

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144503.9 + phase: 0 /pseudo
(346 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
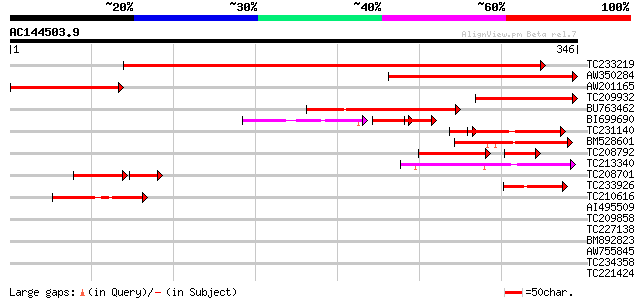
Sequences producing significant alignments: (bits) Value
TC233219 weakly similar to UP|Q8LKI9 (Q8LKI9) NBS/LRR resistance... 410 e-115
AW350284 similar to GP|22946224|gb| CG31868-PA {Drosophila melan... 206 2e-53
AW201165 117 8e-27
TC209932 103 1e-22
BU763462 weakly similar to GP|9757923|dbj| laccase (diphenol oxi... 77 1e-14
BI699690 48 2e-12
TC231140 similar to GB|AAO24550.1|27808540|BT003118 At3g59650 {A... 53 1e-08
BM528601 56 2e-08
TC208792 homologue to UP|Q9S834 (Q9S834) ATP-dependent Clp prote... 49 6e-08
TC213340 54 8e-08
TC208701 weakly similar to UP|Q9SAE5 (Q9SAE5) F3F19.14 protein, ... 44 4e-07
TC233926 45 6e-05
TC210616 similar to UP|Q94JV1 (Q94JV1) At1g69340/F10D13.28, part... 43 2e-04
AI495509 38 0.006
TC209858 38 0.006
TC227138 similar to GB|AAO44062.1|28466907|BT004796 At1g51160 {A... 32 0.017
BM892823 similar to GP|22946353|gb CG6214-PA {Drosophila melanog... 36 0.029
AW755845 32 0.32
TC234358 30 2.1
TC221424 30 2.1
>TC233219 weakly similar to UP|Q8LKI9 (Q8LKI9) NBS/LRR resistance
protein-like protein (Fragment), partial (74%)
Length = 781
Score = 410 bits (1055), Expect = e-115
Identities = 199/259 (76%), Positives = 228/259 (87%), Gaps = 1/259 (0%)
Frame = +3
Query: 70 LWKALEKKGVRIAYIRAIKDMYEGASTSVRTQDGTTEDFPITIGVHQGSTLSPYLFTLVL 129
LWKALEKKGVR+AYIRAI+DMY+ STSVRTQ G ++DFPITIG+HQGSTLSPYLFTL+L
Sbjct: 3 LWKALEKKGVRVAYIRAIQDMYDRVSTSVRTQGGESDDFPITIGLHQGSTLSPYLFTLIL 182
Query: 130 DVLTEHIQELAPRCMLFAD-VVLVGESREEVNGRLETWRQALEAYGFRLSRSTTEYMECN 188
DVLTE IQE+ RCMLFAD +VL+GESRE++N RLETWR+ALE +GFRLSRS +EYMEC
Sbjct: 183 DVLTEQIQEIVSRCMLFADDIVLLGESREKLNERLETWRRALETHGFRLSRSKSEYMECK 362
Query: 189 FSGRRSRSTLEVKVGDHIIPQVTRFKYLGSFV*NDGEIEADVSHRIQAEWLKWRRASGVL 248
F+ RR S EVK+GDHIIPQVTRFKYLGS + +DGEIE DV+HRIQA W+KWR+ASGVL
Sbjct: 363 FNKRRRVSNSEVKIGDHIIPQVTRFKYLGSVIQDDGEIEGDVNHRIQA*WMKWRKASGVL 542
Query: 249 CDKKVPLKLKGKFYRTAVRPALLYGTECWAVKSQHENQVSVAEMRMLRWMSRKTRHDRIK 308
CD KVP+KLKGKFYRTAVRPA+LYGTECWAVKSQHEN+V VA RMLRWM KTR D+I+
Sbjct: 543 CDAKVPIKLKGKFYRTAVRPAILYGTECWAVKSQHENKVGVAXXRMLRWMCGKTRQDKIR 722
Query: 309 NDTIRERVGVAPIVEKLVE 327
N+ ERVGVAPIVEK+VE
Sbjct: 723 NEAXXERVGVAPIVEKMVE 779
>AW350284 similar to GP|22946224|gb| CG31868-PA {Drosophila melanogaster},
partial (1%)
Length = 767
Score = 206 bits (523), Expect = 2e-53
Identities = 96/115 (83%), Positives = 106/115 (91%)
Frame = -1
Query: 232 HRIQAEWLKWRRASGVLCDKKVPLKLKGKFYRTAVRPALLYGTECWAVKSQHENQVSVAE 291
HRIQA W+KWR+ SGVLCD KVP+KLKGKFYRTAVRPA+LYGTECWAVKSQHEN+V VAE
Sbjct: 590 HRIQAGWMKWRKTSGVLCDAKVPIKLKGKFYRTAVRPAILYGTECWAVKSQHENKVGVAE 411
Query: 292 MRMLRWMSRKTRHDRIKNDTIRERVGVAPIVEKLVENRLR*FGHVERRPVDAVVR 346
MRMLRWM KTR D+I+N+ IRERVGVAPIVEK+VENRLR FGHVERRPVD+VVR
Sbjct: 410 MRMLRWMCGKTRQDKIRNEAIRERVGVAPIVEKMVENRLRWFGHVERRPVDSVVR 246
>AW201165
Length = 413
Score = 117 bits (293), Expect = 8e-27
Identities = 56/69 (81%), Positives = 63/69 (91%)
Frame = -1
Query: 1 KLWERVIERRLRKEARVTDNQFGFMPGRLTMEAIYLLRRVIERYRTDKKDLHLVFIDLEK 60
KLWERVIERRLRKE +VT+NQFGFMPGR TMEAIYLLRRV+E+YR ++DLHL+FIDLEK
Sbjct: 209 KLWERVIERRLRKETQVTENQFGFMPGRSTMEAIYLLRRVMEQYRMAQQDLHLIFIDLEK 30
Query: 61 TYDRVPREI 69
DRVPREI
Sbjct: 29 ASDRVPREI 3
>TC209932
Length = 829
Score = 103 bits (257), Expect = 1e-22
Identities = 50/62 (80%), Positives = 56/62 (89%)
Frame = +2
Query: 285 NQVSVAEMRMLRWMSRKTRHDRIKNDTIRERVGVAPIVEKLVENRLR*FGHVERRPVDAV 344
N+V VAEMRMLRWM KTR D+I+N+ IRERVGVAPIVEK+VENRLR FGHVERRPVD+V
Sbjct: 224 NKVGVAEMRMLRWMCGKTRQDKIRNEAIRERVGVAPIVEKMVENRLRWFGHVERRPVDSV 403
Query: 345 VR 346
VR
Sbjct: 404 VR 409
>BU763462 weakly similar to GP|9757923|dbj| laccase (diphenol oxidase)
{Arabidopsis thaliana}, partial (4%)
Length = 421
Score = 77.0 bits (188), Expect = 1e-14
Identities = 41/94 (43%), Positives = 57/94 (60%)
Frame = +1
Query: 182 TEYMECNFSGRRSRSTLEVKVGDHIIPQVTRFKYLGSFV*NDGEIEADVSHRIQAEWLKW 241
T+YM NFS + + LEVK+GD IP V++FKY + N I D +H++Q LK
Sbjct: 4 TKYMHSNFSKK*EENELEVKMGD-AIP*VSKFKYFEPILQNSW*INEDGTHKMQLGCLKE 180
Query: 242 RRASGVLCDKKVPLKLKGKFYRTAVRPALLYGTE 275
+AS + CD KVP +K +FY T + P +LYG E
Sbjct: 181 AKASRINCDHKVPTNIKAQFYCTVIHPNILYGNE 282
>BI699690
Length = 407
Score = 47.8 bits (112), Expect(3) = 2e-12
Identities = 30/80 (37%), Positives = 45/80 (55%), Gaps = 4/80 (5%)
Frame = -1
Query: 143 CMLFADVVLVGESREEVNGRLETWRQALEAYGFRLSRSTTEYMECNFSGRRSRSTLEVKV 202
CML AD + E + + W+Q+ F LSRS T+YM C+FS + + + +
Sbjct: 347 CMLSADDIFFIEESHNMRFGHKLWKQS-----FHLSRSKTKYMYCSFS--KKQEEYK*ED 189
Query: 203 GDHIIPQVT----RFKYLGS 218
G+ +IPQV+ +FKYLGS
Sbjct: 188 GEDVIPQVSKFKFKFKYLGS 129
Score = 32.7 bits (73), Expect(3) = 2e-12
Identities = 13/19 (68%), Positives = 16/19 (83%)
Frame = -3
Query: 242 RRASGVLCDKKVPLKLKGK 260
+R GV+CD KVP+KLKGK
Sbjct: 57 KRQLGVICDHKVPIKLKGK 1
Score = 28.5 bits (62), Expect(3) = 2e-12
Identities = 13/25 (52%), Positives = 18/25 (72%)
Frame = -2
Query: 222 NDGEIEADVSHRIQAEWLKWRRASG 246
N G++ DVSH IQA+ LK ++A G
Sbjct: 118 NYGKMNEDVSHEIQAKRLKCKKAIG 44
>TC231140 similar to GB|AAO24550.1|27808540|BT003118 At3g59650 {Arabidopsis
thaliana;} , partial (93%)
Length = 1123
Score = 52.8 bits (125), Expect(2) = 1e-08
Identities = 28/61 (45%), Positives = 46/61 (74%), Gaps = 1/61 (1%)
Frame = +3
Query: 280 KSQHENQVSVAEMRMLRWMSR-KTRHDRIKNDTIRERVGVAPIVEKLVENRLR*FGHVER 338
K+ + +++VA+MR+L+WMS KTR D ++ +E+VGV PIV+K++E+ LR*F H+ +
Sbjct: 357 KANKKIKLNVAKMRLLKWMSTYKTRWD---DECYKEKVGVTPIVKKMLESLLR*FEHI*K 527
Query: 339 R 339
R
Sbjct: 528 R 530
Score = 24.3 bits (51), Expect(2) = 1e-08
Identities = 11/17 (64%), Positives = 12/17 (69%)
Frame = +2
Query: 269 ALLYGTECWAVKSQHEN 285
A+LYG E W V SQ EN
Sbjct: 323 AMLYGIEYWPV*SQ*EN 373
>BM528601
Length = 426
Score = 56.2 bits (134), Expect = 2e-08
Identities = 37/77 (48%), Positives = 48/77 (62%), Gaps = 5/77 (6%)
Frame = +3
Query: 272 YGTECWAVKSQHENQVSVA---EMRML--RWMSRKTRHDRIKNDTIRERVGVAPIVEKLV 326
Y T+C AVK E++ EMRML MS R +++ N+ IRE+VGVA I EK+V
Sbjct: 156 YCTKCSAVKCHQEHKTKCC*ELEMRML**LMMSIHARIEKM-NECIREKVGVALIQEKMV 332
Query: 327 ENRLR*FGHVERRPVDA 343
E LR F HV RRP++A
Sbjct: 333 EMHLRWFDHVPRRPIEA 383
>TC208792 homologue to UP|Q9S834 (Q9S834) ATP-dependent Clp protease subunit
ClpP (NClpP1) (ATP-dependent Clp protease proteolytic
subunit ClpP5) , partial (14%)
Length = 739
Score = 48.9 bits (115), Expect(2) = 6e-08
Identities = 23/44 (52%), Positives = 31/44 (70%)
Frame = -2
Query: 250 DKKVPLKLKGKFYRTAVRPALLYGTECWAVKSQHENQVSVAEMR 293
+KKVPLKLKGK T++R ++YGT+ W VK Q + +VAE R
Sbjct: 597 NKKVPLKLKGKLQDTSIRSMMVYGTKEWVVKGQQ*RKHNVAETR 466
Score = 25.4 bits (54), Expect(2) = 6e-08
Identities = 11/22 (50%), Positives = 16/22 (72%)
Frame = -3
Query: 303 RHDRIKNDTIRERVGVAPIVEK 324
R D+I+N+ + +VGV IVEK
Sbjct: 473 RQDKIRNECNKGKVGVTLIVEK 408
>TC213340
Length = 501
Score = 54.3 bits (129), Expect = 8e-08
Identities = 35/112 (31%), Positives = 58/112 (51%), Gaps = 5/112 (4%)
Frame = -1
Query: 239 LKWRRASG---VLCDKKVPLKLKGKFYRTAVRPALLYGTECWAVKSQHENQVS--VAEMR 293
L W+++ G V+ KV LK +FY T + P +LY +ECW K HE +VS +
Sbjct: 429 LMWKKSIGGLFVIEICKVLTNLKREFYCTVIEPTILYSSECWGSKG*HEKKVSNRNENTK 250
Query: 294 MLRWMSRKTRHDRIKNDTIRERVGVAPIVEKLVENRLR*FGHVERRPVDAVV 345
+ W +K++ + + + V I E + N + F HV++RP++A V
Sbjct: 249 IDEWPHKKSQDTK---*LYKGNISVTTIEEWTIGN*IICFRHVQKRPLEAPV 103
>TC208701 weakly similar to UP|Q9SAE5 (Q9SAE5) F3F19.14 protein, partial
(19%)
Length = 951
Score = 43.9 bits (102), Expect(2) = 4e-07
Identities = 19/33 (57%), Positives = 24/33 (72%)
Frame = -1
Query: 40 VIERYRTDKKDLHLVFIDLEKTYDRVPREILWK 72
+ +Y DLH+VFIDLEK YD+V +EILWK
Sbjct: 168 ITTKYGG*SSDLHIVFIDLEKFYDQVAKEILWK 70
Score = 27.7 bits (60), Expect(2) = 4e-07
Identities = 12/20 (60%), Positives = 17/20 (85%)
Frame = -3
Query: 74 LEKKGVRIAYIRAIKDMYEG 93
+EKK V++A I AI+DMY+G
Sbjct: 76 MEKKDVQVA*IWAIQDMYDG 17
>TC233926
Length = 570
Score = 44.7 bits (104), Expect = 6e-05
Identities = 20/39 (51%), Positives = 30/39 (76%)
Frame = +3
Query: 302 TRHDRIKNDTIRERVGVAPIVEKLVENRLR*FGHVERRP 340
TR DR++ND IR+ + V PI +K+ +N LR FGH++R+P
Sbjct: 48 TRKDRVQND*IRD-ISVVPIKDKMTQNGLRWFGHMQRKP 161
>TC210616 similar to UP|Q94JV1 (Q94JV1) At1g69340/F10D13.28, partial (6%)
Length = 827
Score = 43.1 bits (100), Expect = 2e-04
Identities = 26/59 (44%), Positives = 40/59 (67%), Gaps = 1/59 (1%)
Frame = +2
Query: 27 GRLTME-AIYLLRRVIERYRTDKKDLHLVFIDLEKTYDRVPREILWKALEKKGVRIAYI 84
GR T + IYLL R++E+Y +K+LH ++L++ D +P+E+ KAL+KKG I YI
Sbjct: 464 GRSTAQWCIYLL*RLMEKYHRKQKELH---VELQED-D*MPKEVF*KALQKKGFCITYI 628
>AI495509
Length = 394
Score = 38.1 bits (87), Expect = 0.006
Identities = 13/27 (48%), Positives = 23/27 (85%)
Frame = +2
Query: 266 VRPALLYGTECWAVKSQHENQVSVAEM 292
++ +L+GTECW VKSQ +N+++VA++
Sbjct: 314 IQXVMLFGTECWTVKSQQDNKLNVADI 394
>TC209858
Length = 1340
Score = 38.1 bits (87), Expect = 0.006
Identities = 19/34 (55%), Positives = 24/34 (69%)
Frame = -3
Query: 248 LCDKKVPLKLKGKFYRTAVRPALLYGTECWAVKS 281
LC + VPLK KGKFY +R A+ GT+C A+KS
Sbjct: 285 LC-QNVPLKTKGKFYHPNIRHAM*CGTKCRAIKS 187
Score = 33.9 bits (76), Expect = 0.11
Identities = 34/118 (28%), Positives = 48/118 (39%), Gaps = 2/118 (1%)
Frame = -2
Query: 229 DVSHRIQAEWLKWRRASGVLCDKKVPLKLKGKFYRTAVRPALLYGTECWAVKSQHENQVS 288
DV+HRI A +KW+ C K+ Y+T W H+
Sbjct: 334 DVNHRIHAGRMKWKILFMPECTT*NQRKILPSKYKTC--------NVVWN*MPCHKKLTK 179
Query: 289 VAEMRMLR--WMSRKTRHDRIKNDTIRERVGVAPIVEKLVENRLR*FGHVERRPVDAV 344
R + T DRI+N I E+VG+AP+VEK++ R F R VD +
Sbjct: 178 RINSM**R*DYYIGCTGQDRIRNVRIGEKVGIAPMVEKML*YYYRWFLENPVRRVDQI 5
>TC227138 similar to GB|AAO44062.1|28466907|BT004796 At1g51160 {Arabidopsis
thaliana;} , partial (97%)
Length = 1259
Score = 32.0 bits (71), Expect(2) = 0.017
Identities = 18/48 (37%), Positives = 29/48 (59%)
Frame = -3
Query: 205 HIIPQVTRFKYLGSFV*NDGEIEADVSHRIQAEWLKWRRASGVLCDKK 252
H+I +VTRF +L + EI+ +++ RIQ +W + V+CDKK
Sbjct: 1209 HVI-RVTRFTHLQFIIYIGREIQGNINCRIQIKW----SITSVICDKK 1081
Score = 23.5 bits (49), Expect(2) = 0.017
Identities = 9/16 (56%), Positives = 11/16 (68%)
Frame = -2
Query: 192 RRSRSTLEVKVGDHII 207
RR EVK+GDHI+
Sbjct: 1258 RRINLXFEVKIGDHIL 1211
>BM892823 similar to GP|22946353|gb CG6214-PA {Drosophila melanogaster},
partial (0%)
Length = 421
Score = 35.8 bits (81), Expect = 0.029
Identities = 15/28 (53%), Positives = 22/28 (78%)
Frame = +1
Query: 316 VGVAPIVEKLVENRLR*FGHVERRPVDA 343
+G+ I EK +E +LR*FGHV+R P++A
Sbjct: 55 IGMTVIEEKKIETQLR*FGHVQRSPLEA 138
>AW755845
Length = 152
Score = 32.3 bits (72), Expect = 0.32
Identities = 13/30 (43%), Positives = 20/30 (66%)
Frame = -2
Query: 210 VTRFKYLGSFV*NDGEIEADVSHRIQAEWL 239
VT F+YLGS + GE + DV+ +I + W+
Sbjct: 94 VT*FRYLGSIIQRKGETKGDVNQKIHSGWM 5
>TC234358
Length = 432
Score = 29.6 bits (65), Expect = 2.1
Identities = 22/66 (33%), Positives = 35/66 (52%)
Frame = -2
Query: 148 DVVLVGESREEVNGRLETWRQALEAYGFRLSRSTTEYMECNFSGRRSRSTLEVKVGDHII 207
DVVL+ +SRE V + + W + + L RS T Y+ + L+VK+ +I+
Sbjct: 308 DVVLIEDSREVVYFKHKLWIYNFDTRSYCLRRSKTVYI---LQFQ*EEYKLKVKI-RNIL 141
Query: 208 PQVTRF 213
QV+RF
Sbjct: 140 SQVSRF 123
>TC221424
Length = 918
Score = 29.6 bits (65), Expect = 2.1
Identities = 14/24 (58%), Positives = 18/24 (74%)
Frame = +2
Query: 236 AEWLKWRRASGVLCDKKVPLKLKG 259
AE L WR+AS V+C+ KV +LKG
Sbjct: 2 AE*L*WRKASRVICNPKVLTELKG 73
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.326 0.140 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,898,169
Number of Sequences: 63676
Number of extensions: 172767
Number of successful extensions: 922
Number of sequences better than 10.0: 45
Number of HSP's better than 10.0 without gapping: 913
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 916
length of query: 346
length of database: 12,639,632
effective HSP length: 98
effective length of query: 248
effective length of database: 6,399,384
effective search space: 1587047232
effective search space used: 1587047232
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 59 (27.3 bits)
Medicago: description of AC144503.9


