

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
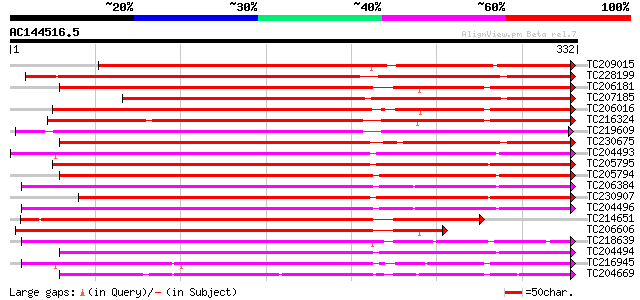
Query= AC144516.5 - phase: 0
(332 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC209015 similar to UP|PE72_ARATH (Q9FJZ9) Peroxidase 72 precurs... 302 2e-82
TC228199 weakly similar to UP|Q9XGV6 (Q9XGV6) Bacterial-induced ... 287 4e-78
TC206181 similar to UP|Q7XYR7 (Q7XYR7) Class III peroxidase, par... 285 3e-77
TC207185 similar to UP|Q9MAX9 (Q9MAX9) Peroxidase precursor , p... 277 5e-75
TC206016 similar to UP|Q6T1C8 (Q6T1C8) Peroxidase precursor, par... 271 3e-73
TC216324 similar to UP|Q9XFL4 (Q9XFL4) Peroxidase 3, complete 269 1e-72
TC219609 similar to UP|PER1_ARAHY (P22195) Cationic peroxidase 1... 265 2e-71
TC230675 similar to PDB|1SCH_A.0|1633130|1SCH_A Chain A, Peanut ... 261 4e-70
TC204493 UP|O22443 (O22443) Seed coat peroxidase precursor , co... 260 7e-70
TC205795 similar to UP|Q9XFL3 (Q9XFL3) Peroxidase 1 (Fragment), ... 257 4e-69
TC205794 similar to UP|Q9XFL3 (Q9XFL3) Peroxidase 1 (Fragment), ... 252 1e-67
TC206384 UP|O23961 (O23961) Peroxidase precursor, complete 250 7e-67
TC230907 similar to UP|O24081 (O24081) Peroxidase1A precursor ,... 249 2e-66
TC204496 homologue to UP|O23961 (O23961) Peroxidase precursor, p... 245 2e-65
TC214651 similar to UP|Q9XGV6 (Q9XGV6) Bacterial-induced peroxid... 239 1e-63
TC206606 similar to UP|Q7XYR7 (Q7XYR7) Class III peroxidase, par... 235 2e-62
TC218639 similar to UP|PE17_ARATH (Q9SJZ2) Peroxidase 17 precurs... 232 2e-61
TC204494 similar to UP|O23961 (O23961) Peroxidase precursor, par... 230 8e-61
TC216945 similar to UP|Q43854 (Q43854) Peroxidase precursor , p... 229 1e-60
TC204669 similar to UP|PER2_ARAHY (P22196) Cationic peroxidase 2... 227 5e-60
>TC209015 similar to UP|PE72_ARATH (Q9FJZ9) Peroxidase 72 precursor (Atperox
P72) (PRXR8) (ATP6a) , partial (81%)
Length = 1099
Score = 302 bits (773), Expect = 2e-82
Identities = 158/281 (56%), Positives = 195/281 (69%), Gaps = 2/281 (0%)
Frame = +3
Query: 53 DPRLAASLLRLHFHDCFVMGCDASVLLDSVEGMTSEKQAGPNVNSLRGFEVIDKIKYLLE 112
+PRLAAS+LRLHFHDCFV GCDAS+LLDS E + SEK + PN NS RGFEVID IK LE
Sbjct: 3 EPRLAASILRLHFHDCFVKGCDASLLLDSSESINSEKGSNPNRNSARGFEVIDAIKAELE 182
Query: 113 KECPLTVSCADILAMVARDAVELRGGPRWEVWLGRKDSLESSFSGANLFIPAPNSSLETL 172
++CP TVSCADIL + ARD+V L GGP WEV LGR+DSL +S SG+N IPAPN++ +T+
Sbjct: 183 RQCPSTVSCADILTLAARDSVVLTGGPNWEVPLGRRDSLGASISGSNNNIPAPNNTFQTI 362
Query: 173 INNFKQQGLDIEDLVVLSGSHTIGRARCLSFRQRIYET--KQEYHHAYDRYKRYTTFRRI 230
+ FK QGLD+ DLV LSG HTIG ARC +FRQR+Y E D+Y +
Sbjct: 363 LTKFKLQGLDLVDLVALSGGHTIGNARCTTFRQRLYNQSGNGEPDSTLDQY-----YAST 527
Query: 231 LQSICPVTGRDDKFAPLDFQTPKRFDNQYFINIIEGKGLLGSDNVLISQDLDGRIRKQVW 290
L++ CP +G D LD+ TP +FDN YF N++ KGLL SD VL + + + + V
Sbjct: 528 LRTRCPSSGGDQNLFFLDYATPYKFDNSYFKNLLAYKGLLSSDQVLFTMNQES--AELVK 701
Query: 291 GYASNEKLFFDSFAKSMIKMGNINVLTGSEGEIRRNCRFVN 331
YA +FF+ FAKSMIKMGNI+ LT S GEIR NCR +N
Sbjct: 702 LYAERNDIFFEHFAKSMIKMGNISPLTNSRGEIRENCRRIN 824
>TC228199 weakly similar to UP|Q9XGV6 (Q9XGV6) Bacterial-induced peroxidase
precursor , partial (81%)
Length = 1075
Score = 287 bits (735), Expect = 4e-78
Identities = 163/324 (50%), Positives = 197/324 (60%), Gaps = 2/324 (0%)
Frame = +3
Query: 10 LITILSNIHTLRGSELLVHEYYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRLHFHDCF 69
L+ +L T G+EL ++Y CP IV+ VA A+ K+PR+ ASLLRLHFHDCF
Sbjct: 9 LLLVLVGATTASGAELCA-DFYSCTCPNLLPIVKKGVAKAIQKEPRMGASLLRLHFHDCF 185
Query: 70 VMGCDASVLLDSVEGMTSEKQAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCADILAMVA 129
V GCDAS+LLD E+ A N S RGF VI+ IK +EKECP VSCADILA+ A
Sbjct: 186 VNGCDASILLDDTSNFIGEQTAAANNQSARGFNVINDIKASVEKECPRVVSCADILALSA 365
Query: 130 RDAVELRGGPRWEVWLGRKDSLESSFSGANLFIPAPNSSLETLINNFKQQGLDIEDLVVL 189
RD+V GGP WEV LGR+DS +S S AN IP P SL LINNF QGL + DLV L
Sbjct: 366 RDSVVYLGGPSWEVGLGRRDSTTASRSDANNSIPGPFLSLTALINNFANQGLSVTDLVAL 545
Query: 190 SGSHTIGRARCLSFRQRIYETKQEYHHAYDRYKRYTTFRRILQSICPV-TGRDDKFAPLD 248
SG+HTIG A C +FR H Y+ ++R+ LQS CP +G D PLD
Sbjct: 546 SGAHTIGLAECKNFRA----------HIYNDSNVDPSYRKFLQSKCPPRSGNDKTLEPLD 695
Query: 249 FQTPKRFDNQYFINIIEGKGLLGSDNVLIS-QDLDGRIRKQVWGYASNEKLFFDSFAKSM 307
QTP FDN YF N++ K LL SD L + D +RK YA+N FF+ FAK M
Sbjct: 696 HQTPIHFDNLYFQNLVSKKALLHSDQELFNGSSTDNLVRK----YATNAAAFFEDFAKGM 863
Query: 308 IKMGNINVLTGSEGEIRRNCRFVN 331
+KM NI LTGS+G+IR NC VN
Sbjct: 864 LKMSNIKPLTGSQGQIRINCGKVN 935
>TC206181 similar to UP|Q7XYR7 (Q7XYR7) Class III peroxidase, partial (94%)
Length = 1219
Score = 285 bits (728), Expect = 3e-77
Identities = 158/305 (51%), Positives = 189/305 (61%), Gaps = 3/305 (0%)
Frame = +2
Query: 30 YYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRLHFHDCFVMGCDASVLLDSVEGMTSEK 89
+Y CP V+ V A+ K+ R+ ASLLRL FHDCFV GCD S+LLD T EK
Sbjct: 146 FYYHSCPNLFSSVKSTVQSAISKETRMGASLLRLFFHDCFVNGCDGSILLDDTSSFTGEK 325
Query: 90 QAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCADILAMVARDAVELRGGPRWEVWLGRKD 149
A PN NS RGFEVID IK +EK CP VSCADILA+ ARD+V++ GGP W V LGR+D
Sbjct: 326 NANPNRNSARGFEVIDNIKSAVEKVCPGVVSCADILAIAARDSVQILGGPSWNVKLGRRD 505
Query: 150 SLESSFSGANLFIPAPNSSLETLINNFKQQGLDIEDLVVLSGSHTIGRARCLSFRQRIY- 208
+ +S S AN IP P S+L LI+ F GL +DLV LSG HTIG+ARC +FR RIY
Sbjct: 506 ARTASQSAANNGIPPPTSNLNQLISRFSALGLSTKDLVALSGGHTIGQARCTNFRARIYN 685
Query: 209 ETKQEYHHAYDRYKRYTTFRRILQSICPVT--GRDDKFAPLDFQTPKRFDNQYFINIIEG 266
ET E T F R Q CP T D+ APLD QTP FDN YF N+++
Sbjct: 686 ETNIE-----------TAFARTRQQSCPRTSGSGDNNLAPLDLQTPTSFDNYYFKNLVQK 832
Query: 267 KGLLGSDNVLISQDLDGRIRKQVWGYASNEKLFFDSFAKSMIKMGNINVLTGSEGEIRRN 326
KGLL SD L + G V GY++N F FA +MIKMG+I+ LTGS GEIR+N
Sbjct: 833 KGLLHSDQQLFN---GGSTDSIVRGYSTNPGTFSSDFAAAMIKMGDISPLTGSNGEIRKN 1003
Query: 327 CRFVN 331
CR +N
Sbjct: 1004CRRIN 1018
>TC207185 similar to UP|Q9MAX9 (Q9MAX9) Peroxidase precursor , partial (77%)
Length = 1072
Score = 277 bits (708), Expect = 5e-75
Identities = 146/266 (54%), Positives = 186/266 (69%), Gaps = 1/266 (0%)
Frame = +3
Query: 67 DCFVMGCDASVLLDSVEGMTSEKQAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCADILA 126
DCFV GCDASVLLDS + SEK++ PN +S RGFEVID+IK LEKECP TVSCADILA
Sbjct: 9 DCFVKGCDASVLLDSSGTIISEKRSNPNRDSARGFEVIDEIKSALEKECPHTVSCADILA 188
Query: 127 MVARDAVELRGGPRWEVWLGRKDSLESSFSGANLFIPAPNSSLETLINNFKQQGLDIEDL 186
+ ARD+ L GGP W V LGR+DSL +S SG+N IPAPN++ +T++ FK +GLDI DL
Sbjct: 189 LAARDSTVLTGGPSWGVPLGRRDSLGASISGSNNNIPAPNNTFQTILTKFKLKGLDIVDL 368
Query: 187 VVLSGSHTIGRARCLSFRQRIYETKQEYHHAYDRYKRYTTFRRILQSICPVTGRDDKFAP 246
V LSGSHTIG +RC SFRQR+Y + + + + L++ CP +G D
Sbjct: 369 VALSGSHTIGNSRCTSFRQRLY---NQTGNGKADFTLDQVYAAELRTRCPRSGGDQNLFV 539
Query: 247 LDFQTPKRFDNQYFINIIEGKGLLGSDNVLISQD-LDGRIRKQVWGYASNEKLFFDSFAK 305
LDF TP +FDN Y+ N++ KGLL SD +L++++ + + KQ YA N LFF+ FAK
Sbjct: 540 LDFVTPIKFDNFYYKNLLANKGLLSSDEILLTKNQVSADLVKQ---YAENNDLFFEQFAK 710
Query: 306 SMIKMGNINVLTGSEGEIRRNCRFVN 331
SM+KMGNI LTGS GEIR+NCR +N
Sbjct: 711 SMVKMGNITPLTGSRGEIRKNCRGIN 788
>TC206016 similar to UP|Q6T1C8 (Q6T1C8) Peroxidase precursor, partial (95%)
Length = 1214
Score = 271 bits (693), Expect = 3e-73
Identities = 152/309 (49%), Positives = 190/309 (61%), Gaps = 3/309 (0%)
Frame = +3
Query: 26 LVHEYYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRLHFHDCFVMGCDASVLLDSVEGM 85
L +Y + CP + V+ V AV+++PR+ AS++RL FHDCFV GCD S+LLD
Sbjct: 117 LSKNFYSKTCPNVFNTVKSVVKSAVVREPRIGASIVRLFFHDCFVQGCDGSILLDDTPTF 296
Query: 86 TSEKQAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCADILAMVARDAVELRGGPRWEVWL 145
EK A N NS+RGFEVID IK +EK CP VSCADIL + +RD+V L GGP W+V L
Sbjct: 297 QGEKTAAANNNSVRGFEVIDAIKSEVEKICPGVVSCADILDIASRDSVVLLGGPFWKVRL 476
Query: 146 GRKDSLESSFSGANL-FIPAPNSSLETLINNFKQQGLDIEDLVVLSGSHTIGRARCLSFR 204
GR+DS ++F+ AN IP P S+L LI F+ QGL D+V LSG+HT G+ARC SFR
Sbjct: 477 GRRDSRTANFTAANTGVIPPPTSNLTNLITRFQDQGLSARDMVALSGAHTFGKARCTSFR 656
Query: 205 QRIYETKQEYHHAYDRYKRYTTFRRILQSICPVTG--RDDKFAPLDFQTPKRFDNQYFIN 262
RIY DR TF Q CP T D+ A LDF+TP FDN YF N
Sbjct: 657 DRIYNQTN-----IDR-----TFALARQRRCPRTNGTGDNNLANLDFRTPNHFDNNYFKN 806
Query: 263 IIEGKGLLGSDNVLISQDLDGRIRKQVWGYASNEKLFFDSFAKSMIKMGNINVLTGSEGE 322
++ +GLL SD VL + G V Y+ N K F F K+MI+MG+I LTGS+GE
Sbjct: 807 LLIKRGLLNSDQVLFN---GGSTDSLVRTYSQNNKAFDTDFVKAMIRMGDIKPLTGSQGE 977
Query: 323 IRRNCRFVN 331
IR+NCR VN
Sbjct: 978 IRKNCRRVN 1004
>TC216324 similar to UP|Q9XFL4 (Q9XFL4) Peroxidase 3, complete
Length = 1246
Score = 269 bits (688), Expect = 1e-72
Identities = 152/314 (48%), Positives = 194/314 (61%), Gaps = 5/314 (1%)
Frame = +2
Query: 23 SELLVHEYYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRLHFHDCFVMGCDASVLLDSV 82
S L +Y KCP V+ + A+ K+PR AS++RL FHDCFV GCD SVLLD
Sbjct: 119 SAQLSENFYDSKCPKVFYAVKSVLQSALAKEPRQGASIVRLFFHDCFVNGCDGSVLLD-- 292
Query: 83 EGMTSEKQAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCADILAMVARDAVELRGGPRWE 142
G +SEK A PN NSLRG+EVID IK +E CP VSCADI+ + ARD+V + GGP W+
Sbjct: 293 -GPSSEKTAPPNNNSLRGYEVIDAIKSKVETVCPGVVSCADIVTIAARDSVAILGGPNWK 469
Query: 143 VWLGRKDSLESSFSGANL-FIPAPNSSLETLINNFKQQGLDIEDLVVLSGSHTIGRARCL 201
V LGR+DS F+ AN +P PNSSL +LI F QGL +D+V LSG+HTIG+ARC+
Sbjct: 470 VKLGRRDSTTGFFNLANSGVLPGPNSSLSSLIQRFDDQGLSTKDMVALSGAHTIGKARCV 649
Query: 202 SFRQRIYETKQEYHHAYDRYKRYTTFRRILQSICPV----TGRDDKFAPLDFQTPKRFDN 257
S+R RI Y+ + F + Q CP T +D+ APLDF+TP FDN
Sbjct: 650 SYRDRI----------YNENNIDSLFAKARQKNCPKGSSGTPKDNNVAPLDFKTPNHFDN 799
Query: 258 QYFINIIEGKGLLGSDNVLISQDLDGRIRKQVWGYASNEKLFFDSFAKSMIKMGNINVLT 317
+YF N+I KGLL SD L + G V Y++N+++F F +MIKMGNI LT
Sbjct: 800 EYFKNLINKKGLLRSDQELFN---GGSTDSLVRTYSNNQRVFEADFVTAMIKMGNIKPLT 970
Query: 318 GSEGEIRRNCRFVN 331
GS G+IR+ CR N
Sbjct: 971 GSNGQIRKQCRRPN 1012
>TC219609 similar to UP|PER1_ARAHY (P22195) Cationic peroxidase 1 precursor
(PNPC1) , partial (79%)
Length = 1036
Score = 265 bits (678), Expect = 2e-71
Identities = 148/327 (45%), Positives = 192/327 (58%)
Frame = +2
Query: 4 LRLLLILITILSNIHTLRGSELLVHEYYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRL 63
L+ LILI+ + + + + L ++Y + CP A +R V AV + R+ ASLLRL
Sbjct: 98 LKFSLILISCVIGVTSAQ----LSSKFYDKSCPKALTTIRKEVERAVRNESRMGASLLRL 265
Query: 64 HFHDCFVMGCDASVLLDSVEGMTSEKQAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCAD 123
HFHDCFV GCDASVLLD T EK + PN NSLRGFEVID IK LE C VSCAD
Sbjct: 266 HFHDCFVQGCDASVLLDDTANFTGEKNSFPNANSLRGFEVIDNIKSKLEGMCKGVVSCAD 445
Query: 124 ILAMVARDAVELRGGPRWEVWLGRKDSLESSFSGANLFIPAPNSSLETLINNFKQQGLDI 183
ILA+ ARDAV GG +WEV +GR+DS +S AN +PAP L LI F ++
Sbjct: 446 ILAVAARDAVVALGGQKWEVQVGRRDSTTASLDEANSDLPAPFLDLSGLITAFAKKNFTT 625
Query: 184 EDLVVLSGSHTIGRARCLSFRQRIYETKQEYHHAYDRYKRYTTFRRILQSICPVTGRDDK 243
++LV LSG HTIG RC FR RI Y+ TF + +Q++CP G DD
Sbjct: 626 QELVTLSGGHTIGLVRCRFFRARI----------YNESNIDPTFAQQMQALCPFEGGDDN 775
Query: 244 FAPLDFQTPKRFDNQYFINIIEGKGLLGSDNVLISQDLDGRIRKQVWGYASNEKLFFDSF 303
+P D TP +FDN ++ N+++ KG++ SD L + + G QV Y+ N F F
Sbjct: 776 LSPFDSTTPFKFDNAFYKNLVQLKGVVHSDQQLFTNNGSGPTNDQVNRYSRNMGNFKKDF 955
Query: 304 AKSMIKMGNINVLTGSEGEIRRNCRFV 330
A +M KM + LTGS G+IR+NCR V
Sbjct: 956 ADAMFKMSMLTPLTGSNGQIRQNCRLV 1036
>TC230675 similar to PDB|1SCH_A.0|1633130|1SCH_A Chain A, Peanut Peroxidase.
{Arachis hypogaea;} , partial (93%)
Length = 1386
Score = 261 bits (666), Expect = 4e-70
Identities = 147/304 (48%), Positives = 188/304 (61%), Gaps = 2/304 (0%)
Frame = +3
Query: 30 YYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRLHFHDCFVMGCDASVLLDSVEGMTSEK 89
+Y KCPL + + V AV K+ R+ ASLLRLHFHDCFV GCDASVLL + T E+
Sbjct: 174 FYLLKCPLGLFTINNLVTAAVRKESRMGASLLRLHFHDCFVQGCDASVLLKNTATFTGEQ 353
Query: 90 QAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCADILAMVARDAVELRGGPRWEVWLGRKD 149
A PN NSLRGFEVID IK LE CP SCADILA+ ARD+V GG W+V LGR+D
Sbjct: 354 GAFPNANSLRGFEVIDNIKAKLEILCPGVFSCADILAVAARDSVVALGGLGWQVRLGRRD 533
Query: 150 SLESSFSGANLFIPAPNSSLETLINNFKQQGLDIEDLVVLSGSHTIGRARCLSFRQRIYE 209
S +S SGAN +PAP L L+ F+++G + ++V LSG+HTIG ARCL+FR R Y
Sbjct: 534 STTASLSGANSDLPAPFLGLTDLVAAFQKKGFTVNEMVALSGAHTIGSARCLTFRSRAYN 713
Query: 210 TKQEYHHAYDRYKRYTTFRRILQSICPVTGRDDKFAPLDFQTPKRFDNQYFINIIEGKGL 269
D Y F L+S CP +G DD +P+D T FDN Y+ N++ KGL
Sbjct: 714 DS-------DIEPSYANF---LRSTCPKSGGDDNLSPIDIATKDIFDNAYYRNLLYKKGL 863
Query: 270 LGSDNVLISQDL-DGRIRKQVWGYASNEKLFFDS-FAKSMIKMGNINVLTGSEGEIRRNC 327
SD L S D +++ YA+ LFF S FA +M+KM N++ LTG++G+IR+ C
Sbjct: 864 FHSDQQLYSGSFTDSKVKY----YATYPSLFFKSDFANAMLKMSNLSPLTGTQGQIRKVC 1031
Query: 328 RFVN 331
VN
Sbjct: 1032SRVN 1043
>TC204493 UP|O22443 (O22443) Seed coat peroxidase precursor , complete
Length = 1294
Score = 260 bits (664), Expect = 7e-70
Identities = 150/333 (45%), Positives = 197/333 (59%), Gaps = 2/333 (0%)
Frame = +1
Query: 1 MEPLRLLLILITILSNIHTLRGSEL--LVHEYYKEKCPLAEDIVRHNVAVAVLKDPRLAA 58
M +RLL++ + +H L +Y+E CP IV + A DPR+ A
Sbjct: 19 MGSMRLLVVALLCAFAMHAGFSVSYAQLTPTFYRETCPNLFPIVFGVIFDASFTDPRIGA 198
Query: 59 SLLRLHFHDCFVMGCDASVLLDSVEGMTSEKQAGPNVNSLRGFEVIDKIKYLLEKECPLT 118
SL+RLHFHDCFV GCD SVLL++ + + SE+ A PN+NS+RG +V++ IK +E CP T
Sbjct: 199 SLMRLHFHDCFVQGCDGSVLLNNTDTIESEQDALPNINSIRGLDVVNDIKTAVENSCPDT 378
Query: 119 VSCADILAMVARDAVELRGGPRWEVWLGRKDSLESSFSGANLFIPAPNSSLETLINNFKQ 178
VSCADILA+ A A L GGP W V LGR+DSL ++ + AN +PAP +L L +F
Sbjct: 379 VSCADILAIAAEIASVLGGGPGWPVPLGRRDSLTANRTLANQNLPAPFFNLTQLKASFAV 558
Query: 179 QGLDIEDLVVLSGSHTIGRARCLSFRQRIYETKQEYHHAYDRYKRYTTFRRILQSICPVT 238
QGL+ DLV LSG HT GRARC +F R+Y + TT+ +L++ CP
Sbjct: 559 QGLNTLDLVTLSGGHTFGRARCSTFINRLYNFS---NTGNPDPTLNTTYLEVLRARCPQN 729
Query: 239 GRDDKFAPLDFQTPKRFDNQYFINIIEGKGLLGSDNVLISQDLDGRIRKQVWGYASNEKL 298
D LD TP +FDN+Y+ N+++ GLL SD L S I V ++SN+
Sbjct: 730 ATGDNLTNLDLSTPDQFDNRYYSNLLQLNGLLQSDQELFSTPGADTI-PIVNSFSSNQNT 906
Query: 299 FFDSFAKSMIKMGNINVLTGSEGEIRRNCRFVN 331
FF +F SMIKMGNI VLTG EGEIR C FVN
Sbjct: 907 FFSNFRVSMIKMGNIGVLTGDEGEIRLQCNFVN 1005
>TC205795 similar to UP|Q9XFL3 (Q9XFL3) Peroxidase 1 (Fragment), partial
(96%)
Length = 1321
Score = 257 bits (657), Expect = 4e-69
Identities = 144/306 (47%), Positives = 186/306 (60%)
Frame = +2
Query: 26 LVHEYYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRLHFHDCFVMGCDASVLLDSVEGM 85
L + +YK+ CP IVR V DPR+ ASL+RLHFHDCFV GCDAS+LL+ +
Sbjct: 116 LDNSFYKDTCPRVHSIVREVVRNVSKSDPRILASLIRLHFHDCFVQGCDASILLNDTATI 295
Query: 86 TSEKQAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCADILAMVARDAVELRGGPRWEVWL 145
SE+ A PN NS+RG +V+++IK +E CP VSCADILA+ A + L GP W+V L
Sbjct: 296 VSEQSAPPNNNSIRGLDVVNQIKTAVENACPGIVSCADILALAAEISSVLAHGPDWKVPL 475
Query: 146 GRKDSLESSFSGANLFIPAPNSSLETLINNFKQQGLDIEDLVVLSGSHTIGRARCLSFRQ 205
GR+DSL SSFS A +P N +L+ L + F +QGL+ DLV LSG+HTIGR++C F
Sbjct: 476 GRRDSLNSSFSLALQNLPGFNFTLDQLKSTFDRQGLNTTDLVALSGAHTIGRSQCRFFAH 655
Query: 206 RIYETKQEYHHAYDRYKRYTTFRRILQSICPVTGRDDKFAPLDFQTPKRFDNQYFINIIE 265
RIY + TT + L++ICP G LD TP RFD+ Y+ N+
Sbjct: 656 RIYNFS---GNGNSDPTLNTTLSQALRAICPNGGPGTNLTNLDLTTPDRFDSNYYSNLQL 826
Query: 266 GKGLLGSDNVLISQDLDGRIRKQVWGYASNEKLFFDSFAKSMIKMGNINVLTGSEGEIRR 325
GLL SD VL S V + SN+ LF++ F SMIKM I VLTGS+GEIR+
Sbjct: 827 QNGLLRSDQVLFSTS-GAETIAIVNSFGSNQTLFYEHFKVSMIKMSIIEVLTGSQGEIRK 1003
Query: 326 NCRFVN 331
+C FVN
Sbjct: 1004HCNFVN 1021
>TC205794 similar to UP|Q9XFL3 (Q9XFL3) Peroxidase 1 (Fragment), partial
(97%)
Length = 1205
Score = 252 bits (644), Expect = 1e-67
Identities = 141/302 (46%), Positives = 185/302 (60%)
Frame = +2
Query: 30 YYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRLHFHDCFVMGCDASVLLDSVEGMTSEK 89
+Y++ CP IVR V DPR+ ASL+RLHFHDCFV GCDAS+LL++ + SE+
Sbjct: 128 FYRDTCPKVHSIVREVVRNVSKSDPRMLASLIRLHFHDCFVQGCDASILLNNTATIESEQ 307
Query: 90 QAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCADILAMVARDAVELRGGPRWEVWLGRKD 149
QA PN NS+RG +V+++IK +E CP VSCADILA+ A + L GP W+V LGR+D
Sbjct: 308 QAFPNNNSIRGLDVVNQIKTAVENACPGVVSCADILALAAEISSVLGHGPDWKVPLGRRD 487
Query: 150 SLESSFSGANLFIPAPNSSLETLINNFKQQGLDIEDLVVLSGSHTIGRARCLSFRQRIYE 209
SL ++ + AN +PAP +L L + F QGL+ DLV LSG+HTIGRA+C F R+Y
Sbjct: 488 SLTANRTLANQNLPAPFLNLTQLKDAFAVQGLNTTDLVALSGAHTIGRAQCRFFVDRLYN 667
Query: 210 TKQEYHHAYDRYKRYTTFRRILQSICPVTGRDDKFAPLDFQTPKRFDNQYFINIIEGKGL 269
TT+ + L +ICP G D TP D+ Y+ N+ KGL
Sbjct: 668 FSST---GNPDPTLNTTYLQTLSAICPNGGPGTNLTNFDPTTPDTVDSNYYSNLQVNKGL 838
Query: 270 LGSDNVLISQDLDGRIRKQVWGYASNEKLFFDSFAKSMIKMGNINVLTGSEGEIRRNCRF 329
L SD L S I V ++SN+ LFF++F SMIKMGNI VLTGS+GEIR+ C F
Sbjct: 839 LQSDQELFSTTGTDTI-AIVNSFSSNQTLFFENFKASMIKMGNIGVLTGSQGEIRQQCNF 1015
Query: 330 VN 331
+N
Sbjct: 1016IN 1021
>TC206384 UP|O23961 (O23961) Peroxidase precursor, complete
Length = 1251
Score = 250 bits (638), Expect = 7e-67
Identities = 147/324 (45%), Positives = 191/324 (58%)
Frame = +1
Query: 8 LILITILSNIHTLRGSELLVHEYYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRLHFHD 67
L + ++ + L L +Y++ CP IVR V KDPR+ ASL+RLHFHD
Sbjct: 55 LCCVVVVLGVLPLSLDAQLDPSFYRDTCPRVHSIVREVVRNVSKKDPRMLASLIRLHFHD 234
Query: 68 CFVMGCDASVLLDSVEGMTSEKQAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCADILAM 127
CFV GCDASVLL++ + SE+QA PN NSLRG +V++ IK +EK CP VSCADIL +
Sbjct: 235 CFVQGCDASVLLNNTATIESEQQALPNNNSLRGLDVVNDIKTAVEKACPGVVSCADILTL 414
Query: 128 VARDAVELRGGPRWEVWLGRKDSLESSFSGANLFIPAPNSSLETLINNFKQQGLDIEDLV 187
+ + L GGP W+V LGR+DSL ++ + AN +PAP +L L F QGLD DLV
Sbjct: 415 ASEISSVLGGGPHWKVPLGRRDSLTANRNLANQNLPAPFFNLSRLKAAFAVQGLDTTDLV 594
Query: 188 VLSGSHTIGRARCLSFRQRIYETKQEYHHAYDRYKRYTTFRRILQSICPVTGRDDKFAPL 247
LSG+HT GRA C R+Y TT+ + L+ ICP G +
Sbjct: 595 ALSGAHTFGRAHCNFILDRLYNFSGT---GKPDPTLDTTYLQQLRQICP-NGGPNNLVNF 762
Query: 248 DFQTPKRFDNQYFINIIEGKGLLGSDNVLISQDLDGRIRKQVWGYASNEKLFFDSFAKSM 307
D TP + D YF N+ KGLL SD L S I V ++S++K+FFD+F SM
Sbjct: 763 DPVTPDKIDRVYFSNLQVKKGLLQSDQELFSTPGADTI-PIVNRFSSDQKVFFDAFEASM 939
Query: 308 IKMGNINVLTGSEGEIRRNCRFVN 331
IKMGNI VLTG +GEIR++C FVN
Sbjct: 940 IKMGNIGVLTGKKGEIRKHCNFVN 1011
>TC230907 similar to UP|O24081 (O24081) Peroxidase1A precursor , partial
(88%)
Length = 944
Score = 249 bits (635), Expect = 2e-66
Identities = 138/291 (47%), Positives = 184/291 (62%)
Frame = +2
Query: 41 IVRHNVAVAVLKDPRLAASLLRLHFHDCFVMGCDASVLLDSVEGMTSEKQAGPNVNSLRG 100
IVR ++ DPR+ ASL+RLHFHDCFV GCDAS+LL+ + + SE+ A PN NS+RG
Sbjct: 5 IVREVLSNVSQSDPRILASLIRLHFHDCFVQGCDASILLNDTDTIVSEQSAVPNNNSIRG 184
Query: 101 FEVIDKIKYLLEKECPLTVSCADILAMVARDAVELRGGPRWEVWLGRKDSLESSFSGANL 160
+V+++IK +E CP VSCADILA+ A+ + +L GP W+V LGR+DSL ++ + AN
Sbjct: 185 LDVVNQIKTAVENACPGIVSCADILALAAQISSDLANGPVWQVPLGRRDSLTANQTLANQ 364
Query: 161 FIPAPNSSLETLINNFKQQGLDIEDLVVLSGSHTIGRARCLSFRQRIYETKQEYHHAYDR 220
+PAP +++ LI +F Q L+I DLV LSG+HTIGRA+C F R+Y +
Sbjct: 365 NLPAPTFTIDQLIESFGNQSLNITDLVALSGAHTIGRAQCRFFVDRLYNFS---NTGNPD 535
Query: 221 YKRYTTFRRILQSICPVTGRDDKFAPLDFQTPKRFDNQYFINIIEGKGLLGSDNVLISQD 280
TT + LQ ICP G LD TP FD+ Y+ N+ GLL SD L+S +
Sbjct: 536 PTLNTTLLQSLQGICPNGGPGTNLTNLDLTTPDTFDSNYYSNLQLQNGLLQSDQELLSAN 715
Query: 281 LDGRIRKQVWGYASNEKLFFDSFAKSMIKMGNINVLTGSEGEIRRNCRFVN 331
+ I V + SN+ LFF++F SMIKMGNI VLTGS+GEIR C VN
Sbjct: 716 -NTDIVAIVNNFISNQTLFFENFKASMIKMGNIGVLTGSQGEIRSQCNSVN 865
>TC204496 homologue to UP|O23961 (O23961) Peroxidase precursor, partial (97%)
Length = 2193
Score = 245 bits (626), Expect = 2e-65
Identities = 145/324 (44%), Positives = 191/324 (58%)
Frame = +3
Query: 8 LILITILSNIHTLRGSELLVHEYYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRLHFHD 67
L + ++ + L L +Y++ CP IVR V KDPR+ ASL+RLHFHD
Sbjct: 894 LCFVVVVVGVLPLSLDAQLDPSFYRDTCPKVHSIVREVVRNVSKKDPRMLASLIRLHFHD 1073
Query: 68 CFVMGCDASVLLDSVEGMTSEKQAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCADILAM 127
CFV GCDASVLL++ + SE+QA PN NSLRG +V++ IK +E+ CP VSCADIL +
Sbjct: 1074 CFVQGCDASVLLNNTATIESEQQALPNNNSLRGLDVVNDIKTAVEQACPGVVSCADILTL 1253
Query: 128 VARDAVELRGGPRWEVWLGRKDSLESSFSGANLFIPAPNSSLETLINNFKQQGLDIEDLV 187
+ + L GGP W+V LGR+DSL ++ + AN +PAP +L L F QGLD DLV
Sbjct: 1254 ASEISSILGGGPDWKVPLGRRDSLTANRTLANQNLPAPFFNLTQLKAAFAVQGLDTTDLV 1433
Query: 188 VLSGSHTIGRARCLSFRQRIYETKQEYHHAYDRYKRYTTFRRILQSICPVTGRDDKFAPL 247
LSG+HT GRA C R+Y TT+ + L+ ICP G +
Sbjct: 1434 ALSGAHTFGRAHCSFILGRLYNFSGT---GKPDPTLDTTYLQQLRQICP-NGGPNNLVNF 1601
Query: 248 DFQTPKRFDNQYFINIIEGKGLLGSDNVLISQDLDGRIRKQVWGYASNEKLFFDSFAKSM 307
D TP + D YF N+ KGLL SD L S I V ++S++ +FFD+F SM
Sbjct: 1602 DPVTPDKIDRVYFSNLQVKKGLLQSDQELFSTPGADTI-PIVNRFSSDQNVFFDAFEASM 1778
Query: 308 IKMGNINVLTGSEGEIRRNCRFVN 331
IKMGNI VLTG++GEIR++C FVN
Sbjct: 1779 IKMGNIGVLTGNKGEIRKHCNFVN 1850
>TC214651 similar to UP|Q9XGV6 (Q9XGV6) Bacterial-induced peroxidase
precursor , partial (79%)
Length = 920
Score = 239 bits (610), Expect = 1e-63
Identities = 131/273 (47%), Positives = 168/273 (60%), Gaps = 1/273 (0%)
Frame = +1
Query: 7 LLILITILSNIHTLRGSELLVHEYYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRLHFH 66
L ++++ILS + + L +Y + CP + IVR + AV K+ R+ AS+LRL FH
Sbjct: 121 LFVVVSILSLL-AFSSNAQLSPTFYAKTCPNLQTIVRSAMRQAVAKEARIGASILRLFFH 297
Query: 67 DCFVMGCDASVLLDSVEGMTSEKQAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCADILA 126
DCFV GCD S+LLD T EK AGPN NS RGFEVID IK +E C TVSCADILA
Sbjct: 298 DCFVNGCDGSILLDDTATFTGEKNAGPNRNSARGFEVIDTIKTNVEASCNATVSCADILA 477
Query: 127 MVARDAVELRGGPRWEVWLGRKDSLESSFSGANLFIPAPNSSLETLINNFKQQGLDIEDL 186
+ RD + L GGP W V LGR+D+ +S S AN IP P+S L TLI+ F +GL DL
Sbjct: 478 LATRDGIVLLGGPSWTVPLGRRDARTASQSAANNQIPGPSSDLSTLISMFASKGLTASDL 657
Query: 187 VVLSGSHTIGRARCLSFRQRIY-ETKQEYHHAYDRYKRYTTFRRILQSICPVTGRDDKFA 245
VLSG+HTIG+A+C FR RIY ET + T F ++ CP TG + A
Sbjct: 658 TVLSGAHTIGQAQCQFFRTRIYNETNID-----------TNFAATRKTTCPATGGNTNLA 804
Query: 246 PLDFQTPKRFDNQYFINIIEGKGLLGSDNVLIS 278
PL+ TP RFDN Y+ +++ +GLL SD VL +
Sbjct: 805 PLETLTPTRFDNNYYADLVNRRGLLHSDQVLFN 903
>TC206606 similar to UP|Q7XYR7 (Q7XYR7) Class III peroxidase, partial (68%)
Length = 787
Score = 235 bits (599), Expect = 2e-62
Identities = 127/256 (49%), Positives = 162/256 (62%), Gaps = 3/256 (1%)
Frame = +3
Query: 4 LRLLLILITILSNIHTLRGSELLVHEYYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRL 63
+ L L+++ + +N + + L +Y CP D V+ V A+ K+ R+ ASLLRL
Sbjct: 51 ITLALLVLVLGTNTSSANANPTLHTNFYYSSCPKLFDTVKRTVESAISKETRMGASLLRL 230
Query: 64 HFHDCFVMGCDASVLLDSVEGMTSEKQAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCAD 123
FHDCFV GCD S+LLD T EK AGPN NS RGFEVID+IK +EK CP VSCAD
Sbjct: 231 FFHDCFVNGCDGSILLDDTSSFTGEKNAGPNRNSARGFEVIDQIKSAVEKVCPGVVSCAD 410
Query: 124 ILAMVARDAVELRGGPRWEVWLGRKDSLESSFSGANLFIPAPNSSLETLINNFKQQGLDI 183
ILA+ ARD+VE+ GGP W+V LGR+DS +S S AN IP P S+L LI+ F GL
Sbjct: 411 ILAIAARDSVEILGGPTWDVKLGRRDSRTASQSAANNDIPRPTSNLNQLISRFNALGLST 590
Query: 184 EDLVVLSGSHTIGRARCLSFRQRIY-ETKQEYHHAYDRYKRYTTFRRILQSICPVT--GR 240
+DLV LSG HTIG+ARC +FR RIY ET + ++F R+ QS CP T R
Sbjct: 591 KDLVALSGGHTIGQARCTTFRARIYNETNID-----------SSFARMRQSKCPQTSRSR 737
Query: 241 DDKFAPLDFQTPKRFD 256
D+ AP++F TP+ FD
Sbjct: 738 DNNLAPINFTTPRFFD 785
>TC218639 similar to UP|PE17_ARATH (Q9SJZ2) Peroxidase 17 precursor (Atperox
P17) (ATP25a) , partial (93%)
Length = 1085
Score = 232 bits (592), Expect = 2e-61
Identities = 132/326 (40%), Positives = 193/326 (58%), Gaps = 2/326 (0%)
Frame = +2
Query: 8 LILITILSNIHTLRGSELLVHEYYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRLHFHD 67
L L+ ++ +I L S L +Y + CP AE IVR + A++++ R AS++R FHD
Sbjct: 5 LFLMFLVLHIAWLVASSDLRAGFYSKTCPKAEVIVRDVMKKALMREARSVASVMRFQFHD 184
Query: 68 CFVMGCDASVLLDSVEGMTSEKQAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCADILAM 127
CFV GCD S+LLD M EK A N+NSLR ++V+D++K LEK+CP VSCADI+ M
Sbjct: 185 CFVNGCDGSMLLDDTATMLGEKMALSNINSLRSYKVVDQVKQALEKDCPGVVSCADIIIM 364
Query: 128 VARDAVELRGGPRWEVWLGRKDSLESSFSGANLFIPAPNSSLETLINNFKQQGLDIEDLV 187
+RDAV L GGP WEV LGR DSL ++ +N +P+P ++ +LI+ F++ L ++DLV
Sbjct: 365 ASRDAVALTGGPEWEVRLGRLDSLSANQEDSNNIMPSPRANASSLIDLFQKYNLTVKDLV 544
Query: 188 VLSGSHTIGRARCLSFRQRIYETK--QEYHHAYDRYKRYTTFRRILQSICPVTGRDDKFA 245
LSGSH+IG+ RC S R+Y A D ++R+ L +CP+ +
Sbjct: 545 ALSGSHSIGQGRCFSVMFRLYNQSGTGRPDPAID-----PSYRQYLNRLCPLDVDQNVTG 709
Query: 246 PLDFQTPKRFDNQYFINIIEGKGLLGSDNVLISQDLDGRIRKQVWGYASNEKLFFDSFAK 305
LD TP FDNQY ++ +G L SD L + R+ V ++ + FF ++ +
Sbjct: 710 NLD-STPLVFDNQYIKDLAARRGFLNSDQTLFTFP---HTREFVRLFSRRKTEFFKAYVE 877
Query: 306 SMIKMGNINVLTGSEGEIRRNCRFVN 331
M+KMG++ +G GE+R NCR VN
Sbjct: 878 GMLKMGDLQ--SGRPGEVRTNCRLVN 949
>TC204494 similar to UP|O23961 (O23961) Peroxidase precursor, partial (93%)
Length = 1326
Score = 230 bits (586), Expect = 8e-61
Identities = 137/302 (45%), Positives = 176/302 (57%)
Frame = +3
Query: 30 YYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRLHFHDCFVMGCDASVLLDSVEGMTSEK 89
+YK+ CP IV V DPR+ ASL+RL FHDCFV GCDAS+LL++ + SE+
Sbjct: 165 FYKKTCPQVHFIVFKVVEKVSRTDPRMPASLVRLFFHDCFVQGCDASILLNNTATIVSEQ 344
Query: 90 QAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCADILAMVARDAVELRGGPRWEVWLGRKD 149
QA PN NS+RG +V+++IK LEK CP VSCADIL + A + L GP + LGR+D
Sbjct: 345 QALPNNNSIRGLDVVNQIKTELEKACPGVVSCADILTLAAEVSSVLAHGPYLKFPLGRRD 524
Query: 150 SLESSFSGANLFIPAPNSSLETLINNFKQQGLDIEDLVVLSGSHTIGRARCLSFRQRIYE 209
SL ++ + AN +PAP +L L F QGLD DLV LSG+H+ GR RCL R+Y
Sbjct: 525 SLTANRTLANQNLPAPFFNLTQLKAAFAVQGLDTTDLVALSGAHSFGRVRCLFILDRLYN 704
Query: 210 TKQEYHHAYDRYKRYTTFRRILQSICPVTGRDDKFAPLDFQTPKRFDNQYFINIIEGKGL 269
TT+ + L+ ICP G + D TP D Y+ N+ KGL
Sbjct: 705 FSGT---GRPDPTLDTTYLKQLRQICPQGGPPNNLVNFDPTTPDTLDKNYYSNLQVKKGL 875
Query: 270 LGSDNVLISQDLDGRIRKQVWGYASNEKLFFDSFAKSMIKMGNINVLTGSEGEIRRNCRF 329
L SD L S I V ++S++ FF SF+ SMIKMGNI VLTG +GEIR+ C F
Sbjct: 876 LQSDQELFSTPGADTI-SIVNKFSSDQIAFFKSFSASMIKMGNIGVLTGKKGEIRKQCNF 1052
Query: 330 VN 331
VN
Sbjct: 1053VN 1058
>TC216945 similar to UP|Q43854 (Q43854) Peroxidase precursor , partial (96%)
Length = 1274
Score = 229 bits (585), Expect = 1e-60
Identities = 136/336 (40%), Positives = 195/336 (57%), Gaps = 12/336 (3%)
Frame = +2
Query: 8 LILITILSNIHTLRGSEL---------LVHEYYKEKCPLAEDIVRHNVAVAVLKDPRLAA 58
L+LI+I +++ + E L + +Y + CP + IVR + KD AA
Sbjct: 29 LLLISIFLSVYNIEVCEAQARPPTAKGLSYTFYDKSCPKLKSIVRSELKKVFNKDIAQAA 208
Query: 59 SLLRLHFHDCFVMGCDASVLLDSVEGMTSEKQAGPNVNSLR--GFEVIDKIKYLLEKECP 116
LLRLHFHDCFV GCD SVLLD EK+A PN+ +LR F++I+ ++ LLEK C
Sbjct: 209 GLLRLHFHDCFVQGCDGSVLLDGSASGPGEKEAPPNL-TLRPEAFKIIENLRGLLEKSCG 385
Query: 117 LTVSCADILAMVARDAVELRGGPRWEVWLGRKDSLESSFSGANL-FIPAPNSSLETLINN 175
VSC+DI A+ ARDAV L GGP +E+ LGR+D L + L +P P+S+ T++++
Sbjct: 386 RVVSCSDITALTARDAVFLSGGPDYEIPLGRRDGLTFATRQVTLDNLPPPSSNASTILSS 565
Query: 176 FKQQGLDIEDLVVLSGSHTIGRARCLSFRQRIYETKQEYHHAYDRYKRYTTFRRILQSIC 235
+ LD D+V LSG HTIG + C SF R+Y T+ D+ TF L+ C
Sbjct: 566 LATKNLDPTDVVALSGGHTIGISHCGSFTNRLYPTQDP---VMDK-----TFGNNLRRTC 721
Query: 236 PVTGRDDKFAPLDFQTPKRFDNQYFINIIEGKGLLGSDNVLISQDLDGRIRKQVWGYASN 295
P D+ LD ++P FDN+Y+++++ +GL SD L + + R + V +A N
Sbjct: 722 PAANTDNT-TVLDIRSPNTFDNKYYVDLMNRQGLFTSDQDLYT---NTRTKGIVTDFAVN 889
Query: 296 EKLFFDSFAKSMIKMGNINVLTGSEGEIRRNCRFVN 331
+ LFFD F +M+KMG +NVLTG++GEIR NC N
Sbjct: 890 QSLFFDKFVFAMLKMGQLNVLTGNQGEIRANCSVRN 997
>TC204669 similar to UP|PER2_ARAHY (P22196) Cationic peroxidase 2 precursor
(PNPC2) , partial (81%)
Length = 1413
Score = 227 bits (579), Expect = 5e-60
Identities = 139/302 (46%), Positives = 178/302 (58%)
Frame = +2
Query: 30 YYKEKCPLAEDIVRHNVAVAVLKDPRLAASLLRLHFHDCFVMGCDASVLLDSVEGMTSEK 89
+Y CP AE IVR V + DP LA +LR+HFHDCFV GCDASVL + G +E+
Sbjct: 257 FYSSTCPRAESIVRSTVESHLRSDPTLAGPILRMHFHDCFVRGCDASVL---IAGAGTER 427
Query: 90 QAGPNVNSLRGFEVIDKIKYLLEKECPLTVSCADILAMVARDAVELRGGPRWEVWLGRKD 149
AGPN+ SLRGF+VID K +E CP VSCADIL++ ARD+V L GG W+V GRKD
Sbjct: 428 TAGPNL-SLRGFDVIDDAKAKIEALCPGVVSCADILSLAARDSVVLSGGLSWQVPTGRKD 604
Query: 150 SLESSFSGANLFIPAPNSSLETLINNFKQQGLDIEDLVVLSGSHTIGRARCLSFRQRIYE 209
S S A L +P PN ++ T + F +GL+ EDLV+L+G HTIG + C SF RIY
Sbjct: 605 GRVSIGSEA-LTLPGPNDTVATQKDKFSNKGLNTEDLVILAGGHTIGTSACRSFADRIYN 781
Query: 210 TKQEYHHAYDRYKRYTTFRRILQSICPVTGRDDKFAPLDFQTPKRFDNQYFINIIEGKGL 269
+ D +F L+ ICP T + K LD + +FD YF +++ G+G+
Sbjct: 782 PNGT-DPSID-----PSFLPFLRQICPQT-QPTKRVALDTGSQFKFDTSYFAHLVRGRGI 940
Query: 270 LGSDNVLISQDLDGRIRKQVWGYASNEKLFFDSFAKSMIKMGNINVLTGSEGEIRRNCRF 329
L SD VL + D R V Y + F F KSMIKM NI V TGS+GEIR+ C
Sbjct: 941 LRSDQVLWT---DASTRGFVQKYLATGP-FKVQFGKSMIKMSNIGVKTGSQGEIRKICSA 1108
Query: 330 VN 331
+N
Sbjct: 1109IN 1114
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.323 0.140 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,445,319
Number of Sequences: 63676
Number of extensions: 185736
Number of successful extensions: 1263
Number of sequences better than 10.0: 204
Number of HSP's better than 10.0 without gapping: 1041
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1056
length of query: 332
length of database: 12,639,632
effective HSP length: 98
effective length of query: 234
effective length of database: 6,399,384
effective search space: 1497455856
effective search space used: 1497455856
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 59 (27.3 bits)
Medicago: description of AC144516.5


