

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
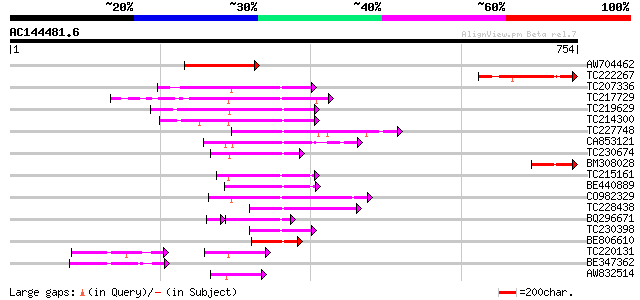
Query= AC144481.6 - phase: 0
(754 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AW704462 homologue to GP|16973451|gb phragmoplast-associated kin... 192 5e-49
TC222267 similar to UP|O23291 (O23291) Kinesin like protein, par... 125 7e-29
TC207336 similar to UP|O24147 (O24147) Kinesin-like protein, par... 98 1e-20
TC217729 similar to UP|Q9CAC9 (Q9CAC9) Kinesin-like protein; 736... 96 4e-20
TC219629 similar to UP|ATK1_ARATH (Q07970) Kinesin 1 (Kinesin-li... 96 8e-20
TC214300 similar to UP|ATK1_ARATH (Q07970) Kinesin 1 (Kinesin-li... 87 4e-17
TC227748 similar to UP|Q940Y8 (Q940Y8) AT3g16060/MSL1_10, partia... 81 1e-15
CA853121 79 7e-15
TC230674 homologue to UP|Q9M0X6 (Q9M0X6) Kinesin-like protein, p... 79 7e-15
BM308028 homologue to GP|16973451|gb| phragmoplast-associated ki... 78 2e-14
TC215161 similar to UP|Q6NQ77 (Q6NQ77) At4g05190, partial (19%) 76 6e-14
BE440889 74 2e-13
CO982329 62 7e-10
TC228438 similar to UP|Q6WJ05 (Q6WJ05) Central motor kinesin 1, ... 62 1e-09
BQ296671 similar to GP|18087642|gb AT5g27950/F15F15_20 {Arabidop... 51 9e-08
TC230398 homologue to UP|O24147 (O24147) Kinesin-like protein, p... 55 2e-07
BE806610 homologue to GP|15450501|gb| AT3g16060/MSL1_10 {Arabido... 54 2e-07
TC220131 similar to UP|ATK2_ARATH (P46864) Kinesin 2 (Kinesin-li... 53 4e-07
BE347362 50 4e-06
AW832514 46 5e-05
>AW704462 homologue to GP|16973451|gb phragmoplast-associated kinesin-related
protein 2 {Arabidopsis thaliana}, partial (11%)
Length = 423
Score = 192 bits (488), Expect = 5e-49
Identities = 99/100 (99%), Positives = 99/100 (99%)
Frame = +1
Query: 233 VKLEVMGKKAKNATYISGNEAGKISKEIQKVEKRRIVKSTLCNDRSSRSHCMVILDVPTV 292
VKLEVMGKKAKNATYISGNEAGKISKEIQKVEKRRIVKSTLCNDRSSRSHCMVILDVPTV
Sbjct: 124 VKLEVMGKKAKNATYISGNEAGKISKEIQKVEKRRIVKSTLCNDRSSRSHCMVILDVPTV 303
Query: 293 GGRLMLVDMAGSENIEQAGQTGFEAKMQTAKINQGNIALK 332
GGRLMLVDMAGSENIEQAGQTGFEAKMQTA INQGNIALK
Sbjct: 304 GGRLMLVDMAGSENIEQAGQTGFEAKMQTA*INQGNIALK 423
>TC222267 similar to UP|O23291 (O23291) Kinesin like protein, partial (3%)
Length = 507
Score = 125 bits (314), Expect = 7e-29
Identities = 77/147 (52%), Positives = 93/147 (62%), Gaps = 16/147 (10%)
Frame = +1
Query: 624 QKACLSTVYEEEGEEEAEQDHDKVEEDEEVEKEVIEEKRVCSVV--------------NK 669
Q+ACLSTVYEEEGE E E DEEVEK +IEEKRVCSV N
Sbjct: 1 QRACLSTVYEEEGEGEGE--------DEEVEKGIIEEKRVCSVEKPSGAGSLGTLNLSNT 156
Query: 670 SPKIEDYTGADKENNGSNRLLRIHNIFTLCGNQRELSQY-GTPIPTKKRSDESFDFKCSP 728
SP E++ + +RLLRI NIFTLCGN R+LSQ+ GTP+ TKKRSDE+F+F SP
Sbjct: 157 SPNKEEFCLKVSSGDNDSRLLRIQNIFTLCGNHRQLSQHIGTPVSTKKRSDETFEF--SP 330
Query: 729 VKSSEKKDSVLRVSNKENLEA-YVIGN 754
+ KDS+ + SNKEN E +V+GN
Sbjct: 331 ---ANDKDSIFKTSNKENFEPHHVLGN 402
>TC207336 similar to UP|O24147 (O24147) Kinesin-like protein, partial (22%)
Length = 1055
Score = 98.2 bits (243), Expect = 1e-20
Identities = 64/221 (28%), Positives = 114/221 (50%), Gaps = 9/221 (4%)
Frame = +3
Query: 197 VLEIYNEEIYDLLSTNGGGGGGGGFGFGWSKSNASKVKLEVMGKKA-KNATYISGNEAGK 255
++E+Y + + DLL G K +K + G +N T +S + +
Sbjct: 93 MVELYQDTLIDLLLPKNG------------KPLKLDIKKDSTGMVVVENVTVMSISTIEE 236
Query: 256 ISKEIQKVEKRRIVKSTLCNDRSSRSHCMVILDVPTVG--------GRLMLVDMAGSENI 307
++ IQ+ +RR + T ND SSRSH ++ + + + G+L VD+AGSE +
Sbjct: 237 LNSIIQRGSERRHISGTQMNDESSRSHLILSIVIESTNLQSQSVAKGKLSFVDLAGSERV 416
Query: 308 EQAGQTGFEAKMQTAKINQGNIALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKI 367
+++G TG + K + IN+ AL V+ S+++G H P+R+ KLTML+ DS +K
Sbjct: 417 KKSGSTGSQLK-EAQSINKSLSALGDVISSLSSGGQHTPYRNHKLTMLMSDSL-GGNAKT 590
Query: 368 LMILCASPDPKEIHKTISTLEYGAKAKCIVRGPHTPVKEED 408
LM + +P + +T ++L Y ++ + IV P+ V ++
Sbjct: 591 LMFVNVAPTESNLDETNNSLMYASRVRSIVNDPNKNVSSKE 713
>TC217729 similar to UP|Q9CAC9 (Q9CAC9) Kinesin-like protein; 73641-79546,
partial (26%)
Length = 1075
Score = 96.3 bits (238), Expect = 4e-20
Identities = 95/310 (30%), Positives = 144/310 (45%), Gaps = 13/310 (4%)
Frame = +1
Query: 134 TIMMYGPTGSGKSHTMFGCSKQAGIVYRALRDILGDGDTDSEGSDGDSSKGFCLGVRTFV 193
T M GP S KS G+ YRAL D+ + SS + +GV+
Sbjct: 40 TYTMSGPGLSSKSDW--------GVNYRALHDLFHISQSRR------SSIVYEVGVQ--- 168
Query: 194 QVTVLEIYNEEIYDLLSTNGGGGGGGGFGFGWSKSNASKVKLEVMGKKAKNATYISGNEA 253
++EIYNE++ DLLS+NG G W+ + + G +A+ S N
Sbjct: 169 ---MVEIYNEQVRDLLSSNGPQKRLGI----WNTAQPN-------GLAVPDASMHSVNSM 306
Query: 254 GKISKEIQKVEKRRIVKSTLCNDRSSRSHCMVILDV--------PTVGGRLMLVDMAGSE 305
+ + + R +T N+RSSRSH ++ + V + G L LVD+AGSE
Sbjct: 307 ADVLELMNIGLMNRATSATALNERSSRSHSVLSVHVRGTDLKTNTLLRGCLHLVDLAGSE 486
Query: 306 NIEQAGQTGFEAKMQTAKINQGNIALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKS 365
++++ TG K + IN+ AL V+ +++ SHVP+R+SKLT LLQ S ++
Sbjct: 487 RVDRSEATGDMLK-EAQHINKSLSALGDVIFALSQKSSHVPYRNSKLTQLLQSSL-GGQA 660
Query: 366 KILMILCASPDPKEIHKTISTLEYGAKAKCIVRGPHTPVKE-----EDSSSTVILGSRIA 420
K LM + +PD +T+STL++ + + G KE E L IA
Sbjct: 661 KTLMFVQLNPDVASYSETVSTLKFAERVSGVELGAARSNKEGRDVRELMEQLASLKDVIA 840
Query: 421 AMDEFIMKLQ 430
DE I +LQ
Sbjct: 841 RKDEEIERLQ 870
>TC219629 similar to UP|ATK1_ARATH (Q07970) Kinesin 1 (Kinesin-like protein
A), partial (26%)
Length = 923
Score = 95.5 bits (236), Expect = 8e-20
Identities = 74/233 (31%), Positives = 116/233 (49%), Gaps = 9/233 (3%)
Frame = +1
Query: 188 GVRTFVQVTVLEIYNEEIYDLLSTNGGGGGGGGFGFGWSKSNASKVKLEVMGKKAKNATY 247
G + + V++ EIYNE I DLLS+N G ++++A + K +
Sbjct: 43 GWKYTMHVSIYEIYNETIRDLLSSNRSSGND----HTRTENSAPTPSKQHTIKHESDLAT 210
Query: 248 ISGNEAGKISKEIQKVEKRRIVKSTLCNDRSSRSHCMVILDVP--------TVGGRLMLV 299
+ A +IS +Q+ + R V T N++SSRSH + L + V G L L+
Sbjct: 211 LEVCSAEEISSLLQQAAQSRSVGRTQMNEQSSRSHFVFKLRISGRNEKTEKQVQGVLNLI 390
Query: 300 DMAGSENIEQAGQTGFEAKMQTAKINQGNIALKRVVESIANGDSHVPFRDSKLTMLLQDS 359
D+AGSE + ++G TG K +T IN+ +L V+ ++A + HVPFR+SKLT LQ
Sbjct: 391 DLAGSERLSRSGATGDRLK-ETQAINKSLSSLSDVIFALAKKEEHVPFRNSKLTHFLQPY 567
Query: 360 FEDDKSKILMILCASPDPKEIHKTISTLEYGAKAK-CIVRGPHTPVKEEDSSS 411
D SK LM + SPD +++ +L + A+ C + P + SS
Sbjct: 568 LGGD-SKTLMFVNISPDQSSAGESLCSLRFAARVNACEIGIPRRQTQTSTRSS 723
>TC214300 similar to UP|ATK1_ARATH (Q07970) Kinesin 1 (Kinesin-like protein
A), partial (27%)
Length = 903
Score = 86.7 bits (213), Expect = 4e-17
Identities = 71/230 (30%), Positives = 111/230 (47%), Gaps = 17/230 (7%)
Frame = +2
Query: 200 IYNEEIYDLLSTNGGGGGGGGFGFGWSKSNASKVKLEVMGKKAKNATYISG------NEA 253
IYNE I DLL+TN G N + K ++ A T++S
Sbjct: 2 IYNETIRDLLATNKSSADGTPTRV----ENGTPGKQYMIKHDANGNTHVSDLTVVDVQSV 169
Query: 254 GKISKEIQKVEKRRIVKSTLCNDRSSRSHCMVILDV--------PTVGGRLMLVDMAGSE 305
+++ + + R V T N++SSRSH + L + V G L L+D+AGSE
Sbjct: 170 KEVAFLLNQAASSRSVGKTQMNEQSSRSHFVFTLRIYGVNESTDQQVQGILNLIDLAGSE 349
Query: 306 NIEQAGQTGFEAKMQTAKINQGNIALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKS 365
+ ++G TG K +T IN+ +L V+ ++A + H+PFR+SKLT LLQ D S
Sbjct: 350 RLSRSGSTGDRLK-ETQAINKSLSSLSDVIFALAKKEDHIPFRNSKLTYLLQPCLGGD-S 523
Query: 366 KILMILCASPDPKEIHKTISTLEYGAKAK-CIVRGP--HTPVKEEDSSST 412
K LM + SPD +++ +L + ++ C + P HT + +S T
Sbjct: 524 KTLMFVNISPDQASSGESLCSLRFASRVNACEIGTPRRHTNGRPIESRLT 673
>TC227748 similar to UP|Q940Y8 (Q940Y8) AT3g16060/MSL1_10, partial (34%)
Length = 1192
Score = 81.3 bits (199), Expect = 1e-15
Identities = 66/244 (27%), Positives = 115/244 (47%), Gaps = 16/244 (6%)
Frame = +3
Query: 295 RLMLVDMAGSENIEQAGQTGFEAKMQTAKINQGNIALKRVVESIANGDSHVPFRDSKLTM 354
+L +D+AGSE + +++ A+IN+ +ALK + ++ N H+PFR SKLT
Sbjct: 3 KLSFIDLAGSERGADTTDNDKQTRIEGAEINKSLLALKECIRALDNDQGHIPFRGSKLTE 182
Query: 355 LLQDSFEDDKSKILMILCASPDPKEIHKTISTLEYGAKAKCIVRGPHTPVKEEDS----- 409
+L+DSF + S+ +MI C SP T++TL Y + K + +G ++ S
Sbjct: 183 VLRDSFVGN-SRTVMISCISPSTGSCEHTLNTLRYADRVKSLSKGNNSKKDVLSSNFNLK 359
Query: 410 -SSTVILGSRIAA------MDEFIMKLQMENKLREKERNEAHKKLMKKEEEIAALRTKVE 462
SSTV L S + D + + + ++ +E E K +KK ++ + T +
Sbjct: 360 ESSTVPLSSVTGSAYEDRVTDGWPDENEGDDFSPSEEYYEQVKPPLKKNGKMESYATTDD 539
Query: 463 TAPASEEEINL----KVNERTRHLRQELEKKLEECQRMTNEFVELERKRMEERILQQQEE 518
+I KV +T H +L L+E + V R ++EE + +EE
Sbjct: 540 KLKKPSGQIKWKDLPKVEPQTTHAEDDLNALLQE----EEDLVNAHRTQVEETMNIVREE 707
Query: 519 VEIL 522
+ +L
Sbjct: 708 MNLL 719
>CA853121
Length = 677
Score = 79.0 bits (193), Expect = 7e-15
Identities = 62/220 (28%), Positives = 110/220 (49%), Gaps = 8/220 (3%)
Frame = +3
Query: 258 KEIQKVEKRRIVKSTLCNDRSSRSHCMV---ILDVPTVGGR-----LMLVDMAGSENIEQ 309
++++ + R V ST N+ SSRSHC++ +L + G+ L LVD+AGSE + +
Sbjct: 84 EKLKSGNQARSVGSTSANELSSRSHCLLRVTVLGENLINGQKTRSHLWLVDLAGSERVGK 263
Query: 310 AGQTGFEAKMQTAKINQGNIALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILM 369
G K ++ IN+ AL V+ ++A+ +H+P+R+SKLT +LQ S D K LM
Sbjct: 264 TEAEGERLK-ESQFINKSLSALGDVISALASKSAHIPYRNSKLTHILQSSLGGD-CKTLM 437
Query: 370 ILCASPDPKEIHKTISTLEYGAKAKCIVRGPHTPVKEEDSSSTVILGSRIAAMDEFIMKL 429
+ SP ++ +T+ +L + + + I GP K+ D + E
Sbjct: 438 FVQISPGAADLTETLCSLNFATRVRGIESGPAR--KQTD-------------LTELNKYK 572
Query: 430 QMENKLREKERNEAHKKLMKKEEEIAALRTKVETAPASEE 469
QM K++ E K+ K ++ + A++ ++ T + E
Sbjct: 573 QMVEKVKHDE-----KETRKLQDNLQAMQMRLTTRETTRE 677
>TC230674 homologue to UP|Q9M0X6 (Q9M0X6) Kinesin-like protein, partial (20%)
Length = 811
Score = 79.0 bits (193), Expect = 7e-15
Identities = 52/133 (39%), Positives = 74/133 (55%), Gaps = 8/133 (6%)
Frame = +1
Query: 267 RIVKSTLCNDRSSRSHCMVILDVP--------TVGGRLMLVDMAGSENIEQAGQTGFEAK 318
R V T N++SSRSH + L + V G L L+D+AGSE + ++G TG K
Sbjct: 94 RSVGRTHMNEQSSRSHFVXTLRISGTNSNTDQQVQGVLNLIDLAGSERLSRSGATGDRLK 273
Query: 319 MQTAKINQGNIALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPK 378
+T IN+ +L V+ ++A HVPFR+SKLT LLQ D SK LM + SPDP
Sbjct: 274 -ETQAINKSLSSLSDVIFALAKKQEHVPFRNSKLTYLLQPCLGGD-SKTLMFVNISPDPS 447
Query: 379 EIHKTISTLEYGA 391
+++ +L + A
Sbjct: 448 STGESLCSLRFAA 486
>BM308028 homologue to GP|16973451|gb| phragmoplast-associated
kinesin-related protein 2 {Arabidopsis thaliana},
partial (1%)
Length = 366
Score = 77.8 bits (190), Expect = 2e-14
Identities = 44/64 (68%), Positives = 54/64 (83%), Gaps = 3/64 (4%)
Frame = +1
Query: 694 NIFTLCGNQRELSQY-GTPI-PTKKRSDESFDFKCSPVKSSEKKDSVLRVSNKENLEAY- 750
NIFTLCGNQRELSQ+ GTP+ PTKKRSDE+F+F SP K+S+ KDSV ++SNKEN E +
Sbjct: 1 NIFTLCGNQRELSQHIGTPVQPTKKRSDETFEF--SPAKTSD-KDSVFKISNKENFEPHN 171
Query: 751 VIGN 754
V+GN
Sbjct: 172 VLGN 183
>TC215161 similar to UP|Q6NQ77 (Q6NQ77) At4g05190, partial (19%)
Length = 692
Score = 75.9 bits (185), Expect = 6e-14
Identities = 50/145 (34%), Positives = 79/145 (54%), Gaps = 8/145 (5%)
Frame = +2
Query: 275 NDRSSRSHCMVILDV--------PTVGGRLMLVDMAGSENIEQAGQTGFEAKMQTAKINQ 326
N++SSRSH + L + V G L L+D+AGSE + ++G TG K +T IN+
Sbjct: 44 NEQSSRSHFVFTLRIYGVNESTDXXVQGVLNLIDLAGSERLSKSGSTGDRLK-ETQAINK 220
Query: 327 GNIALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTIST 386
+L V+ ++A + HVPFR+SKLT LLQ S LM + SPDP + +++ +
Sbjct: 221 SLSSLSDVIFALAKKEDHVPFRNSKLTYLLQPCL-GGYSNTLMFVNISPDPSSVGESLCS 397
Query: 387 LEYGAKAKCIVRGPHTPVKEEDSSS 411
L + ++ G TP ++ + S
Sbjct: 398 LRFASRVNACEIG--TPRRQTNGRS 466
>BE440889
Length = 484
Score = 74.3 bits (181), Expect = 2e-13
Identities = 46/128 (35%), Positives = 75/128 (57%)
Frame = +2
Query: 286 ILDVPTVGGRLMLVDMAGSENIEQAGQTGFEAKMQTAKINQGNIALKRVVESIANGDSHV 345
+L+ + +L LVD+AGSE + + G K + IN+ AL V+ ++A SH+
Sbjct: 59 LLNGESTKSKLWLVDLAGSERLAKTDVQGERLK-EAQNINRSLSALGDVISALAAKSSHI 235
Query: 346 PFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIHKTISTLEYGAKAKCIVRGPHTPVK 405
P+R+SKLT LLQDS D SK LM + SP +++ +T+S+L + + + + G PVK
Sbjct: 236 PYRNSKLTHLLQDSLGGD-SKTLMFVQISPSDQDVGETLSSLNFATRVRGVELG---PVK 403
Query: 406 EEDSSSTV 413
++ +S V
Sbjct: 404 KQIDTSEV 427
>CO982329
Length = 852
Score = 62.4 bits (150), Expect = 7e-10
Identities = 64/232 (27%), Positives = 106/232 (45%), Gaps = 14/232 (6%)
Frame = -3
Query: 265 KRRIVKSTLCNDRSSRSHCMVILDVPTVG------------GRLMLVDMAGSENIEQAGQ 312
+ R V ST N SSRSH + L + + +L L+D+AGSE+ +A
Sbjct: 838 EHRHVGSTNFNLLSSRSHTIFSLTIESSPCGENNEGEAITLSQLNLIDLAGSES-SRAET 662
Query: 313 TGFEAKMQTAKINQGNIALKRVVESIANGD-SHVPFRDSKLTMLLQDSFEDDKSKILMIL 371
TG + + + IN+ + L V+ + G SH+P+RDSKLT LLQ S +I +I
Sbjct: 661 TGMRRR-EGSYINKSLLTLGTVISKLTEGKASHIPYRDSKLTRLLQSSL-SGHGRISLIC 488
Query: 372 CASPDPKEIHKTISTLEYGAKAKCI-VRGPHTPVKEEDSSSTVILGSRIAAMDEFIMKLQ 430
+P +T +TL++ +AK I ++ + +E S +E Q
Sbjct: 487 TVTPSSSNAEETHNTLKFAHRAKHIEIQAAQNTIIDEKSLIKKYQHEIQFLKEELEQMRQ 308
Query: 431 MENKLREKERNEAHKKLMKKEEEIAALRTKVETAPASEEEINLKVNERTRHL 482
++ KE E L+K++ E + K+++ EEE + R + L
Sbjct: 307 GIVSVQSKETVEVDFVLLKQKLEDG--QVKLQSRLEEEEEAKAALLGRIQRL 158
>TC228438 similar to UP|Q6WJ05 (Q6WJ05) Central motor kinesin 1, partial
(13%)
Length = 809
Score = 61.6 bits (148), Expect = 1e-09
Identities = 41/152 (26%), Positives = 76/152 (49%), Gaps = 3/152 (1%)
Frame = +2
Query: 319 MQTAKINQGNIALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPK 378
++ A IN+ +ALK + ++ N H+PFR SKLT +L+DSF + SK +MI C SP
Sbjct: 2 IEGA*INKSLLALKECIRALDNDQIHIPFRGSKLTEVLRDSFVGN-SKTVMISCISPGAG 178
Query: 379 EIHKTISTLEYGAKAKCIVRGPHTPVKEEDSSSTVILGSRIAAMDEFIMKLQME--NKLR 436
T++TL Y + K + + + + ++ +++ F E N R
Sbjct: 179 SCEHTLNTLRYADRVKSLSKSGNPRKDQVPNAVPQTNNKDVSSTSSFPASSGAEDFNDQR 358
Query: 437 EKERNEAHKKLMKKEEEI-AALRTKVETAPAS 467
+++ + +K ++KE + ++ V+ P S
Sbjct: 359 QEKTIDMGRKFVEKENSLHSSAAASVDKQPVS 454
>BQ296671 similar to GP|18087642|gb AT5g27950/F15F15_20 {Arabidopsis
thaliana}, partial (20%)
Length = 422
Score = 51.2 bits (121), Expect(2) = 9e-08
Identities = 32/94 (34%), Positives = 52/94 (55%)
Frame = +3
Query: 287 LDVPTVGGRLMLVDMAGSENIEQAGQTGFEAKMQTAKINQGNIALKRVVESIANGDSHVP 346
L+ + +L ++D+ GSE + + G G A IN AL VV ++ HVP
Sbjct: 141 LEAKSEVSKLWMIDLGGSERLLKTGAKGLTLDEGRA-INLSLSALADVVAALKRKRCHVP 317
Query: 347 FRDSKLTMLLQDSFEDDKSKILMILCASPDPKEI 380
+R+SKLT +L+DS SK+LM++ SP +++
Sbjct: 318 YRNSKLTQILKDSL-GYGSKVLMLVHISPSEEDV 416
Score = 23.9 bits (50), Expect(2) = 9e-08
Identities = 11/26 (42%), Positives = 15/26 (57%)
Frame = +1
Query: 262 KVEKRRIVKSTLCNDRSSRSHCMVIL 287
K ++ R T N+ SSRSHC + L
Sbjct: 25 KGKRFRSTSWTNVNEASSRSHCFIYL 102
>TC230398 homologue to UP|O24147 (O24147) Kinesin-like protein, partial (10%)
Length = 599
Score = 54.7 bits (130), Expect = 2e-07
Identities = 28/89 (31%), Positives = 50/89 (55%)
Frame = +3
Query: 320 QTAKINQGNIALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKE 379
+ IN+ AL V+ ++++G H+P+R+ KLTML+ DS +K LM + SP
Sbjct: 6 EAQSINKSLSALGDVISALSSGGQHIPYRNHKLTMLMSDSL-GGNAKTLMFVNVSPVESS 182
Query: 380 IHKTISTLEYGAKAKCIVRGPHTPVKEED 408
+ +T ++L Y ++ + IV P V ++
Sbjct: 183 LDETHNSLMYASRVRSIVNDPSKNVSSKE 269
>BE806610 homologue to GP|15450501|gb| AT3g16060/MSL1_10 {Arabidopsis
thaliana}, partial (10%)
Length = 220
Score = 54.3 bits (129), Expect = 2e-07
Identities = 28/68 (41%), Positives = 42/68 (61%)
Frame = +2
Query: 322 AKINQGNIALKRVVESIANGDSHVPFRDSKLTMLLQDSFEDDKSKILMILCASPDPKEIH 381
A+IN+ + LK + ++ N H+PFR SKLT +L+DSF + S+ +MI C SP
Sbjct: 11 AEINKSLLVLKECIRALDNDQGHIPFRGSKLTEVLRDSFVGN-SRTVMISCISPMFGSCE 187
Query: 382 KTISTLEY 389
T++TL Y
Sbjct: 188 HTLNTLRY 211
>TC220131 similar to UP|ATK2_ARATH (P46864) Kinesin 2 (Kinesin-like protein
B), partial (37%)
Length = 1044
Score = 53.1 bits (126), Expect = 4e-07
Identities = 41/134 (30%), Positives = 68/134 (50%), Gaps = 5/134 (3%)
Frame = +3
Query: 83 SNSRSIRVKADFGYRDFTLDGVSVSEEEELDLFYK--KFVESRINGVKLGDKCTIMMYGP 140
++ R+I + + FT D V E + ++F + + V+S ++G K+ I YG
Sbjct: 270 TSGRAIDLAQNGQKHAFTFDKVFTPEASQEEVFVEISQLVQSALDGYKV----CIFAYGQ 437
Query: 141 TGSGKSHTMFGCS---KQAGIVYRALRDILGDGDTDSEGSDGDSSKGFCLGVRTFVQVTV 197
TGSGK++TM G ++ G++ R+L I + G + +QV++
Sbjct: 438 TGSGKTYTMMGRPGHPEEKGLIPRSLEQIFQTKQSQQPQ-----------GWKYEMQVSM 584
Query: 198 LEIYNEEIYDLLST 211
LEIYNE I DL+ST
Sbjct: 585 LEIYNETIRDLIST 626
Score = 52.4 bits (124), Expect = 7e-07
Identities = 33/96 (34%), Positives = 51/96 (52%), Gaps = 8/96 (8%)
Frame = +1
Query: 260 IQKVEKRRIVKSTLCNDRSSRSHCMVILDV--------PTVGGRLMLVDMAGSENIEQAG 311
+ + R V T N++SSRSH + L + V G L L+D+AGSE + ++G
Sbjct: 760 LNQAANSRSVGKTQMNEQSSRSHFVFTLRIYGVNESTDQQVQGVLNLIDLAGSERLSKSG 939
Query: 312 QTGFEAKMQTAKINQGNIALKRVVESIANGDSHVPF 347
TG K +T IN+ +L V+ ++A + HVPF
Sbjct: 940 STGDRLK-ETQAINKSLSSLSDVIFALAKKEDHVPF 1044
>BE347362
Length = 448
Score = 50.1 bits (118), Expect = 4e-06
Identities = 39/135 (28%), Positives = 67/135 (48%), Gaps = 2/135 (1%)
Frame = +1
Query: 80 QASSNSRSIRVKADFGYRDFTLDGVSVSEEEELDLFY--KKFVESRINGVKLGDKCTIMM 137
++SS++ + AD + F D V E+ + +F K V S ++G + I
Sbjct: 7 ESSSDNELQVICADSSKKQFKFDHVFGPEDNQETVFQQTKPIVTSVLDGYNV----CIFA 174
Query: 138 YGPTGSGKSHTMFGCSKQAGIVYRALRDILGDGDTDSEGSDGDSSKGFCLGVRTFVQVTV 197
YG TG+GK+ TM G + G+ YR L ++ + + G+ ++ + V++
Sbjct: 175 YGQTGTGKTFTMEGTPEHRGVNYRTLEELFRITE-ERHGT-----------MKYELSVSM 318
Query: 198 LEIYNEEIYDLLSTN 212
LE+YNE+I DLL N
Sbjct: 319 LEVYNEKIRDLLVEN 363
>AW832514
Length = 412
Score = 46.2 bits (108), Expect = 5e-05
Identities = 32/84 (38%), Positives = 42/84 (49%), Gaps = 9/84 (10%)
Frame = +3
Query: 267 RIVKSTLCNDRSSRSHCMVIL---------DVPTVGGRLMLVDMAGSENIEQAGQTGFEA 317
R V T N SSRSHC+ I D T G+L+LVD+AGSE +E+ G G
Sbjct: 156 RAVGETQMNVASSRSHCIYIFTIQQEFLSRDKRTRFGKLILVDLAGSEKVEKTGAEG-RV 332
Query: 318 KMQTAKINQGNIALKRVVESIANG 341
+ IN+ AL V+ S+ G
Sbjct: 333 LEEAKTINKSLSALGNVINSLTCG 404
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.312 0.131 0.366
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 28,908,230
Number of Sequences: 63676
Number of extensions: 461724
Number of successful extensions: 9919
Number of sequences better than 10.0: 245
Number of HSP's better than 10.0 without gapping: 5046
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 8055
length of query: 754
length of database: 12,639,632
effective HSP length: 104
effective length of query: 650
effective length of database: 6,017,328
effective search space: 3911263200
effective search space used: 3911263200
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 63 (28.9 bits)
Medicago: description of AC144481.6


