVPVSSPPAPVPVAAPTTPVAPAPAPSKHKKKGKKHGAPAPSP 529
Query: 128 SAINSPPSPPPHASPAAAPAITPSSISTPPTKAPSSISTPPTKAPSTPTNGATMNGFNV- 186
S + PP+P P AP + + S P A + S T G G+
Sbjct: 530 SLL-GPPAP-----PTGAPGPSEDASSPGPATAANDESGAETIMCLKKVLGGLGLGWATL 691
Query: 187 -----AGYAAAVVFIAALI 200
G+ A ++F+ +I
Sbjct: 692 VLVF*TGFVAVIIFLYCII 748
>TC225829 weakly similar to UP|Q9C5Q6 (Q9C5Q6) Fasciclin-like
arabinogalactan-protein 2, partial (57%)
Length = 1652
Score = 80.1 bits (196), Expect = 6e-16
Identities = 59/160 (36%), Positives = 80/160 (49%), Gaps = 6/160 (3%)
Frame = +2
Query: 23 SLSSSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPP----ASFGSPAS 78
SLSSS + P P P SP +SP + P+ P P+ P A+ SP+
Sbjct: 23 SLSSSSSS*PPPPSPP---SPPTTSPACWPHTPASPPSTTTSPSPTWPKRSTAARPSPSW 193
Query: 79 PVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSPPSPPP 138
P P P S ST PS PS +PSTS+++ TS+P S + +PPS PP
Sbjct: 194 PSTMPPCPPSSTSTSPSP-PSKTSSPSTSSST---------TSAPKSSTRSTTAPPSSPP 343
Query: 139 HASPAAAPAITPSSISTPPTKAPSSISTPP--TKAPSTPT 176
+ P A P P++ ST PT P+ ++PP T APSTP+
Sbjct: 344 CSRPPAPPPAPPAT-STSPTSRPARSASPPKTTTAPSTPS 460
Score = 57.0 bits (136), Expect = 6e-09
Identities = 40/117 (34%), Positives = 56/117 (47%), Gaps = 7/117 (5%)
Frame = +2
Query: 68 PPPASFGSPASPVAAPAVTPLSISTPPSQ--APSPAIAPSTSANSPVASSPVPVTSSPSP 125
PPP S P+ P +PA P + ++PPS +PSP ++A P S P + P
Sbjct: 50 PPPPS--PPSPPTTSPACWPHTPASPPSTTTSPSPTWPKRSTAARPSPSWPSTMPPCPPS 223
Query: 126 SPSAINSPPS----PPPHASPAAAPAITPSSISTPPTKAP-SSISTPPTKAPSTPTN 177
S S SPPS P +S +AP + S + PP+ P S PP P+T T+
Sbjct: 224 STSTSPSPPSKTSSPSTSSSTTSAPKSSTRSTTAPPSSPPCSRPPAPPPAPPATSTS 394
Score = 56.2 bits (134), Expect = 1e-08
Identities = 53/185 (28%), Positives = 84/185 (44%), Gaps = 21/185 (11%)
Frame = +2
Query: 17 SSTFARSLSSSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSP 76
S ++ ++ P ++ S + SP + + + S+ +AP S + P S PP S P
Sbjct: 182 SPSWPSTMPPCPPSSTSTSPSPPSKTSSPSTSSSTTSAPKSSTRSTTAPPSSPPCS--RP 355
Query: 77 ASPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSP---VPVTSSPSPS------P 127
+P AP T S ++ P+++ SP +T+ ++P S+P P TS S S P
Sbjct: 356 PAPPPAPPATSTSPTSRPARSASPP-KTTTAPSTPSTSNPSRKCPTTSPSSKSAPPLAPP 532
Query: 128 SAINSPPSPPPHA------SPAAAPAIT------PSSISTPPTKAPSSISTPPTKAPSTP 175
+ + PP PPP + AA P+ T P +S A S PPT APS
Sbjct: 533 TPRHPPPPPPPST*SPLCPNKAARPSRTF*EAPRPFHLSRKTLTAA*RFSAPPT-APSAV 709
Query: 176 TNGAT 180
+ +T
Sbjct: 710 SRQST 724
Score = 47.8 bits (112), Expect = 3e-06
Identities = 36/101 (35%), Positives = 48/101 (46%), Gaps = 9/101 (8%)
Frame = +2
Query: 89 SISTPPSQAPSPAIAPSTSANSPVA-----SSPVPVTSSPSPSPSAINSPPSPPPHASPA 143
S+S+ S P P PS SP +SP T+SPSP+ ++ P P + P+
Sbjct: 23 SLSSSSSS*PPPPSPPSPPTTSPACWPHTPASPPSTTTSPSPTWPKRSTAARPSP-SWPS 199
Query: 144 AAPAITPSSIST---PPTKAPS-SISTPPTKAPSTPTNGAT 180
P PSS ST PP+K S S S+ T AP + T T
Sbjct: 200 TMPPCPPSSTSTSPSPPSKTSSPSTSSSTTSAPKSSTRSTT 322
Score = 35.8 bits (81), Expect = 0.014
Identities = 27/74 (36%), Positives = 37/74 (49%), Gaps = 6/74 (8%)
Frame = +2
Query: 113 ASSPVPVTSSPSPSPSAINSPPSPPPHA---SPAAAPAITPSSISTPP---TKAPSSIST 166
ASS + +SS P P + SPP+ P +PA+ P+ T S T P T A S S
Sbjct: 14 ASSSLSSSSSS*PPPPSPPSPPTTSPACWPHTPASPPSTTTSPSPTWPKRSTAARPSPSW 193
Query: 167 PPTKAPSTPTNGAT 180
P T P P++ +T
Sbjct: 194 PSTMPPCPPSSTST 235
Score = 35.4 bits (80), Expect = 0.018
Identities = 22/64 (34%), Positives = 34/64 (52%), Gaps = 3/64 (4%)
Frame = +2
Query: 121 SSPSPSPSAINSPPSPPPHASPAAAPAITPSSISTPPTKAPSSISTPP---TKAPSTPTN 177
+S S S S+ + PP P P + P +PA P + ++PP+ S T P T A +P+
Sbjct: 14 ASSSLSSSSSS*PPPPSPPSPPTTSPACWPHTPASPPSTTTSPSPTWPKRSTAARPSPSW 193
Query: 178 GATM 181
+TM
Sbjct: 194 PSTM 205
>TC204251 weakly similar to UP|O36027 (O36027) Wiskott-Aldrich Syndrome
protein homolog, partial (8%)
Length = 1045
Score = 80.1 bits (196), Expect = 6e-16
Identities = 51/138 (36%), Positives = 76/138 (54%)
Frame = +3
Query: 38 PLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVAAPAVTPLSISTPPSQA 97
PL I P+++ P ++ +P+P A +P S PP++ SP++P+ AP+ +P +PPS
Sbjct: 153 PLPILPSLALP---RSSTAPTPTAPTFPPSAPPST--SPSTPLTAPSPSP----SPPSLP 305
Query: 98 PSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSPPSPPPHASPAAAPAITPSSISTPP 157
P+PA +P S +S +A + P SSP SPP PPP SP+AA P+ ST
Sbjct: 306 PAPAGSPGASTSSAMACA-APKPSSP--------SPPPPPPPPSPSAATTSPPTKPSTKS 458
Query: 158 TKAPSSISTPPTKAPSTP 175
PS+ T K P+ P
Sbjct: 459 KPLPSTRGTSRPKNPTAP 512
Score = 53.9 bits (128), Expect = 5e-08
Identities = 47/141 (33%), Positives = 62/141 (43%), Gaps = 5/141 (3%)
Frame = +3
Query: 23 SLSSSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSP----AS 78
S + +P A P +P + SP+ P +PSPS P S PPA GSP +S
Sbjct: 192 STAPTPTAPTFPPSAPPSTSPSTPLTAP---SPSPS------PPSLPPAPAGSPGASTSS 344
Query: 79 PVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAI-NSPPSPP 137
+A A P S S PP P P +PS + SP P ++ P PS S P P
Sbjct: 345 AMACAAPKPSSPSPPP---PPPPPSPSAATTSP----PTKPSTKSKPLPSTRGTSRPKNP 503
Query: 138 PHASPAAAPAITPSSISTPPT 158
P+ AP+ +P T T
Sbjct: 504 TAP*PSTAPSRSPIRRGTSAT 566
Score = 48.5 bits (114), Expect = 2e-06
Identities = 34/97 (35%), Positives = 48/97 (49%), Gaps = 6/97 (6%)
Frame = +3
Query: 90 ISTPPSQA-PSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSPPSPPPH-----ASPA 143
+S PP PS A+ S++A +P A + P ++ PS SPS + PSP P +PA
Sbjct: 141 LSFPPLPILPSLALPRSSTAPTPTAPT-FPPSAPPSTSPSTPLTAPSPSPSPPSLPPAPA 317
Query: 144 AAPAITPSSISTPPTKAPSSISTPPTKAPSTPTNGAT 180
+P + SS PSS S PP P +P+ T
Sbjct: 318 GSPGASTSSAMACAAPKPSSPSPPPPPPPPSPSAATT 428
>TC210299 homologue to UP|EFT2_SOYBN (P46280) Elongation factor Tu,
chloroplast precursor (EF-Tu), complete
Length = 1615
Score = 80.1 bits (196), Expect = 6e-16
Identities = 55/160 (34%), Positives = 82/160 (50%), Gaps = 18/160 (11%)
Frame = +2
Query: 39 LAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVAAPAVTPLSISTPPSQAP 98
L +S ++ P P++++ P T+PPP + P+S + +PL + PP+ AP
Sbjct: 47 LLMSTSLLHPPPIISSLKP--------TNPPPHTSPLPSSTPPQSSTSPLP-TPPPAAAP 199
Query: 99 SPAIAP--STSANSPVASSPVPVTSS----PSPSPSAINSPPS------------PPPHA 140
SP+ P S+SA SP ++S TS+ PSP PS SPPS PPP +
Sbjct: 200 SPSAPPAASSSARSPTSTSAPSATSTTVRPPSPPPSPWPSPPSAIVPLKNTTRSTPPPRS 379
Query: 141 SPAAAPAITPSSISTPPTKAPSSISTPPTKAPSTPTNGAT 180
+PAA+P+ P P P + +TP + AP+TPT T
Sbjct: 380 APAASPSTPP-----PSNTRPRTATTPTSTAPATPTTSKT 484
Score = 67.0 bits (162), Expect = 5e-12
Identities = 58/161 (36%), Positives = 80/161 (49%), Gaps = 4/161 (2%)
Frame = +2
Query: 24 LSSSPVATPSPAGSPLAIS-PAVSSPVPLMNAPSPSPFAVNYPTSPPPASFG--SPASPV 80
+SS P P SPL S P SS PL P+P P A P++PP AS SP S
Sbjct: 86 ISSLKPTNPPPHTSPLPSSTPPQSSTSPL---PTPPPAAAPSPSAPPAASSSARSPTS-T 253
Query: 81 AAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSPPSPPP-H 139
+AP+ T ++ PPS PSP +P SA P+ ++ S+P P + SP +PPP +
Sbjct: 254 SAPSATSTTVR-PPSPPPSPWPSPP-SAIVPLKNT---TRSTPPPRSAPAASPSTPPPSN 418
Query: 140 ASPAAAPAITPSSISTPPTKAPSSISTPPTKAPSTPTNGAT 180
P A T ++ +TP T S + P APS+ + T
Sbjct: 419 TRPRTATTPTSTAPATPTTSKT*SPAPPRWTAPSSSSPAPT 541
Score = 60.5 bits (145), Expect = 5e-10
Identities = 56/174 (32%), Positives = 87/174 (49%), Gaps = 17/174 (9%)
Frame = +2
Query: 13 SLLVSSTFARSLSSSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVN---YPTSPP 69
S L SST +S S+SP+ TP PA +P +P +S + +P A + P SPP
Sbjct: 125 SPLPSSTPPQS-STSPLPTPPPAAAPSPSAPPAASSSARSPTSTSAPSATSTTVRPPSPP 301
Query: 70 PASFGSPASPVAAPAVTPL---SISTPPSQAPSPAIAPST---------SANSPVASSPV 117
P+ + SP S A+ PL + STPP ++ +PA +PST +A +P +++P
Sbjct: 302 PSPWPSPPS-----AIVPLKNTTRSTPPPRS-APAASPSTPPPSNTRPRTATTPTSTAPA 463
Query: 118 PVTSSP--SPSPSAINSPPSPPPHASPAAAPAITPSSISTPPTKAPSSISTPPT 169
T+S SP+P +P S P +P A ++ S P A ++S+ T
Sbjct: 464 TPTTSKT*SPAPPRWTAPSSSSP--APTAQCPKPKNTFSWPNKSAFPTLSSSST 619
>TC215396 weakly similar to UP|Q8H9F4 (Q8H9F4) 41 kD chloroplast nucleoid DNA
binding protein (CND41), partial (69%)
Length = 1808
Score = 79.7 bits (195), Expect = 8e-16
Identities = 60/163 (36%), Positives = 80/163 (48%), Gaps = 2/163 (1%)
Frame = +1
Query: 12 ASLLVSSTFARSLSSSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPA 71
+S SS A++ +S VA P+ + S SP+ + P P + SP+ SPPP
Sbjct: 772 SSTTSSSVAAKTTRASLVAPPASSASAATPSPSSNKPPPYIVKSSPT-------ASPPP- 927
Query: 72 SFGSPASPVAAP-AVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSP-SPSA 129
PA P A+P A PL S+ P PSPA PST++ SP + P + SP P SP A
Sbjct: 928 ----PAPPAASPSAPPPLPTSSTPPSPPSPAAPPSTASTSPASPWAAPNSQSPPPPSPPA 1095
Query: 130 INSPPSPPPHASPAAAPAITPSSISTPPTKAPSSISTPPTKAP 172
+ S S P SPA+ P TPP+ PS + P T P
Sbjct: 1096VQS--STPAP*SPASPPP------PTPPSAPPSGKACPNTPPP 1200
Score = 63.5 bits (153), Expect = 6e-11
Identities = 54/160 (33%), Positives = 74/160 (45%), Gaps = 18/160 (11%)
Frame = +1
Query: 37 SPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVAAPAVTPLSISTPPSQ 96
SPLA S A SP P P+ S + + AS +P + +A A TP +P S
Sbjct: 721 SPLATSAASGSP*P---PPTSSTTSSSVAAKTTRASLVAPPAS-SASAATP----SPSSN 876
Query: 97 APSPAIAPST-SANSPVASSPVPVTSSPSPSPSAINSPPSPPPHASP-------AAAPAI 148
P P I S+ +A+ P + P S+P P P++ ++PPSPP A+P A+P
Sbjct: 877 KPPPYIVKSSPTASPPPPAPPAASPSAPPPLPTS-STPPSPPSPAAPPSTASTSPASPWA 1053
Query: 149 TPSSISTPPTKAP----------SSISTPPTKAPSTPTNG 178
P+S S PP P S S PP PS P +G
Sbjct: 1054APNSQSPPPPSPPAVQSSTPAP*SPASPPPPTPPSAPPSG 1173
Score = 46.2 bits (108), Expect = 1e-05
Identities = 33/105 (31%), Positives = 52/105 (49%), Gaps = 2/105 (1%)
Frame = +1
Query: 78 SPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSPPSPP 137
SP+A A + P S S ++A T+ S VA P S+ +PSPS+ N PP
Sbjct: 721 SPLATSAASGSP*PPPTSSTTSSSVAAKTTRASLVAP-PASSASAATPSPSS-NKPPPYI 894
Query: 138 PHASPAAA--PAITPSSISTPPTKAPSSISTPPTKAPSTPTNGAT 180
+SP A+ P P++ + P P+S + P +P+ P + A+
Sbjct: 895 VKSSPTASPPPPAPPAASPSAPPPLPTSSTPPSPPSPAAPPSTAS 1029
Score = 42.0 bits (97), Expect = 2e-04
Identities = 37/132 (28%), Positives = 56/132 (42%), Gaps = 42/132 (31%)
Frame = +1
Query: 88 LSISTPPSQAPSPAIAPSTSANSPVASS-------PVPVTSSPS---------------- 124
+S P Q + + + + SP+A+S P P +S+ S
Sbjct: 655 MSRDVQPPQKHAYTASSTVTVRSPLATSAASGSP*PPPTSSTTSSSVAAKTTRASLVAPP 834
Query: 125 --------PSPSAINSPP---------SPPPHASPAAAPAITP--SSISTPPTKAPSSIS 165
PSPS+ PP SPPP A PAA+P+ P + STPP +P S +
Sbjct: 835 ASSASAATPSPSSNKPPPYIVKSSPTASPPPPAPPAASPSAPPPLPTSSTPP--SPPSPA 1008
Query: 166 TPPTKAPSTPTN 177
PP+ A ++P +
Sbjct: 1009APPSTASTSPAS 1044
>TC226993 homologue to UP|GMD1_ARATH (Q9SNY3) GDP-mannose 4,6 dehydratase
(GDP-D-mannose dehydratase) (GMD) , partial (94%)
Length = 1388
Score = 79.3 bits (194), Expect = 1e-15
Identities = 56/148 (37%), Positives = 71/148 (47%), Gaps = 3/148 (2%)
Frame = +2
Query: 27 SPVATPSPAGSPLAISPAVSSPVPL-MNAPSPSPFAVNYPTSPPPASFGS--PASPVAAP 83
SP PS AGS + SP S+ PL + +PSPS + PTS PPA S P+ P + P
Sbjct: 329 SPTPPPSAAGSTPS-SPTRSTTSPLSLTSPSPSRYPTTPPTSSPPAPSASSRPSVPTSPP 505
Query: 84 AVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSPPSPPPHASPA 143
P S +T P AP AP P+ SP P PS +PP+PPP+A P
Sbjct: 506 PAAPTSATTKP--APPRCSAP-------------PLRRSPKPPPST-PAPPTPPPNAPPT 637
Query: 144 AAPAITPSSISTPPTKAPSSISTPPTKA 171
P T + P A SS ++PP A
Sbjct: 638 GTP*TTARLTPSSPATASSSTTSPPAAA 721
Score = 64.3 bits (155), Expect = 4e-11
Identities = 60/181 (33%), Positives = 86/181 (47%), Gaps = 7/181 (3%)
Frame = +2
Query: 19 TFARSLSSSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPAS 78
T RS S++ + P+ PA SS P+ +P+P P A S P S SP S
Sbjct: 245 TSTRSASTTSMWIPTTP-----TRPA*SSTTPI--SPTPPPSAAGSTPSSPTRSTTSPLS 403
Query: 79 PVA-APAVTPLSISTPPSQAPSPAIAPSTSANSPVASSP---VPVTSSPSPSPSAINSPP 134
+ +P+ P +TPP+ +P APS S+ V +SP P +++ P+P ++PP
Sbjct: 404 LTSPSPSRYP---TTPPTSSPP---APSASSRPSVPTSPPPAAPTSATTKPAPPRCSAPP 565
Query: 135 ---SPPPHASPAAAPAITPSSISTPPTKAPSSISTPPTKAPSTPTNGATMNGFNVAGYAA 191
SP P S A P TPP AP + TP T A TP++ AT + + AA
Sbjct: 566 LRRSPKPPPSTPAPP--------TPPPNAPPT-GTP*TTARLTPSSPATASSSTTSPPAA 718
Query: 192 A 192
A
Sbjct: 719 A 721
Score = 44.7 bits (104), Expect = 3e-05
Identities = 35/103 (33%), Positives = 48/103 (45%), Gaps = 23/103 (22%)
Frame = +2
Query: 101 AIAPSTSANS--------------PVASSPVPVTSSPSPSPSAINSPPSPPPHA--SPAA 144
++AP TS S P SS P+ SP+P PSA S PS P + SP +
Sbjct: 230 SVAPRTSTRSASTTSMWIPTTPTRPA*SSTTPI--SPTPPPSAAGSTPSSPTRSTTSPLS 403
Query: 145 APAITPSSI-STPPTKAP------SSISTPPTKAPSTPTNGAT 180
+ +PS +TPPT +P S S P + P+ PT+ T
Sbjct: 404 LTSPSPSRYPTTPPTSSPPAPSASSRPSVPTSPPPAAPTSATT 532
Score = 44.3 bits (103), Expect = 4e-05
Identities = 29/95 (30%), Positives = 45/95 (46%)
Frame = +2
Query: 82 APAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSAINSPPSPPPHAS 141
AP + S ST P+ P+ S+ +P+ SP+P PSA S PS P ++
Sbjct: 236 APRTSTRSASTTSMWIPTTPTRPA*SSTTPI---------SPTPPPSAAGSTPSSPTRST 388
Query: 142 PAAAPAITPSSISTPPTKAPSSISTPPTKAPSTPT 176
+ +PS P++ P+ TPPT +P P+
Sbjct: 389 TSPLSLTSPS-----PSRYPT---TPPTSSPPAPS 469
>TC209975 weakly similar to UP|Q7T9Y7 (Q7T9Y7) Odv-e66, partial (9%)
Length = 537
Score = 79.3 bits (194), Expect = 1e-15
Identities = 55/166 (33%), Positives = 83/166 (49%), Gaps = 12/166 (7%)
Frame = +1
Query: 27 SPVATPSPAGSPLAISPAVSSPVPLMNAPS-PSPFAVNYPTSPPPASFGSPASPVAAPAV 85
+P PSP P + +P P + P+ P+P P+S PP + P +P +P
Sbjct: 28 APPIAPSPGNHPPYVPTPPKTPSPSYSPPNVPTPPKTPSPSSQPPFTPTPPKTP--SPTS 201
Query: 86 TPLSISTPPS---QAPSPAIAPSTSANSPVASSPVPVTSSPS--------PSPSAINSPP 134
P I TPPS Q PS A P+T SP + P P T PS PSPS ++P
Sbjct: 202 QPPYIPTPPSPISQPPSIATPPNTL--SPTSQPPYPSTIPPSTSLSPSYPPSPSPSSAPT 375
Query: 135 SPPPHASPAAAPAITPSSISTPPTKAPSSISTPPTKAPSTPTNGAT 180
PP + +PA +P +PS+ +T PT +P++ + P+ + S+ + T
Sbjct: 376 YPPSYLAPATSPP-SPSTSATTPTISPAATTPSPSSSSSSGASNET 510
Score = 78.2 bits (191), Expect = 2e-15
Identities = 56/158 (35%), Positives = 85/158 (53%), Gaps = 4/158 (2%)
Frame = +1
Query: 27 SPVATPSPAGSPLAISPAVSSPVPLMNAP-SPSPFAVNYPTSPPPASFGSPASPVAAPA- 84
+P TPSP+ SP + +P P P +P+P PTS PP +P SP++ P
Sbjct: 76 TPPKTPSPSYSPPNVPTPPKTPSPSSQPPFTPTPPKTPSPTSQPPY-IPTPPSPISQPPS 252
Query: 85 -VTPLSISTPPSQAPSPA-IAPSTSANSPVASSPVPVTSSPSPSPSAINSPPSPPPHASP 142
TP + +P SQ P P+ I PSTS + SP P +S+P+ PS + SPP ++
Sbjct: 253 IATPPNTLSPTSQPPYPSTIPPSTSLSPSYPPSPSP-SSAPTYPPSYLAPATSPPSPSTS 429
Query: 143 AAAPAITPSSISTPPTKAPSSISTPPTKAPSTPTNGAT 180
A P I+P++ + P+ + SS ++ T P+ P NGA+
Sbjct: 430 ATTPTISPAATTPSPSSSSSSGASNET-TPARP-NGAS 537
Score = 65.9 bits (159), Expect = 1e-11
Identities = 52/161 (32%), Positives = 71/161 (43%), Gaps = 8/161 (4%)
Frame = +1
Query: 28 PVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVAAPAVTP 87
P TPSP +P P+ +P P V P P S+ P P P TP
Sbjct: 1 PPKTPSPIYAP-----------PIAPSPGNHPPYVPTPPKTPSPSYSPPNVPT--PPKTP 141
Query: 88 LSISTPPSQAPSPAIAPSTSANSP-VASSPVPVTSSPS-PSPSAINSPPSPPPHASPAAA 145
S PP P+P PS ++ P + + P P++ PS +P SP S PP+ P+
Sbjct: 142 SPSSQPPF-TPTPPKTPSPTSQPPYIPTPPSPISQPPSIATPPNTLSPTSQPPY--PSTI 312
Query: 146 PAITPSSISTPPTKAPSSIST------PPTKAPSTPTNGAT 180
P T S S PP+ +PSS T P +P +P+ AT
Sbjct: 313 PPSTSLSPSYPPSPSPSSAPTYPPSYLAPATSPPSPSTSAT 435
>TC225559 similar to UP|Q9XIV1 (Q9XIV1) Cucumis sativus mRNA expressed in
cucumber hypocotyls, complete cds (Arabinogalactan
protein), partial (47%)
Length = 870
Score = 79.0 bits (193), Expect = 1e-15
Identities = 56/167 (33%), Positives = 80/167 (47%), Gaps = 20/167 (11%)
Frame = +3
Query: 31 TPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPAS-PVAAPAVTPLS 89
+P+ A S +PA ++P + SP V P S PPA+ +PAS P A+P TP +
Sbjct: 138 SPAAAPSNTPATPAAATPAQAPSTVPKSPAPVASPKSSPPAA--TPASTPAASPTTTPAA 311
Query: 90 ---ISTPPSQAPSPAIAPSTS---ANSPVASSPVPVTSSPSPSPSAINSP-----PSPPP 138
++ PP+ +P PA P + A PV+S P PV S P+P + +P PSP P
Sbjct: 312 PAPVTKPPAASPPPATPPPATPPPAPVPVSSPPAPVPVSSPPAPVPVAAPTTPVAPSPAP 491
Query: 139 --------HASPAAAPAITPSSISTPPTKAPSSISTPPTKAPSTPTN 177
H +PA +P + P + PPT AP + P T N
Sbjct: 492 KHKKKGKKHGAPAPSPLLGPPA---PPTGAPGPSEDASSPGPGTAAN 623
Score = 60.8 bits (146), Expect = 4e-10
Identities = 51/144 (35%), Positives = 68/144 (46%), Gaps = 19/144 (13%)
Frame = +3
Query: 25 SSSPVATPS--PAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPASPVAA 82
SS P ATP+ PA SP +PA +PV A SP P A P +PPPA PV++
Sbjct: 246 SSPPAATPASTPAASPTT-TPAAPAPVTKPPAASPPP-ATPPPATPPPAPV-----PVSS 404
Query: 83 PAVTPLSISTPPSQ----APSPAIAPSTS-------------ANSPVASSPVPVTSSPSP 125
P P+ +S+PP+ AP+ +APS + A SP+ P P T +P P
Sbjct: 405 PPA-PVPVSSPPAPVPVAAPTTPVAPSPAPKHKKKGKKHGAPAPSPLLGPPAPPTGAPGP 581
Query: 126 SPSAINSPPSPPPHASPAAAPAIT 149
S A S P P A+ + T
Sbjct: 582 SEDA--SSPGPGTAANDESGAETT 647
Score = 60.5 bits (145), Expect = 5e-10
Identities = 47/139 (33%), Positives = 65/139 (45%), Gaps = 5/139 (3%)
Frame = +3
Query: 17 SSTFARSLSSSPVATPSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSP 76
+ST A S +++P A P+P P A SP ++P P P+P P S PPA
Sbjct: 270 ASTPAASPTTTPAA-PAPVTKPPAASPPPATPPPATPPPAP------VPVSSPPA----- 413
Query: 77 ASPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSP-----SPSPSAIN 131
P+ +S+PP AP P AP+T PVA SP P +P+PS +
Sbjct: 414 ----------PVPVSSPP--APVPVAAPTT----PVAPSPAPKHKKKGKKHGAPAPSPLL 545
Query: 132 SPPSPPPHASPAAAPAITP 150
PP+PP A + A +P
Sbjct: 546 GPPAPPTGAPGPSEDASSP 602
>TC226468 weakly similar to UP|O65450 (O65450) Glycine-rich protein, partial
(20%)
Length = 1041
Score = 77.4 bits (189), Expect = 4e-15
Identities = 54/164 (32%), Positives = 79/164 (47%), Gaps = 15/164 (9%)
Frame = -2
Query: 25 SSSPVATPSP----AGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSP---- 76
SS+P+A P A P +P +S P AP P+P + P P +S +P
Sbjct: 908 SSTPLAVTQPPTVVASPPTTSTPPTTSQPPANVAPKPAPVKPSAPKVAPASSPKAPPPQI 729
Query: 77 -------ASPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSPSPSA 129
SPV+ P + P +++PP P P IAP T A +P + +P T +P+P+P+
Sbjct: 728 PIPQPPKPSPVSPPPLPPPPVASPP-PLPPPKIAP-TPAKTPPSPAPAKATPAPAPAPTK 555
Query: 130 INSPPSPPPHASPAAAPAITPSSISTPPTKAPSSISTPPTKAPS 173
PSP P PAA PA P+ + P AP+ + P AP+
Sbjct: 554 PAPTPSPVP-PPPAATPA--PAPVIEVPAPAPAPVIEVPAPAPA 432
Score = 75.1 bits (183), Expect = 2e-14
Identities = 56/161 (34%), Positives = 77/161 (47%), Gaps = 6/161 (3%)
Frame = -2
Query: 21 ARSLSSSPVATPS---PAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPASFGSPA 77
A++ ++ PV P P +P A SP SS PL P+ A TS PP + PA
Sbjct: 989 AQAPATVPVKLPPTTLPPTTPTATSP--SSSTPLAVTQPPTVVASPPTTSTPPTTSQPPA 816
Query: 78 SPVAAPAVTPLSISTPPSQAPSPAIAPSTSANSPVASSPVPVTSSPSP-SPSAINSPP-- 134
+ PA P + +P +AP++S +P P+P PSP SP + PP
Sbjct: 815 NVAPKPA---------PVKPSAPKVAPASSPKAPPPQIPIPQPPKPSPVSPPPLPPPPVA 663
Query: 135 SPPPHASPAAAPAITPSSISTPPTKAPSSISTPPTKAPSTP 175
SPPP P AP TP+ TPP+ AP+ + P AP+ P
Sbjct: 662 SPPPLPPPKIAP--TPA--KTPPSPAPAKATPAPAPAPTKP 552
Score = 63.5 bits (153), Expect = 6e-11
Identities = 49/142 (34%), Positives = 64/142 (44%), Gaps = 22/142 (15%)
Frame = -2
Query: 32 PSPAGSPLAISPAVSSPVPLMNAPSPSPFAVNYPTSPPPA-SFGSPASPVAAPAVTPLSI 90
PSP P P V+SP PL P +P P SP PA + +PA PA TP +
Sbjct: 707 PSPVSPPPLPPPPVASPPPLP-PPKIAPTPAKTPPSPAPAKATPAPAPAPTKPAPTPSPV 531
Query: 91 STPPSQAPSPA--IAPSTSANSPVASSPVPVTS-----------------SPSPSPSAI- 130
PP+ P+PA I A +PV P P + +P+P+P+ I
Sbjct: 530 PPPPAATPAPAPVIEVPAPAPAPVIEVPAPAPAHHRHRRHRHRHRHRRHQAPAPAPTIIR 351
Query: 131 NSPPSPPPH-ASPAAAPAITPS 151
SPP+PP A APA +PS
Sbjct: 350 KSPPAPPDSTAESDTAPAPSPS 285
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.306 0.120 0.341
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,622,377
Number of Sequences: 63676
Number of extensions: 330818
Number of successful extensions: 38999
Number of sequences better than 10.0: 3718
Number of HSP's better than 10.0 without gapping: 8654
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 19312
length of query: 202
length of database: 12,639,632
effective HSP length: 93
effective length of query: 109
effective length of database: 6,717,764
effective search space: 732236276
effective search space used: 732236276
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.6 bits)
S2: 56 (26.2 bits)
Medicago: description of AC144517.1


