

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144513.4 - phase: 0 /pseudo
(1015 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
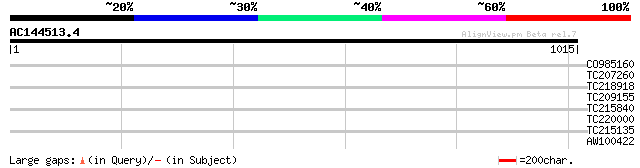
Score E
Sequences producing significant alignments: (bits) Value
CO985160 34 0.28
TC207260 homologue to UP|IF2C_PHAVU (P57997) Translation initiat... 32 1.8
TC218918 UP|Q39892 (Q39892) Nucleosome assembly protein 1, complete 31 2.4
TC209155 31 3.1
TC215840 homologue to UP|Q41122 (Q41122) Proline-rich protein pr... 30 4.1
TC220000 weakly similar to UP|O95199 (O95199) RCC1-like G exchan... 30 5.4
TC215135 similar to GB|AAM65351.1|21655279|AY103299 AT3g27090/MO... 30 5.4
AW100422 similar to GP|5929964|gb| malate dehydrogenase {Glycine... 30 7.0
>CO985160
Length = 777
Score = 34.3 bits (77), Expect = 0.28
Identities = 28/73 (38%), Positives = 40/73 (54%)
Frame = +3
Query: 370 MRSLLPILRRCLLVLDLLFPLMLWRWLVTELIMICLVTLQLNSLRKRLWLLFFNLRGPLL 429
+R LL + R LL+LDLL L+ R L+ L+ + + L L LRK L LL LR LL
Sbjct: 426 LRKLLLLQLRKLLLLDLLLLLLQLRKLLLLLLQLRKLLLLLLQLRK-LLLLLLELRKLLL 602
Query: 430 LVQMV*HLFFTII 442
L+ + L ++
Sbjct: 603 LLLQLRKLLLLLL 641
Score = 29.6 bits (65), Expect = 7.0
Identities = 20/65 (30%), Positives = 35/65 (53%), Gaps = 1/65 (1%)
Frame = +3
Query: 687 LNTKLFLARKLILISQIWSLAQIP-VLVLKIYSELISLSTSRIIFLSIWGFLLSLEDLKL 745
L +L RKL+L+ +W L ++ +L+L +L+ L +++ L + LL L L L
Sbjct: 333 LRLQLL*LRKLLLLL*LWELLKLR*LLLLLDLRKLLLLQLRKLLLLDLLLLLLQLRKLLL 512
Query: 746 KILSL 750
+L L
Sbjct: 513 LLLQL 527
>TC207260 homologue to UP|IF2C_PHAVU (P57997) Translation initiation factor
IF-2, chloroplast precursor (PvIF2cp), partial (21%)
Length = 1107
Score = 31.6 bits (70), Expect = 1.8
Identities = 11/28 (39%), Positives = 18/28 (64%)
Frame = +3
Query: 395 WLVTELIMICLVTLQLNSLRKRLWLLFF 422
WL +ICL+ ++LN + K WL+F+
Sbjct: 1023 WLKNICSLICLLMIRLNEIEKAYWLMFY 1106
>TC218918 UP|Q39892 (Q39892) Nucleosome assembly protein 1, complete
Length = 1507
Score = 31.2 bits (69), Expect = 2.4
Identities = 21/59 (35%), Positives = 30/59 (50%)
Frame = -2
Query: 387 LFPLMLWRWLVTELIMICLVTLQLNSLRKRLWLLFFNLRGPLLLVQMV*HLFFTIIIGI 445
L L LWR L + +CL TL L SL L + F L +L+ ++ LF II+ +
Sbjct: 1173 LLLLALWRSLTITITSLCLTTLLLFSLSVILLFIVFVLVILFVLIFVLIFLFVLIILQV 997
>TC209155
Length = 694
Score = 30.8 bits (68), Expect = 3.1
Identities = 22/77 (28%), Positives = 41/77 (52%), Gaps = 18/77 (23%)
Frame = +1
Query: 360 KKFGEITITFMRS--------LLPILRRCL--------LVLDLLFPLMLWRWLV--TELI 401
+++GE TI F+R + I CL L+LDL P+++W+ L+ + L+
Sbjct: 427 ERYGEFTILFLRIYQRLK*LLVYSIFVVCLERSIHEAHLILDLFRPILVWKNLIV*SHLV 606
Query: 402 MICLVTLQLNSLRKRLW 418
+I + QL S+++ L+
Sbjct: 607 IISCLFHQLKSVQELLY 657
>TC215840 homologue to UP|Q41122 (Q41122) Proline-rich protein precursor,
partial (74%)
Length = 1171
Score = 30.4 bits (67), Expect = 4.1
Identities = 36/116 (31%), Positives = 58/116 (49%), Gaps = 3/116 (2%)
Frame = +3
Query: 365 ITITFMRSLLPILRRCLLVLDLLFPLMLWRWLVTELIMICLVTLQLNSLRKRLWLLFFNL 424
ITIT + L+ +L R +L L+LLF L +L ++ TLQLN L + LL NL
Sbjct: 348 ITITLLLLLIHLLFRFILQLNLLFRFTL------QLNLLFRFTLQLNLLFRLTLLL--NL 503
Query: 425 RGPLLLVQMV*HLF-FTIIIGI**GRIFYYPPSMF*ITMVTLIWSILP--ILALFP 477
R LF FT+++ + R+F + P + ++ ++ + P +L L P
Sbjct: 504 R-----------LFRFTLLLNL---RLFRFTPFLEALSQFKELFMLSPASMLGLTP 629
>TC220000 weakly similar to UP|O95199 (O95199) RCC1-like G exchanging factor
RLG (Chromosome condensation 1-like), partial (6%)
Length = 985
Score = 30.0 bits (66), Expect = 5.4
Identities = 19/60 (31%), Positives = 30/60 (49%)
Frame = +2
Query: 356 LTHMKKFGEITITFMRSLLPILRRCLLVLDLLFPLMLWRWLVTELIMICLVTLQLNSLRK 415
L H + + + + + L +LR L +L + MLWR V +L ICL+ LN R+
Sbjct: 557 LIH*RSYASLYLQIV*LYLMLLR--LRILHIELDQMLWRAFVEDLESICLLVELLNKRRR 730
>TC215135 similar to GB|AAM65351.1|21655279|AY103299 AT3g27090/MOJ10_18
{Arabidopsis thaliana;} , partial (74%)
Length = 1358
Score = 30.0 bits (66), Expect = 5.4
Identities = 16/35 (45%), Positives = 26/35 (73%)
Frame = -2
Query: 16 KLIKNLSHVFSMVLSLILLLLIVLLVIAIDQVVLP 50
K++ LS F ++LSL+L+LL VLL+I++ +LP
Sbjct: 514 KVLNLLSVAFVLLLSLVLMLLPVLLLISLLC*ILP 410
>AW100422 similar to GP|5929964|gb| malate dehydrogenase {Glycine max},
partial (46%)
Length = 484
Score = 29.6 bits (65), Expect = 7.0
Identities = 13/44 (29%), Positives = 26/44 (58%)
Frame = +3
Query: 365 ITITFMRSLLPILRRCLLVLDLLFPLMLWRWLVTELIMICLVTL 408
+ + F+ S+L ++RRC+L+L + + + WL +CL+ L
Sbjct: 348 LVMIFLTSMLALIRRCVLLLLSIVLMYMVSWLAML*TPLCLLLL 479
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.368 0.167 0.619
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 51,438,812
Number of Sequences: 63676
Number of extensions: 842156
Number of successful extensions: 13899
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 5922
Number of HSP's successfully gapped in prelim test: 681
Number of HSP's that attempted gapping in prelim test: 7475
Number of HSP's gapped (non-prelim): 7455
length of query: 1015
length of database: 12,639,632
effective HSP length: 107
effective length of query: 908
effective length of database: 5,826,300
effective search space: 5290280400
effective search space used: 5290280400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.7 bits)
S2: 64 (29.3 bits)
Medicago: description of AC144513.4


