

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144513.2 - phase: 0
(215 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
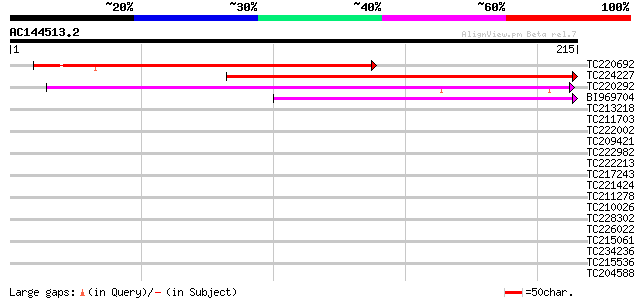
Score E
Sequences producing significant alignments: (bits) Value
TC220692 similar to UP|Q9M342 (Q9M342) Protein kinase-like prote... 205 1e-53
TC224227 199 8e-52
TC220292 weakly similar to UP|O94096 (O94096) Kexin, partial (4%) 155 2e-38
BI969704 50 8e-07
TC213218 38 0.004
TC211703 37 0.009
TC222002 34 0.057
TC209421 33 0.075
TC222982 UP|Q9SJF7 (Q9SJF7) T27G7.3, partial (5%) 32 0.22
TC222213 32 0.29
TC217243 similar to UP|Q8L8V0 (Q8L8V0) Transcription co-activato... 31 0.37
TC221424 30 0.64
TC211278 weakly similar to GB|BAB02861.1|9294580|AB026644 recept... 30 0.83
TC210026 homologue to UP|Q9LXT8 (Q9LXT8) Pre-mRNA splicing facto... 28 2.4
TC228302 homologue to UP|DCP1_PEA (P51850) Pyruvate decarboxylas... 28 2.4
TC226022 28 3.2
TC215061 similar to UP|O24550 (O24550) Malate dehydrogenase , p... 28 3.2
TC234236 similar to UP|Q9LW21 (Q9LW21) Arabidopsis thaliana geno... 28 4.1
TC215536 similar to UP|SNA2_ARATH (Q9SPE6) Alpha-soluble NSF att... 27 7.0
TC204588 similar to UP|Q7X9G7 (Q7X9G7) Alpha-L-arabinofuranosida... 27 7.0
>TC220692 similar to UP|Q9M342 (Q9M342) Protein kinase-like protein, partial
(3%)
Length = 432
Score = 205 bits (521), Expect = 1e-53
Identities = 104/133 (78%), Positives = 112/133 (84%), Gaps = 3/133 (2%)
Frame = +1
Query: 10 KMVILLLFLLITGLGSVINASP---CSNCGDFEVPYPLSTNDDCGDKRYKIYCNNDSLEF 66
KM+ LLFLL T LG VINAS CSNCGDF VPYPLSTN+DCGD RYK+YCN+ +LEF
Sbjct: 37 KMIAFLLFLL-TSLGLVINASTFSACSNCGDFVVPYPLSTNEDCGDNRYKVYCNDGNLEF 213
Query: 67 LSATGTYYKILKIDTSANKLVIKPPNIFKHTCYSSDLIGGGLVLDESLPFNISTLNTVML 126
LSATGTYYKIL+ID SANKLVI PP I K+TCYSSDL GGL+LDESLPFNIST NTVML
Sbjct: 214 LSATGTYYKILRIDPSANKLVISPPPILKNTCYSSDLYMGGLLLDESLPFNISTQNTVML 393
Query: 127 LNCSDNILQSPLN 139
NCS NILQSPLN
Sbjct: 394 FNCSYNILQSPLN 432
>TC224227
Length = 719
Score = 199 bits (506), Expect = 8e-52
Identities = 97/133 (72%), Positives = 103/133 (76%)
Frame = +3
Query: 83 ANKLVIKPPNIFKHTCYSSDLIGGGLVLDESLPFNISTLNTVMLLNCSDNILQSPLNCSS 142
ANKLVI I K SSDL GGL+LDESLPFNI VML NCS NILQSPLNCSS
Sbjct: 3 ANKLVISXXPILKXXXXSSDLYMGGLLLDESLPFNIFXXXXVMLFNCSYNILQSPLNCSS 182
Query: 143 NSICRQFEEKVEEGNGCMNTLCCHYLKDSVMNSHKIRLRVGSCTAYTCLVDFKPDEPFKK 202
NSICRQFEEKVEEG GCM TLCCHYLKDS MN KIR++VGSC AYTCLV FKP++ +
Sbjct: 183 NSICRQFEEKVEEGTGCMXTLCCHYLKDSAMNFPKIRVKVGSCPAYTCLVGFKPNDQLET 362
Query: 203 WNYGIELQWKPPN 215
WNYGIELQW PN
Sbjct: 363 WNYGIELQWLSPN 401
>TC220292 weakly similar to UP|O94096 (O94096) Kexin, partial (4%)
Length = 1094
Score = 155 bits (391), Expect = 2e-38
Identities = 76/203 (37%), Positives = 114/203 (55%), Gaps = 3/203 (1%)
Frame = +3
Query: 15 LLFLLITGLGSVINASPCSNCGDFEVPYPLSTNDDCGDKRYKIYCNNDSLEFLSATGTYY 74
LL LL++ V +A+ C CG+ VP+PLST CGD YKI C++ + Y
Sbjct: 309 LLLLLLSRATHVSSATLCPPCGNTTVPFPLSTTPTCGDPSYKIRCSSSNTLVFDTLNNSY 488
Query: 75 KILKIDTSANKLVIKPPNIFKHTCYSSDLIGGGLVLDESLPFNISTLNTVMLLNCSDNIL 134
I ID ++ + VI+P + +TC S+D + G+ L+ +LPFNI++ NT++ LNC+ +L
Sbjct: 489 PIESIDPNSQRFVIRPAPLLTNTCVSTDKVHQGIQLNTTLPFNITSSNTIVYLNCTTTLL 668
Query: 135 QSPLNCSSNSICRQFEEKVEEGNGCMNT--LCCHYLKDSVMNSHKIRLRVGSCTAYTCLV 192
QSPLNCS+ S C + + C LCC Y NS+ +R+R C+AY+ V
Sbjct: 669 QSPLNCSAASACHSYIKATASAAACQGAGPLCCTYRTGGSSNSYMLRVRDSGCSAYSSFV 848
Query: 193 DFKPDEPFKKW-NYGIELQWKPP 214
+ P P +W G+E+QW P
Sbjct: 849 NLNPALPVNRWPEPGLEIQWLSP 917
>BI969704
Length = 612
Score = 50.1 bits (118), Expect = 8e-07
Identities = 41/115 (35%), Positives = 51/115 (43%)
Frame = -1
Query: 101 SDLIGGGLVLDESLPFNISTLNTVMLLNCSDNILQSPLNCSSNSICRQFEEKVEEGNGCM 160
SDL L+ + SLPFNI V LN S N L+SP N SN FE+K ++ G
Sbjct: 612 SDLXRXXLLXNXSLPFNIXPQTRVRWLNGSKNTLRSPWNGFSNGXAGHFEKKXKKELGGR 433
Query: 161 NTLCCHYLKDSVMNSHKIRLRVGSCTAYTCLVDFKPDEPFKKWNYGIELQWKPPN 215
L+ S M S K R+R GS V K + + G E Q PN
Sbjct: 432 APFGSLSLRASPMISPKFRVRFGSAPVNPGWVGSKLNAHLELGI*GFEFQGWFPN 268
>TC213218
Length = 835
Score = 37.7 bits (86), Expect = 0.004
Identities = 26/119 (21%), Positives = 51/119 (42%), Gaps = 9/119 (7%)
Frame = +1
Query: 34 NCGDFEV-PYPLSTNDD---CGDKRYKIYCNNDSLEFLSATGTYYKILKIDTSANKLVIK 89
NCG + YP + D CG ++ + C N + L+ + Y+++ ID+ + L +
Sbjct: 172 NCGSITILSYPFTGGDRPSFCGPPQFLLNCRNGVVAELNISSVSYRVIDIDSEDHTLTLA 351
Query: 90 PPNIFKHTCYSSDLIGGGLVLDESLPFNISTLNTVMLLNC--SDNILQSP---LNCSSN 143
+++ TC +D+ + N + C + I+++P NC SN
Sbjct: 352 RLDLWNETC--TDVYVNSTFDGPVFSYGSGNQNLTLFYECKPTSRIIETPENLFNCWSN 522
>TC211703
Length = 683
Score = 36.6 bits (83), Expect = 0.009
Identities = 22/77 (28%), Positives = 37/77 (47%), Gaps = 4/77 (5%)
Frame = +3
Query: 28 NASPCSNCGDF-EVPYPLSTNDD---CGDKRYKIYCNNDSLEFLSATGTYYKILKIDTSA 83
+A P S+CG + YP D CGD RY++ C N+ S +G Y+ + I+ +
Sbjct: 9 HACPPSSCGKITNITYPFRLRGDPKGCGDNRYELACENNVTVLYSYSGKYH-VQAINYNN 185
Query: 84 NKLVIKPPNIFKHTCYS 100
+ + P + + C S
Sbjct: 186 FTIRVVDPGVQQPNCSS 236
>TC222002
Length = 726
Score = 33.9 bits (76), Expect = 0.057
Identities = 18/44 (40%), Positives = 24/44 (53%)
Frame = +1
Query: 124 VMLLNCSDNILQSPLNCSSNSICRQFEEKVEEGNGCMNTLCCHY 167
V+ +D SPL C+S+S+C QFE K + TLC HY
Sbjct: 130 VLFAT*TDTFCYSPL-CASDSVCPQFEVKCFDSLT*NRTLCLHY 258
>TC209421
Length = 1028
Score = 33.5 bits (75), Expect = 0.075
Identities = 27/97 (27%), Positives = 44/97 (44%), Gaps = 6/97 (6%)
Frame = +3
Query: 11 MVILLLFLL--ITGLGSVINASPCSNCGDF-EVPYPLSTND---DCGDKRYKIYCNNDSL 64
+VILLL L+ I + P S+CG + YP CGD RY++ C N+
Sbjct: 15 LVILLLLLVQQICATKKQDHGCPLSSCGKITNITYPFRLKGHPKSCGDNRYELACENNVT 194
Query: 65 EFLSATGTYYKILKIDTSANKLVIKPPNIFKHTCYSS 101
+G Y+ + I+ + + + P + + T SS
Sbjct: 195 VLHLYSGKYH-VQAINYNNFTIRVVDPGVDQQTNCSS 302
>TC222982 UP|Q9SJF7 (Q9SJF7) T27G7.3, partial (5%)
Length = 1161
Score = 32.0 bits (71), Expect = 0.22
Identities = 27/85 (31%), Positives = 36/85 (41%), Gaps = 10/85 (11%)
Frame = +1
Query: 8 SLKMVILLLFLLITGLGSVINASPCSNCGDF-EVPYPLSTNDD---CGDKRYKIYCNNDS 63
S+ ILLLF P S+CG + +P DD CGD RY++ C N++
Sbjct: 178 SIFAAILLLFQPNCIAKQQHQPCPPSSCGIIGNISFPFRLKDDPSHCGDTRYQLDCVNNA 357
Query: 64 LEFLSATGTY------YKILKIDTS 82
+G Y YK KI S
Sbjct: 358 TLLTLFSGKYHVQDIDYKRYKIKVS 432
>TC222213
Length = 592
Score = 31.6 bits (70), Expect = 0.29
Identities = 21/60 (35%), Positives = 29/60 (48%)
Frame = -2
Query: 84 NKLVIKPPNIFKHTCYSSDLIGGGLVLDESLPFNISTLNTVMLLNCSDNILQSPLNCSSN 143
NKL +KP F TC+S I + L PF S N ++ +N + I+ S CS N
Sbjct: 417 NKLTLKPKFCFVFTCHSFSSI---INLQSLPPFLFS--NNLLYMNRNKTIVNSKTTCSRN 253
>TC217243 similar to UP|Q8L8V0 (Q8L8V0) Transcription co-activator-like
protein, complete
Length = 934
Score = 31.2 bits (69), Expect = 0.37
Identities = 17/60 (28%), Positives = 30/60 (49%)
Frame = -2
Query: 119 STLNTVMLLNCSDNILQSPLNCSSNSICRQFEEKVEEGNGCMNTLCCHYLKDSVMNSHKI 178
+TLNT++L S ++LQ+ L+ ++ + C N C L+D + S+KI
Sbjct: 843 NTLNTILLFLISLHMLQNSLHQEHETVMSSDHFQGSRTTSCFNLFIC*NLEDKIHFSNKI 664
>TC221424
Length = 918
Score = 30.4 bits (67), Expect = 0.64
Identities = 14/36 (38%), Positives = 19/36 (51%)
Frame = +3
Query: 32 CSNCGDFEVPYPLSTNDDCGDKRYKIYCNNDSLEFL 67
C CGD + L+ + C D IYCN+D L+ L
Sbjct: 81 CDICGDQGIEEYLAICNKCPDGAEHIYCNDDKLDKL 188
>TC211278 weakly similar to GB|BAB02861.1|9294580|AB026644 receptor-like
protein kinase-like protein {Arabidopsis thaliana;} ,
partial (15%)
Length = 875
Score = 30.0 bits (66), Expect = 0.83
Identities = 22/88 (25%), Positives = 39/88 (44%)
Frame = +3
Query: 76 ILKIDTSANKLVIKPPNIFKHTCYSSDLIGGGLVLDESLPFNISTLNTVMLLNCSDNILQ 135
++ ID S+NKL P + Y ++L+ L S+P +LN + + ++N L
Sbjct: 492 LVSIDLSSNKLSGHIPPTIVNCSYLNELVLSNNQLSGSIPSEFGSLNRLKKFSVANNRLS 671
Query: 136 SPLNCSSNSICRQFEEKVEEGNGCMNTL 163
+ I F+ + EGN N +
Sbjct: 672 GTI----PEIFNGFDREGFEGNSKKNDI 743
>TC210026 homologue to UP|Q9LXT8 (Q9LXT8) Pre-mRNA splicing factor
ATP-dependent RNA helicase-like protein, partial (20%)
Length = 750
Score = 28.5 bits (62), Expect = 2.4
Identities = 13/36 (36%), Positives = 21/36 (58%)
Frame = +3
Query: 1 MYPCIWNSLKMVILLLFLLITGLGSVINASPCSNCG 36
M+ C N LK+ I L+ + L +I + PC++CG
Sbjct: 87 MWSCC*NLLKLKIYLILISWIHLLKIIFSIPCTSCG 194
>TC228302 homologue to UP|DCP1_PEA (P51850) Pyruvate decarboxylase isozyme 1
(PDC) , partial (42%)
Length = 1014
Score = 28.5 bits (62), Expect = 2.4
Identities = 15/48 (31%), Positives = 24/48 (49%)
Frame = -2
Query: 133 ILQSPLNCSSNSICRQFEEKVEEGNGCMNTLCCHYLKDSVMNSHKIRL 180
I+ NC ++ I + + + N C N LCC YLK ++ N+ L
Sbjct: 506 IMNLNFNCITSIIDEKDDGLLSTANHC*NILCC-YLKTAITNASNYTL 366
>TC226022
Length = 1066
Score = 28.1 bits (61), Expect = 3.2
Identities = 13/39 (33%), Positives = 23/39 (58%), Gaps = 2/39 (5%)
Frame = +2
Query: 161 NTLCCHY--LKDSVMNSHKIRLRVGSCTAYTCLVDFKPD 197
+TLC Y + + ++S+ R GSC ++C V F+P+
Sbjct: 233 STLCMEY*YMGEKKISSYTTTRRKGSCW*HSCAVPFRPN 349
>TC215061 similar to UP|O24550 (O24550) Malate dehydrogenase , partial (58%)
Length = 1146
Score = 28.1 bits (61), Expect = 3.2
Identities = 14/40 (35%), Positives = 23/40 (57%)
Frame = +2
Query: 96 HTCYSSDLIGGGLVLDESLPFNISTLNTVMLLNCSDNILQ 135
++C SS L+ +++ F++ L T MLLNC N+ Q
Sbjct: 320 NSCKSS*LLSSK-TMEKKFLFSLKILQTTMLLNCLQNMAQ 436
>TC234236 similar to UP|Q9LW21 (Q9LW21) Arabidopsis thaliana genomic DNA,
chromosome 3, P1 clone: MDJ14, partial (4%)
Length = 586
Score = 27.7 bits (60), Expect = 4.1
Identities = 14/35 (40%), Positives = 22/35 (62%), Gaps = 2/35 (5%)
Frame = +2
Query: 8 SLKMVILLLFLLITGLGSVINASPCSN--CGDFEV 40
S M+I +L LLI+ +G + N S CS+ C D+ +
Sbjct: 356 SCAMLISILVLLISNIGYIFNLSFCSHDICADWHL 460
>TC215536 similar to UP|SNA2_ARATH (Q9SPE6) Alpha-soluble NSF attachment
protein 2 (Alpha-SNAP2) (N-ethylmaleimide-sensitive
factor attachment protein, alpha 2), complete
Length = 1264
Score = 26.9 bits (58), Expect = 7.0
Identities = 12/33 (36%), Positives = 16/33 (48%)
Frame = -3
Query: 117 NISTLNTVMLLNCSDNILQSPLNCSSNSICRQF 149
NIS L + +NCS N+ Q + S QF
Sbjct: 968 NISKLRNIFFINCSSNVRQKSIFTCSRKCWIQF 870
>TC204588 similar to UP|Q7X9G7 (Q7X9G7) Alpha-L-arabinofuranosidase, partial
(84%)
Length = 2003
Score = 26.9 bits (58), Expect = 7.0
Identities = 16/67 (23%), Positives = 37/67 (54%), Gaps = 3/67 (4%)
Frame = +2
Query: 38 FEVPYPLSTNDDCGDKRYKIYCNNDSLEFLSATGTYYKILKIDTS---ANKLVIKPPNIF 94
F++ Y N+DCG K Y+ + L+F SA + Y ++I ++ +++ + P +++
Sbjct: 938 FDLRYVAVGNEDCGKKNYR----GNYLKFYSAIKSAYPDIQIISNCDGSSRPLDHPADMY 1105
Query: 95 KHTCYSS 101
+ Y++
Sbjct: 1106 DYHVYTN 1126
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.321 0.139 0.446
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,563,908
Number of Sequences: 63676
Number of extensions: 214518
Number of successful extensions: 1125
Number of sequences better than 10.0: 43
Number of HSP's better than 10.0 without gapping: 1122
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1124
length of query: 215
length of database: 12,639,632
effective HSP length: 93
effective length of query: 122
effective length of database: 6,717,764
effective search space: 819567208
effective search space used: 819567208
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 57 (26.6 bits)
Medicago: description of AC144513.2


