

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144484.11 - phase: 0
(796 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
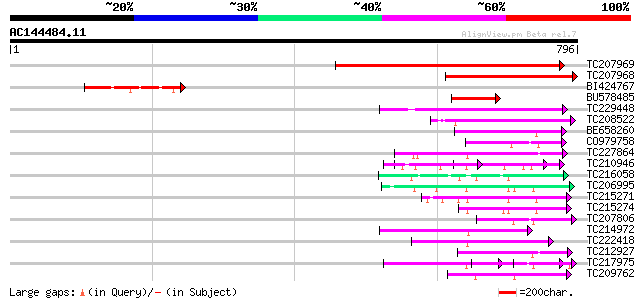
Score E
Sequences producing significant alignments: (bits) Value
TC207969 similar to UP|Q9SJ70 (Q9SJ70) Probable calcium-binding ... 549 e-156
TC207968 similar to UP|Q9SJ70 (Q9SJ70) Probable calcium-binding ... 327 1e-89
BI424767 176 4e-44
BU578485 136 4e-32
TC229448 similar to UP|Q6YVE7 (Q6YVE7) Mitochondrial aspartate-g... 127 2e-29
TC208522 similar to UP|Q6YVE7 (Q6YVE7) Mitochondrial aspartate-g... 107 2e-23
BE658260 similar to GP|9757958|dbj mitochondrial carrier protein... 87 3e-17
CO979758 76 5e-14
TC227864 uncoupling protein 1a [Glycine max] 72 1e-12
TC210946 similar to UP|Q9FI43 (Q9FI43) Calcium-binding transport... 70 3e-12
TC216058 similar to UP|MCAT_ARATH (Q93XM7) Mitochondrial carniti... 70 5e-12
TC206995 similar to UP|Q9SSP4 (Q9SSP4) F3N23.2 protein (At1g7282... 69 1e-11
TC215271 similar to PIR|T01729|T01729 mitochondrial solute carri... 67 3e-11
TC215274 similar to PIR|T01729|T01729 mitochondrial solute carri... 66 5e-11
TC207806 similar to GB|AAP42736.1|30984546|BT008723 At1g07030 {A... 66 7e-11
TC214972 similar to UP|Q43649 (Q43649) Oxoglutarate malate trans... 65 1e-10
TC222418 UP|Q8W1A3 (Q8W1A3) Uncoupling protein 1b (Fragment), pa... 64 2e-10
TC212927 similar to UP|Q9ZNY4 (Q9ZNY4) Mitochondrial energy tran... 64 3e-10
TC217975 similar to UP|Q9LY28 (Q9LY28) Peroxisomal Ca-dependent ... 63 4e-10
TC209762 62 1e-09
>TC207969 similar to UP|Q9SJ70 (Q9SJ70) Probable calcium-binding
mitochondrial carrier At2g35800, partial (36%)
Length = 968
Score = 549 bits (1414), Expect = e-156
Identities = 276/322 (85%), Positives = 299/322 (92%)
Frame = +3
Query: 458 EDNAAAMMRFLNADTEESISYGHFRNFMLLLPSDRLQEDPRSIWFEAATVVAVPPSVEIP 517
EDNA AMMRFL ADTEESISYGHFRNFMLLLPSDRLQEDPRSIWFEAATVVAVPP+VEIP
Sbjct: 3 EDNAVAMMRFLKADTEESISYGHFRNFMLLLPSDRLQEDPRSIWFEAATVVAVPPAVEIP 182
Query: 518 AGSVLRSALAGGLSCALSCALLHPVDSIKTRVQASSMSFPEIIAKLPEIGTRGLYRGSIP 577
AGSVLRSALAGGLSCALSCALLHPVD+IKTRVQAS+MSFPEII+KLPEIG RGLYRGSIP
Sbjct: 183 AGSVLRSALAGGLSCALSCALLHPVDTIKTRVQASTMSFPEIISKLPEIGRRGLYRGSIP 362
Query: 578 AILGQFSSHGLRTGIFEASKLVLVNVAPNLPELQVQSIASFCSTFLGTAVRIPCEVLKQR 637
AILGQFSSHGLRTGIFEASKLVL+N+AP LPELQVQS+ASFCSTFLGTAVRIPC+VLKQ+
Sbjct: 363 AILGQFSSHGLRTGIFEASKLVLINIAPTLPELQVQSVASFCSTFLGTAVRIPCQVLKQK 542
Query: 638 LQAGLFNNVGEALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKGVQKLLGR 697
LQAGL +NVG+A V TW+QDGL+GFFRGTGATLC EVPFYV GMGLYAESKK ++LL R
Sbjct: 543 LQAGLCDNVGKAFVATWEQDGLRGFFRGTGATLCHEVPFYVDGMGLYAESKKVAERLLER 722
Query: 698 ELEAWETIAVGALSGGLAAVVTTPFDVMKTRMMTAQGRSVSMSIVAFSILRHEGPLGLFK 757
EL ETIA GALSGGLAAVVTTPFDVMKTRMMTA GRSVSM+++AF IL+ +GPLGLFK
Sbjct: 723 ELSPLETIADGALSGGLAAVVTTPFDVMKTRMMTAHGRSVSMTLIAFFILKLDGPLGLFK 902
Query: 758 GAVPRFFWIAPLGAMNFAGYEL 779
GA PRFFWIAP+ A+ FAG +L
Sbjct: 903 GA*PRFFWIAPVCAIIFAGLDL 968
>TC207968 similar to UP|Q9SJ70 (Q9SJ70) Probable calcium-binding
mitochondrial carrier At2g35800, partial (21%)
Length = 884
Score = 327 bits (839), Expect = 1e-89
Identities = 161/185 (87%), Positives = 174/185 (94%)
Frame = +3
Query: 612 VQSIASFCSTFLGTAVRIPCEVLKQRLQAGLFNNVGEALVGTWQQDGLKGFFRGTGATLC 671
VQS+ASFCSTFLGTAVRIPCEVLKQRLQAGLF+NVGEA V TW+QDGL+GFFRGTGATLC
Sbjct: 120 VQSVASFCSTFLGTAVRIPCEVLKQRLQAGLFDNVGEAFVATWEQDGLRGFFRGTGATLC 299
Query: 672 REVPFYVAGMGLYAESKKGVQKLLGRELEAWETIAVGALSGGLAAVVTTPFDVMKTRMMT 731
REVPFYVAGMGLYAESKK ++LL REL ETIAVGALSGGLAAVVTTPFDVMKTRMMT
Sbjct: 300 REVPFYVAGMGLYAESKKVAERLLERELGPLETIAVGALSGGLAAVVTTPFDVMKTRMMT 479
Query: 732 AQGRSVSMSIVAFSILRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYELARKAMNKNDEAK 791
AQGRSVSM+++AFSIL+HEGPLGLFKGAVPRFFWIAPLGAMNFAGYELA+KAMNKN+E K
Sbjct: 480 AQGRSVSMTLIAFSILKHEGPLGLFKGAVPRFFWIAPLGAMNFAGYELAKKAMNKNEEGK 659
Query: 792 TGNLE 796
G+ E
Sbjct: 660 AGSSE 674
>BI424767
Length = 428
Score = 176 bits (446), Expect = 4e-44
Identities = 94/149 (63%), Positives = 115/149 (77%), Gaps = 7/149 (4%)
Frame = +2
Query: 105 SNCLKFSVTWSLLVSGFIQSLPIPFKSVKKRGQKVCDEDSHKEKCSCMKPSLSPCEMKHN 164
+NCL+F+VTWSLLV+GF+QSLP+PFKS KK+ QKVCDED + CSC KP++S CE+K N
Sbjct: 2 TNCLQFAVTWSLLVNGFLQSLPLPFKSGKKKCQKVCDED---KLCSCTKPTVSSCEVKQN 172
Query: 165 ESK----GRTIKEKVVKRKDGKEHVSLECVIGFIFDQLSHTLQSLDQGINGLQEKNDELE 220
ESK GR ++EK V+RKDGK +VSLEC+IGFIFDQLS TLQSLD G++ E ND+L+
Sbjct: 173 ESKGGQFGRAVREKGVRRKDGK-NVSLECLIGFIFDQLSQTLQSLDYGVH---ENNDDLD 340
Query: 221 CGKASLDS---APFGHVNAFTSFLEGHKV 246
GK SL + FGHVNA FLE HKV
Sbjct: 341 NGKTSLPQPSFSHFGHVNALAGFLEEHKV 427
>BU578485
Length = 428
Score = 136 bits (342), Expect = 4e-32
Identities = 64/69 (92%), Positives = 67/69 (96%)
Frame = +1
Query: 621 TFLGTAVRIPCEVLKQRLQAGLFNNVGEALVGTWQQDGLKGFFRGTGATLCREVPFYVAG 680
TFLGTAVRIPCEVLKQRLQAGLF+NVGEA V TW+QDGL+GFFRGTGATLCREVPFYVAG
Sbjct: 1 TFLGTAVRIPCEVLKQRLQAGLFDNVGEAFVATWEQDGLRGFFRGTGATLCREVPFYVAG 180
Query: 681 MGLYAESKK 689
MGLYAESKK
Sbjct: 181 MGLYAESKK 207
>TC229448 similar to UP|Q6YVE7 (Q6YVE7) Mitochondrial aspartate-glutamate
carrier protein-like, partial (96%)
Length = 1000
Score = 127 bits (319), Expect = 2e-29
Identities = 79/265 (29%), Positives = 127/265 (47%), Gaps = 2/265 (0%)
Frame = +1
Query: 520 SVLRSALAGGLSCALSCALLHPVDSIKTRVQASSMSFPEIIAKLPEIGTRGLYRGSIPAI 579
++ +AGG + + L+P+D+IKTR+QA+ I+ +GLY G +
Sbjct: 136 TLFEGVIAGGTAGVVVETALYPIDTIKTRLQAARGGEKLIL--------KGLYSGLAGNL 291
Query: 580 LGQFSSHGLRTGIFEASKLVLVNVAPNLPELQVQSIASFCSTFLGTAVRIPCEVLKQRLQ 639
+G + L G++E K L+ + P A + +R+P EV+KQR+Q
Sbjct: 292 VGVLPASALFVGVYEPIKQKLLRIFPEHLSAFTHLTAGAIGGIAASLIRVPTEVIKQRMQ 471
Query: 640 AGLFNNVGEALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKGVQKLLGREL 699
G F + A+ ++G KGF+ G G+ L R++PF +Y + + G R L
Sbjct: 472 TGQFASASGAVRFIASKEGFKGFYAGYGSFLLRDLPFDAIQFCIYEQIRIGYMLAAQRNL 651
Query: 700 EAWETIAVGALSGGLAAVVTTPFDVMKTRMMTAQGRSVSMSIV--AFSILRHEGPLGLFK 757
E +GA +G L +TTP DV+KTR+M + IV +I++ EGP K
Sbjct: 652 NDPENAIIGAFAGALTGAITTPLDVIKTRLMVQGSANQYKGIVDCVQTIIKEEGPRAFLK 831
Query: 758 GAVPRFFWIAPLGAMNFAGYELARK 782
G PR WI G++ F E ++
Sbjct: 832 GIGPRVLWIGIGGSIFFGVLESTKR 906
>TC208522 similar to UP|Q6YVE7 (Q6YVE7) Mitochondrial aspartate-glutamate
carrier protein-like, partial (72%)
Length = 1068
Score = 107 bits (268), Expect = 2e-23
Identities = 70/212 (33%), Positives = 115/212 (54%), Gaps = 8/212 (3%)
Frame = +2
Query: 591 GIFEASKLVLVNVAPNLPELQVQSIASFCSTFLG----TAVRIPCEVLKQRLQAGLFNNV 646
G++E +K L+ +LPE + ++A F + +G + VR+P EV+KQR+Q G F +
Sbjct: 23 GVYEPTKQQLLK---SLPE-NLSAVAHFAAGAIGGIASSVVRVPTEVVKQRMQIGQFKSA 190
Query: 647 GEALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKGVQKLLGRELEAWETIA 706
+A+ +G KG F G G+ L R++PF + +Y + + G + R+ E
Sbjct: 191 PDAVRLIVANEGFKGLFAGYGSFLLRDLPFDAIELCIYEQLRIGYKLAAKRDPNDPENAM 370
Query: 707 VGALSGGLAAVVTTPFDVMKTRMMT--AQGRSVSMSIVAFSILRHEGPLGLFKGAVPRFF 764
+GA++G + VTTP DV+KTR+M +Q +S +I++ EG LFKG PR
Sbjct: 371 LGAVAGAVTGAVTTPLDVVKTRLMVQGSQNHYKGISDCVRTIVKEEGSHALFKGIGPRVL 550
Query: 765 WIAPLGAMNFAGYELARK--AMNKNDEAKTGN 794
WI G++ F E +K A ++ +A+T N
Sbjct: 551 WIGIGGSIFFCVLEKTKKILAQKRHSKAETQN 646
>BE658260 similar to GP|9757958|dbj mitochondrial carrier protein-like
{Arabidopsis thaliana}, partial (46%)
Length = 775
Score = 87.0 bits (214), Expect = 3e-17
Identities = 57/168 (33%), Positives = 86/168 (50%), Gaps = 11/168 (6%)
Frame = -3
Query: 625 TAVRIPCEVLKQRLQAGLFNNVGEALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLY 684
+A+ +P E++ QR+QAG + Q DG+ G + G ATL R +P V +
Sbjct: 737 SAIMVPKELITQRMQAGAKXRSXQVFAEIIQXDGVMGLYAGYSATLLRNLPAGVLSYSSF 558
Query: 685 AESKKGV-QKLLGRELEAWETIAVGALSGGLAAVVTTPFDVMKTRMMT-AQGRSVS---- 738
K V QK +E +++ GAL+G ++A +TTP DV+KTR+MT +G VS
Sbjct: 557 EYLKAAVLQKTKQSYMEPVQSVLCGALAGAISASLTTPLDVVKTRLMTQVRGEGVSKVAA 378
Query: 739 -----MSIVAFSILRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYELAR 781
+S IL+ EG +GL +G PR A A+ + +E AR
Sbjct: 377 VMYDGVSATVKQILKEEGWVGLTRGMGPRVLHSACFSALGYFAFETAR 234
Score = 37.4 bits (85), Expect = 0.026
Identities = 36/131 (27%), Positives = 60/131 (45%), Gaps = 14/131 (10%)
Frame = -3
Query: 501 WFEAATVVAVPPSVEIPAGSVLRSALAGGLSCALSCALLHPVDSIKTRV----------Q 550
+ +AA + S P SVL ALAG A+S +L P+D +KTR+ +
Sbjct: 554 YLKAAVLQKTKQSYMEPVQSVLCGALAG----AISASLTTPLDVVKTRLMTQVRGEGVSK 387
Query: 551 ASSMSFPEIIAKLPEI----GTRGLYRGSIPAILGQFSSHGLRTGIFEASKLVLVNVAPN 606
+++ + + A + +I G GL RG P +L L FE ++L ++
Sbjct: 386 VAAVMYDGVSATVKQILKEEGWVGLTRGMGPRVLHSACFSALGYFAFETARLSILREYLR 207
Query: 607 LPELQVQSIAS 617
EL+ S++S
Sbjct: 206 SKELREVSVSS 174
>CO979758
Length = 751
Score = 76.3 bits (186), Expect = 5e-14
Identities = 50/152 (32%), Positives = 75/152 (48%), Gaps = 11/152 (7%)
Frame = -1
Query: 641 GLFNNVGEALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKGVQKLLGRELE 700
G + + A W+ GLKG + G +TL R+VPF + Y K + R +
Sbjct: 706 GYYTGMLHAGCSIWKAQGLKGLYAGYLSTLARDVPFAGLMVVFYEALKDAKDYVEQRWIS 527
Query: 701 A--W------ETIAVGALSGGLAAVVTTPFDVMKTRMMTAQGRSVSMS---IVAFSILRH 749
+ W E + +G L+GGL+A +TTP DV+KTR+ QG ++ + +I
Sbjct: 526 SPNWHVNNSVEGLVLGGLAGGLSAYLTTPLDVVKTRLQ-VQGSTLRYNGWLDAIHNIWAT 350
Query: 750 EGPLGLFKGAVPRFFWIAPLGAMNFAGYELAR 781
EG G+F+G+VPR W P A+ F E R
Sbjct: 349 EGMKGMFRGSVPRITWYIPASALTFMAVEFLR 254
Score = 36.6 bits (83), Expect = 0.045
Identities = 22/58 (37%), Positives = 34/58 (57%), Gaps = 6/58 (10%)
Frame = -1
Query: 528 GGLSCALSCALLHPVDSIKTR--VQASSMSFPEIIAKLPEI----GTRGLYRGSIPAI 579
GGL+ LS L P+D +KTR VQ S++ + + + I G +G++RGS+P I
Sbjct: 481 GGLAGGLSAYLTTPLDVVKTRLQVQGSTLRYNGWLDAIHNIWATEGMKGMFRGSVPRI 308
Score = 36.2 bits (82), Expect = 0.058
Identities = 34/125 (27%), Positives = 53/125 (42%), Gaps = 16/125 (12%)
Frame = -1
Query: 567 GTRGLYRGSIPAILGQFSSHGLRTGIFEASKLVLVNV------APN------LPELQVQS 614
G +GLY G + + GL +EA K V +PN + L +
Sbjct: 655 GLKGLYAGYLSTLARDVPFAGLMVVFYEALKDAKDYVEQRWISSPNWHVNNSVEGLVLGG 476
Query: 615 IASFCSTFLGTAVRIPCEVLKQRLQAG----LFNNVGEALVGTWQQDGLKGFFRGTGATL 670
+A S +L T P +V+K RLQ +N +A+ W +G+KG FRG+ +
Sbjct: 475 LAGGLSAYLTT----PLDVVKTRLQVQGSTLRYNGWLDAIHNIWATEGMKGMFRGSVPRI 308
Query: 671 CREVP 675
+P
Sbjct: 307 TWYIP 293
>TC227864 uncoupling protein 1a [Glycine max]
Length = 1434
Score = 71.6 bits (174), Expect = 1e-12
Identities = 65/267 (24%), Positives = 115/267 (42%), Gaps = 25/267 (9%)
Frame = +1
Query: 541 PVDSIKTRVQASSMSFPEIIAKLPE----IGTRG----------LYRGSIPAILGQFSSH 586
P+D+ K R+Q + + LP+ +GT G L++G +P + Q
Sbjct: 310 PLDTAKVRLQLQKQAVAGDVVSLPKYKGMLGTVGTIAREEGLSALWKGIVPGLHRQCLYG 489
Query: 587 GLRTGIFEASKLVLVNV--APNLPELQVQSIASFCSTFLGTAVRIPCEVLKQRLQAG--- 641
GLR G++E K V ++P L + +A+F + AV P +++K RLQA
Sbjct: 490 GLRIGLYEPVKTFYVGKDHVGDVP-LSKKILAAFTTGAFAIAVANPTDLVKVRLQAEGKL 666
Query: 642 ------LFNNVGEALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKGVQKLL 695
++ A +Q+G+ + G G + R A + Y + K+ + K+
Sbjct: 667 PPGVPRRYSGSLNAYSTIVRQEGVGALWTGLGPNIARNGIINAAELASYDQVKQTILKIP 846
Query: 696 GRELEAWETIAVGALSGGLAAVVTTPFDVMKTRMMTAQGRSVSMSIVAFSILRHEGPLGL 755
G + G +G A + +P DV+K+RMM ++ L+++GPL
Sbjct: 847 GFTDNVVTHLLAGLGAGFFAVCIGSPVDVVKSRMMGDSSYKNTLDCF-IKTLKNDGPLAF 1023
Query: 756 FKGAVPRFFWIAPLGAMNFAGYELARK 782
+KG +P F + + F E +K
Sbjct: 1024YKGFLPNFGRLGSWNVIMFLTLEQTKK 1104
>TC210946 similar to UP|Q9FI43 (Q9FI43) Calcium-binding transporter-like
protein, partial (42%)
Length = 714
Score = 70.5 bits (171), Expect = 3e-12
Identities = 50/164 (30%), Positives = 76/164 (45%), Gaps = 9/164 (5%)
Frame = +3
Query: 624 GTAVRIPCEVLKQRLQAGLF-NNVGEALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMG 682
GT P + LK LQ +++ A+ W++ GL GFFRG G + + P
Sbjct: 6 GTRATAPLDRLKVVLQIQTTQSHIMPAIKDIWKKGGLLGFFRGNGLNVLKVAPESAIRFY 185
Query: 683 LYAESKKGVQKLLGRELEAWETIAVG-ALSGGLAAVVTT----PFDVMKTRMMT---AQG 734
Y K + + G E +A A+G L+GG+A V P D++KTR+ T G
Sbjct: 186 SYEMLKTFITRAKGEEAKAANIGAMGRLLAGGIAGAVAQTAIYPMDLVKTRLQTHACKSG 365
Query: 735 RSVSMSIVAFSILRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYE 778
R S+ ++ I EGP ++G +P I P ++ A YE
Sbjct: 366 RIPSLGTLSKDIWVQEGPRAFYRGLIPSLLGIIPYAGIDLAAYE 497
Score = 53.5 bits (127), Expect = 4e-07
Identities = 41/152 (26%), Positives = 71/152 (45%), Gaps = 14/152 (9%)
Frame = +3
Query: 526 LAGGLSCALSCALLHPVDSIKTRVQASSMSFPEIIAKLPEIGT-----------RGLYRG 574
LAGG++ A++ ++P+D +KTR+Q + ++P +GT R YRG
Sbjct: 270 LAGGIAGAVAQTAIYPMDLVKTRLQTHACK----SGRIPSLGTLSKDIWVQEGPRAFYRG 437
Query: 575 SIPAILGQFSSHGLRTGIFEASKLVLVN--VAPNLPELQVQSIASFCSTFLGTAVRIPCE 632
IP++LG G+ +E K + + P VQ S LG P +
Sbjct: 438 LIPSLLGIIPYAGIDLAAYETLKDMSKQYILHDGEPGPLVQLGCGTVSGTLGATCVYPLQ 617
Query: 633 VLKQRLQA-GLFNNVGEALVGTWQQDGLKGFF 663
V++ R+QA + + + T + +GL+GF+
Sbjct: 618 VVRTRMQAQRSYKGMADVFRKTLEHEGLRGFY 713
Score = 50.8 bits (120), Expect = 2e-06
Identities = 62/234 (26%), Positives = 96/234 (40%), Gaps = 18/234 (7%)
Frame = +3
Query: 541 PVDSIKTRVQ---ASSMSFPEIIAKLPEIGTRGLYRGSIPAILGQFSSHGLRTGIFEASK 597
P+D +K +Q S P I + G G +RG+ +L +R +E K
Sbjct: 24 PLDRLKVVLQIQTTQSHIMPAIKDIWKKGGLLGFFRGNGLNVLKVAPESAIRFYSYEMLK 203
Query: 598 LVLVNV------APNLPELQVQSIASFCSTFLGTAVRIPCEVLKQRLQ-----AGLFNNV 646
+ A N+ + TA+ P +++K RLQ +G ++
Sbjct: 204 TFITRAKGEEAKAANIGAMGRLLAGGIAGAVAQTAI-YPMDLVKTRLQTHACKSGRIPSL 380
Query: 647 GEALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYA-ESKKGVQK---LLGRELEAW 702
G W Q+G + F+RG +L +P+ AG+ L A E+ K + K L E
Sbjct: 381 GTLSKDIWVQEGPRAFYRGLIPSLLGIIPY--AGIDLAAYETLKDMSKQYILHDGEPGPL 554
Query: 703 ETIAVGALSGGLAAVVTTPFDVMKTRMMTAQGRSVSMSIVAFSILRHEGPLGLF 756
+ G +SG L A P V++TRM AQ M+ V L HEG G +
Sbjct: 555 VQLGCGTVSGTLGATCVYPLQVVRTRMQ-AQRSYKGMADVFRKTLEHEGLRGFY 713
Score = 33.1 bits (74), Expect = 0.49
Identities = 23/79 (29%), Positives = 36/79 (45%), Gaps = 2/79 (2%)
Frame = +3
Query: 719 TTPFDVMKTRMMTAQGRSVSMSIVAFSILRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYE 778
T P D +K + +S M + I + G LG F+G +AP A+ F YE
Sbjct: 18 TAPLDRLKVVLQIQTTQSHIMPAIK-DIWKKGGLLGFFRGNGLNVLKVAPESAIRFYSYE 194
Query: 779 LARKAMN--KNDEAKTGNL 795
+ + + K +EAK N+
Sbjct: 195 MLKTFITRAKGEEAKAANI 251
>TC216058 similar to UP|MCAT_ARATH (Q93XM7) Mitochondrial
carnitine/acylcarnitine carrier-like protein (A BOUT DE
SOUFFLE) (Carnitine/acylcarnitine translocase-like
protein) (CAC-like protein), partial (97%)
Length = 1441
Score = 69.7 bits (169), Expect = 5e-12
Identities = 84/307 (27%), Positives = 117/307 (37%), Gaps = 41/307 (13%)
Frame = +2
Query: 519 GSVLRSALAGGLSCALSCALLHPVDSIKTRVQASSMSFPEIIAK-----------LPEIG 567
G V + AG + A HP D+IK ++Q+ P + K + G
Sbjct: 383 GDVAKDLAAGTVGGAAQLICGHPFDTIKVKLQSQPAPLPGQLPKYSGAFDAVKQTIAAEG 562
Query: 568 TRGLYRGSIPAILGQFSSHGLRTGIFEASKLVLVNVAPNLPELQVQSIASFCSTFLGTAV 627
RGLY+G + A L ++ LV P P Q + C G AV
Sbjct: 563 ARGLYKG-MGAPLATVAAFNAVLFTVRGQMETLVRSNPGAPLTVDQQVV--CGAGAGVAV 733
Query: 628 RI---PCEVLKQRLQAGLFNNVGEALVGTW--------------------QQDGLKGFFR 664
I P E++K RLQA AL G+ + G++G F+
Sbjct: 734 SILACPTELIKCRLQAQ------SALAGSETATVAVKYGGPMDVARHVLKSEGGMRGLFK 895
Query: 665 GTGATLCREVPFYVAGMGLYAESKKGVQKLLGRE----LEAWETIAVGALSGGLAAVVTT 720
G T+ RE+P G+Y K QK G L I G L+G +
Sbjct: 896 GLVPTMGREIPGNAIMFGVYEALK---QKFAGGTDTSGLSRGSLIVAGGLAGASFWFLVY 1066
Query: 721 PFDVMKTRMMTAQGRS--VSMSIVAFSILRH-EGPLGLFKGAVPRFFWIAPLGAMNFAGY 777
P DV+K+ + R+ S S AF +R EG GL+KG P P A F Y
Sbjct: 1067PTDVIKSVIQVDDHRNPKFSGSFDAFRKIRATEGFKGLYKGFGPAMARSVPANAACFLAY 1246
Query: 778 ELARKAM 784
E+ R A+
Sbjct: 1247EMTRSAL 1267
>TC206995 similar to UP|Q9SSP4 (Q9SSP4) F3N23.2 protein (At1g72820/F3N23_2),
partial (89%)
Length = 1357
Score = 68.6 bits (166), Expect = 1e-11
Identities = 76/312 (24%), Positives = 123/312 (39%), Gaps = 41/312 (13%)
Frame = +2
Query: 522 LRSALAGGLSCALSCALLHPVDSIKTRVQASSMSFPEIIAKLPEI---GTRGLYRGSIPA 578
L +AL G+S AL +PV +KTR Q + I I G R LYRG +
Sbjct: 221 LGAALFSGVSAAL-----YPVVVLKTRQQVAQSKVSCINTAFSLIRGEGFRALYRGFGTS 385
Query: 579 ILGQFSSHGLRTGIFEASK------LVLVNVAPNLPELQVQSIASFCSTFLGTAVRIPCE 632
++G + L E +K V +A + A + V P +
Sbjct: 386 LMGTIPARALYMAALEVTKSNVGTATVRFGLAEPTAAAVANAAAGLSAAMAAQLVWTPVD 565
Query: 633 VLKQRLQ-------------AGLFNNVGEALVGTWQQDGLKGFFRGTGATLCREVPFYVA 679
V+ QRL A + N +A DGL+G +RG G ++ P
Sbjct: 566 VVSQRLMVQGVCDSGNSKASALRYINGIDAFRKILSSDGLRGLYRGFGISILTYAPSNAV 745
Query: 680 GMGLYAESKKGVQKLLGREL----------EAWETIAV----GALSGGLAAVVTTPFDVM 725
Y+ +++ V +G L + +AV A++GG++A++T P D +
Sbjct: 746 WWASYSVAQRMVWGGVGYYLCKGNDSALKPDTKTVMAVQGVSAAVAGGMSALITMPLDTI 925
Query: 726 KTRMMTAQG-----RSVSMSIVAFSILRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYELA 780
KTR+ G R + S++R G + ++G PR+ ++ YEL
Sbjct: 926 KTRLQVLDGDENGRRGPTAMQTVRSLVREGGWMACYRGLGPRWASMSMSATTMITTYELL 1105
Query: 781 RKAMNKNDEAKT 792
++ KN E T
Sbjct: 1106KRLSAKNQEVLT 1141
>TC215271 similar to PIR|T01729|T01729 mitochondrial solute carrier protein
homolog - Arabidopsis thaliana {Arabidopsis thaliana;} ,
partial (78%)
Length = 1089
Score = 67.0 bits (162), Expect = 3e-11
Identities = 62/246 (25%), Positives = 104/246 (42%), Gaps = 35/246 (14%)
Frame = +3
Query: 578 AILGQFS----SHGLRTGIFEASKLVLVNVAPN-------LPELQVQSIASFCSTFLGTA 626
AI+ FS S ++TG +++ +VN+A + + ++AS C + +
Sbjct: 180 AIIRSFSLFMASENVKTG--DSAVTTIVNLAEEAKLAREGVVKAPSYALASICKSLVAGG 353
Query: 627 VR--------IPCEVLKQRLQAG-----LFNNVGEALVGTWQQDGLKGFFRGTGATLCRE 673
V P E LK LQ +N + L W+ +G +G F+G G R
Sbjct: 354 VAGGVSRTAVAPLERLKILLQVQNPHNIKYNGTVQGLKYIWRTEGFRGLFKGNGTNCARI 533
Query: 674 VPFYVAGMGLYAESKKGVQKLLGRE-------LEAWETIAVGALSGGLAAVVTTPFDVMK 726
VP Y ++ KG+ L ++ L + GA +G +A T P D+++
Sbjct: 534 VPNSAVKFFSYEQASKGILHLYKQQTGNEDAQLTPLLRLGAGACAGIIAMSATYPMDMVR 713
Query: 727 TRMMTAQGRSV----SMSIVAFSILRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYELARK 782
R+ S M ++LR EGP L+KG +P + P +NFA YE +
Sbjct: 714 GRITVQTEASPYQYRGMFHALSTVLREEGPRALYKGWLPSVIGVIPYVGLNFAVYESLKD 893
Query: 783 AMNKND 788
+ K++
Sbjct: 894 YLIKSN 911
Score = 37.4 bits (85), Expect = 0.026
Identities = 25/95 (26%), Positives = 42/95 (43%), Gaps = 3/95 (3%)
Frame = +3
Query: 703 ETIAVGALSGGLAAVVTTPFDVMKTRMMTAQGRSVSMSIVAFS---ILRHEGPLGLFKGA 759
+++ G ++GG++ P + +K + ++ + I R EG GLFKG
Sbjct: 333 KSLVAGGVAGGVSRTAVAPLERLKILLQVQNPHNIKYNGTVQGLKYIWRTEGFRGLFKGN 512
Query: 760 VPRFFWIAPLGAMNFAGYELARKAMNKNDEAKTGN 794
I P A+ F YE A K + + +TGN
Sbjct: 513 GTNCARIVPNSAVKFFSYEQASKGILHLYKQQTGN 617
>TC215274 similar to PIR|T01729|T01729 mitochondrial solute carrier protein
homolog - Arabidopsis thaliana {Arabidopsis thaliana;} ,
partial (77%)
Length = 1034
Score = 66.2 bits (160), Expect = 5e-11
Identities = 50/175 (28%), Positives = 78/175 (44%), Gaps = 16/175 (9%)
Frame = +2
Query: 630 PCEVLKQRLQAG-----LFNNVGEALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLY 684
P E LK LQ +N + L W+ +G +G F+G G R VP Y
Sbjct: 326 PLERLKILLQVQNPHSIKYNGTIQGLKYIWRTEGFRGLFKGNGTNCARIVPNSAVKFFSY 505
Query: 685 AESKKGV----QKLLGRE---LEAWETIAVGALSGGLAAVVTTPFDVMKTRMMTAQGRSV 737
++ KG+ +K G E L + GA +G +A T P D+++ R+ +S
Sbjct: 506 EQASKGILHLYRKQTGNEDAQLTPLLRLGAGACAGIIAMSATYPMDMVRGRITVQTEKSP 685
Query: 738 ----SMSIVAFSILRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYELARKAMNKND 788
M ++LR EGP L+KG +P + P +NFA YE + + K++
Sbjct: 686 YQYRGMFHALSTVLREEGPRALYKGWLPSVIGVIPYVGLNFAVYESLKDWLVKSN 850
Score = 37.7 bits (86), Expect = 0.020
Identities = 26/95 (27%), Positives = 41/95 (42%), Gaps = 3/95 (3%)
Frame = +2
Query: 703 ETIAVGALSGGLAAVVTTPFDVMKTRMMTAQGRSVSMSIVAFS---ILRHEGPLGLFKGA 759
+++ G ++GG++ P + +K + S+ + I R EG GLFKG
Sbjct: 272 KSLVAGGVAGGVSRTAVAPLERLKILLQVQNPHSIKYNGTIQGLKYIWRTEGFRGLFKGN 451
Query: 760 VPRFFWIAPLGAMNFAGYELARKAMNKNDEAKTGN 794
I P A+ F YE A K + +TGN
Sbjct: 452 GTNCARIVPNSAVKFFSYEQASKGILHLYRKQTGN 556
>TC207806 similar to GB|AAP42736.1|30984546|BT008723 At1g07030 {Arabidopsis
thaliana;} , partial (44%)
Length = 872
Score = 65.9 bits (159), Expect = 7e-11
Identities = 44/151 (29%), Positives = 74/151 (48%), Gaps = 11/151 (7%)
Frame = +1
Query: 656 QDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKGVQKLLGRELEAWETIA---VGALSG 712
++G F+ T+ PF Y +K+G+ ++ ++ + GA +G
Sbjct: 31 EEGFGAFYASYRTTVLMNAPFTAVHFTTYEAAKRGLMEVSPESVDDERLVVHATAGAAAG 210
Query: 713 GLAAVVTTPFDVMKTRMMTAQG-------RSVSMSIVAFSILRHEGPLGLFKGAVPRFFW 765
GLAAVVTTP DV+KT++ QG S S+ V +I++ +G GL +G +PR +
Sbjct: 211 GLAAVVTTPLDVVKTQLQ-CQGVCGCDRFTSGSIGDVIRTIVKKDGYRGLMRGWIPRMLF 387
Query: 766 IAPLGAMNFAGYELARKAMNK-NDEAKTGNL 795
AP A+ ++ YE + N + TG +
Sbjct: 388 HAPAAAICWSTYEAGKSLFQDFNQQKDTGTV 480
Score = 31.6 bits (70), Expect = 1.4
Identities = 24/83 (28%), Positives = 40/83 (47%), Gaps = 10/83 (12%)
Frame = +1
Query: 525 ALAGGLSCALSCALLHPVDSIKTRVQA---------SSMSFPEIIAKL-PEIGTRGLYRG 574
A AG + L+ + P+D +KT++Q +S S ++I + + G RGL RG
Sbjct: 187 ATAGAAAGGLAAVVTTPLDVVKTQLQCQGVCGCDRFTSGSIGDVIRTIVKKDGYRGLMRG 366
Query: 575 SIPAILGQFSSHGLRTGIFEASK 597
IP +L + + +EA K
Sbjct: 367 WIPRMLFHAPAAAICWSTYEAGK 435
>TC214972 similar to UP|Q43649 (Q43649) Oxoglutarate malate translocator,
complete
Length = 1370
Score = 65.1 bits (157), Expect = 1e-10
Identities = 54/226 (23%), Positives = 100/226 (43%), Gaps = 12/226 (5%)
Frame = +2
Query: 520 SVLRSALAGGLSCALSCALLHPVDSIKTRVQASSMSFPEIIA-KLPEIGTRGLYRGSIPA 578
S ++ + GG S L+ ++ P+D IK R+Q S ++ + L G Y+G
Sbjct: 182 STIKPFVNGGASGMLATCVIQPIDMIKVRIQLGQGSAAQVTSTMLKNEGFAAFYKGLSAG 361
Query: 579 ILGQFSSHGLRTGIFE--ASKLVLVNVAPNLPELQVQSIASFCSTFLGTAVRIPCEVLKQ 636
+L Q + R G F+ +K + N LP L +++ + +G +V P ++
Sbjct: 362 LLRQATYTTARLGSFKILTAKAIEANDGKPLP-LYQKALCGLTAGAIGASVGSPADLALI 538
Query: 637 RLQAGL---------FNNVGEALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYAES 687
R+QA + N AL +G+ ++G G T+ R + + + Y +S
Sbjct: 539 RMQADATLPAAQRRNYTNAFHALYRITADEGVLALWKGAGPTVVRAMALNMGMLASYDQS 718
Query: 688 KKGVQKLLGRELEAWETIAVGALSGGLAAVVTTPFDVMKTRMMTAQ 733
+ + +G E + ++SG AA + PFD +KT++ Q
Sbjct: 719 VEFFRDSVGLG-EGATVLGASSVSGFFAAACSLPFDYVKTQIQKMQ 853
>TC222418 UP|Q8W1A3 (Q8W1A3) Uncoupling protein 1b (Fragment), partial (98%)
Length = 884
Score = 64.3 bits (155), Expect = 2e-10
Identities = 53/210 (25%), Positives = 93/210 (44%), Gaps = 11/210 (5%)
Frame = +3
Query: 565 EIGTRGLYRGSIPAILGQFSSHGLRTGIFEASKLVLVNV--APNLPELQVQSIASFCSTF 622
E G L++G +P + Q + GLR ++E K V ++P L + +A F +
Sbjct: 78 EEGFSALWKGIVPGLHRQCLNGGLRIALYEPVKNFYVGADHVGDVP-LSKKILAGFTTGA 254
Query: 623 LGTAVRIPCEVLKQRLQAG---------LFNNVGEALVGTWQQDGLKGFFRGTGATLCRE 673
+ AV P +++K RLQA ++ A +Q+G+ + G G + R
Sbjct: 255 MAIAVANPTDLVKVRLQAEGKLPPGVPRRYSGSLNAYSTIVRQEGVGALWTGIGPNIARN 434
Query: 674 VPFYVAGMGLYAESKKGVQKLLGRELEAWETIAVGALSGGLAAVVTTPFDVMKTRMMTAQ 733
A + Y + K+ + K+ G + G +G A V +P DV+K+RMM
Sbjct: 435 GIINAAELASYDQVKQTILKIPGFTDNVVTHLLAGLGAGFFAVCVGSPVDVVKSRMMGDS 614
Query: 734 GRSVSMSIVAFSILRHEGPLGLFKGAVPRF 763
++ L+++GP +KG +P F
Sbjct: 615 SYKSTLDCFV-KTLKNDGPFAFYKGFIPNF 701
Score = 37.0 bits (84), Expect = 0.034
Identities = 31/155 (20%), Positives = 62/155 (40%), Gaps = 12/155 (7%)
Frame = +3
Query: 523 RSALAGGLSCALSCALLHPVDSIKTRVQASSMSFPEIIAK-----------LPEIGTRGL 571
+ LAG + A++ A+ +P D +K R+QA P + + + + G L
Sbjct: 222 KKILAGFTTGAMAIAVANPTDLVKVRLQAEGKLPPGVPRRYSGSLNAYSTIVRQEGVGAL 401
Query: 572 YRGSIPAILGQFSSHGLRTGIFEASKLVLVNVAPNLPELQVQSIASFCSTFLGTAVRIPC 631
+ G P I + ++ K ++ + + +A + F V P
Sbjct: 402 WTGIGPNIARNGIINAAELASYDQVKQTILKIPGFTDNVVTHLLAGLGAGFFAVCVGSPV 581
Query: 632 EVLKQRLQA-GLFNNVGEALVGTWQQDGLKGFFRG 665
+V+K R+ + + + V T + DG F++G
Sbjct: 582 DVVKSRMMGDSSYKSTLDCFVKTLKNDGPFAFYKG 686
>TC212927 similar to UP|Q9ZNY4 (Q9ZNY4) Mitochondrial energy transfer protein
precursor, partial (43%)
Length = 762
Score = 63.9 bits (154), Expect = 3e-10
Identities = 47/169 (27%), Positives = 85/169 (49%), Gaps = 8/169 (4%)
Frame = +2
Query: 629 IPCEVLKQRL--QAGLFNNVGEALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYAE 686
IP E+LK RL Q G++ N+ +A V Q++G +RG ++L VP+ A Y
Sbjct: 62 IPLELLKTRLTVQRGVYKNLLDAFVRIIQEEGPAELYRGLTSSLIGVVPYAAANYLAYDT 241
Query: 687 SKKGVQKLLGR-ELEAWETIAVGALSGGLAAVVTTPFDVMKTRMMTA-----QGRSVSMS 740
+K +K + E+ T+ +G+ +G +++ T P +V M Q R++ +
Sbjct: 242 LRKAYKKAFKK*EIGNVMTLLIGSAAGAISSSATFPLEVACEHMQAGALNGRQYRNLLHA 421
Query: 741 IVAFSILRHEGPLGLFKGAVPRFFWIAPLGAMNFAGYELARKAMNKNDE 789
+V SIL EG GL++G + P ++F YE ++ + +N++
Sbjct: 422 LV--SILEKEGVGGLYRGL*LSCLKLVPAAGISFMCYEACKRVLVENEQ 562
>TC217975 similar to UP|Q9LY28 (Q9LY28) Peroxisomal Ca-dependent solute
carrier-like protein, partial (44%)
Length = 1187
Score = 63.2 bits (152), Expect = 4e-10
Identities = 36/135 (26%), Positives = 61/135 (44%), Gaps = 5/135 (3%)
Frame = +1
Query: 649 ALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKGVQKLLGRELEAWET--IA 706
A+ W+QDGL GFFRG G + + P + KK + + G + + +
Sbjct: 142 AVTKIWKQDGLLGFFRGNGLNVVKVSPESAIKFYAFEMLKKVIGEAHGNKSDIGTAGRLV 321
Query: 707 VGALSGGLAAVVTTPFDVMKTRMMTAQ---GRSVSMSIVAFSILRHEGPLGLFKGAVPRF 763
G +G +A P D++KTR+ T G+ + + +I EGP ++G VP
Sbjct: 322 AGGTAGAIAQAAIYPMDLIKTRLQTCPSEGGKVPKLGTLTMNIWVQEGPRAFYRGLVPSL 501
Query: 764 FWIAPLGAMNFAGYE 778
+ P A++ Y+
Sbjct: 502 LGMIPYAAIDLTAYD 546
Score = 48.1 bits (113), Expect = 1e-05
Identities = 41/178 (23%), Positives = 78/178 (43%), Gaps = 10/178 (5%)
Frame = +1
Query: 526 LAGGLSCALSCALLHPVDSIKT--RVQASSMSFPEIIAKL-PEIGTRGLYRGSIPAILGQ 582
LAGG++ +S P+D +K +VQ+ S + K+ + G G +RG+ ++
Sbjct: 37 LAGGIAGGISRTATAPLDRLKVVLQVQSEPASIMPAVTKIWKQDGLLGFFRGNGLNVVKV 216
Query: 583 FSSHGLRTGIFEASKLVLVNVAPNLPELQVQS--IASFCSTFLGTAVRIPCEVLKQRLQA 640
++ FE K V+ N ++ +A + + A P +++K RLQ
Sbjct: 217 SPESAIKFYAFEMLKKVIGEAHGNKSDIGTAGRLVAGGTAGAIAQAAIYPMDLIKTRLQT 396
Query: 641 -----GLFNNVGEALVGTWQQDGLKGFFRGTGATLCREVPFYVAGMGLYAESKKGVQK 693
G +G + W Q+G + F+RG +L +P+ + Y ++ K + K
Sbjct: 397 CPSEGGKVPKLGTLTMNIWVQEGPRAFYRGLVPSLLGMIPYAAIDLTAY-DTMKDISK 567
Score = 43.1 bits (100), Expect = 5e-04
Identities = 25/93 (26%), Positives = 44/93 (46%), Gaps = 5/93 (5%)
Frame = +1
Query: 708 GALSGGLAAVVTTPFDVMKTRMMTAQGRSVSMSIVAFSILRHEGPLGLFKGAVPRFFWIA 767
G ++GG++ T P D +K ++ Q S+ I + +G LG F+G ++
Sbjct: 43 GGIAGGISRTATAPLDRLKV-VLQVQSEPASIMPAVTKIWKQDGLLGFFRGNGLNVVKVS 219
Query: 768 PLGAMNFAGYELARKAM-----NKNDEAKTGNL 795
P A+ F +E+ +K + NK+D G L
Sbjct: 220 PESAIKFYAFEMLKKVIGEAHGNKSDIGTAGRL 318
>TC209762
Length = 950
Score = 62.0 bits (149), Expect = 1e-09
Identities = 45/193 (23%), Positives = 86/193 (44%), Gaps = 19/193 (9%)
Frame = +2
Query: 615 IASFCSTFLGTAVRIPCEVLKQRLQAGLFNNVGEALVG------TWQQDGLKGFFRGTGA 668
+A S+ +V +P +V+ Q+L ++ + G + DG++G +RG G
Sbjct: 20 VAGMTSSLFAQSVFVPIDVVSQKLMVQGYSGHAQYSGGLDVVRQVLRTDGIRGLYRGFGL 199
Query: 669 TLCREVPFYVAGMGLYAESKKGVQKLLGRELEAWETIAV-----------GALSGGLAAV 717
+ P Y S++ + + L + E G ++G ++
Sbjct: 200 SAITYAPASAVWWASYGSSQRFIWRFLDHGAKYDEVAPSLQKIMLVQATGGIIAGATSSC 379
Query: 718 VTTPFDVMKTRM--MTAQGRSVSMSIVAFSILRHEGPLGLFKGAVPRFFWIAPLGAMNFA 775
+TTP D +KTR+ M + RS S+ VA ++ +G G ++G PRFF ++ G
Sbjct: 380 ITTPLDTIKTRLQVMGHENRS-SIKQVAKDLINEDGWRGFYRGFGPRFFSMSAWGTSMIL 556
Query: 776 GYELARKAMNKND 788
YE ++ +K++
Sbjct: 557 TYEYLKRVCSKDE 595
Score = 31.6 bits (70), Expect = 1.4
Identities = 23/62 (37%), Positives = 34/62 (54%), Gaps = 5/62 (8%)
Frame = +2
Query: 521 VLRSALAGGLSCALSCALLHPVDSIKTRVQA---SSMSFPEIIAK--LPEIGTRGLYRGS 575
+L A G ++ A S + P+D+IKTR+Q + S + +AK + E G RG YRG
Sbjct: 329 MLVQATGGIIAGATSSCITTPLDTIKTRLQVMGHENRSSIKQVAKDLINEDGWRGFYRGF 508
Query: 576 IP 577
P
Sbjct: 509 GP 514
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.318 0.134 0.385
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 31,586,040
Number of Sequences: 63676
Number of extensions: 417023
Number of successful extensions: 2382
Number of sequences better than 10.0: 129
Number of HSP's better than 10.0 without gapping: 2238
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2330
length of query: 796
length of database: 12,639,632
effective HSP length: 105
effective length of query: 691
effective length of database: 5,953,652
effective search space: 4113973532
effective search space used: 4113973532
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 63 (28.9 bits)
Medicago: description of AC144484.11


