

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144481.4 + phase: 0
(152 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
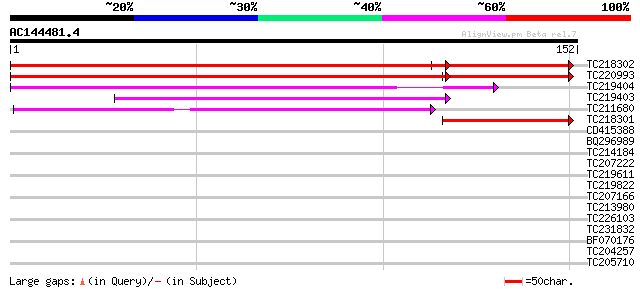
TC218302 similar to UP|Q6K1Q5 (Q6K1Q5) Glycolipid transfer prote... 215 3e-65
TC220993 similar to UP|Q6K1Q5 (Q6K1Q5) Glycolipid transfer prote... 203 7e-59
TC219404 weakly similar to UP|Q9LU33 (Q9LU33) Arabidopsis thalia... 75 1e-14
TC219403 weakly similar to UP|Q7FB31 (Q7FB31) OJ000114_01.1 prot... 69 7e-13
TC211680 weakly similar to UP|Q01494 (Q01494) HET-C2, partial (91%) 62 1e-10
TC218301 49 9e-07
CD415388 similar to GP|22022510|gb| AT4g39670/T19P19_60 {Arabido... 29 0.76
BQ296989 29 0.76
TC214184 similar to UP|Q6NLW5 (Q6NLW5) At5g48720, partial (29%) 29 0.99
TC207222 28 1.3
TC219611 similar to UP|Q8DLK1 (Q8DLK1) ABC transporter subunit, ... 28 2.2
TC219822 homologue to UP|CENB_HUMAN (P07199) Major centromere au... 27 2.9
TC207166 similar to UP|O23207 (O23207) Minor allergen, partial (... 27 3.8
TC213980 27 3.8
TC226103 weakly similar to UP|Q9FGQ2 (Q9FGQ2) MtN3-like protein,... 27 4.9
TC231832 weakly similar to UP|Q8TVZ4 (Q8TVZ4) Uncharacterized pr... 26 6.4
BF070176 GP|310561|gb|A ascorbate peroxidase {Glycine max}, part... 26 6.4
TC204257 UP|Q76LA8 (Q76LA8) Cytosolic ascorbate peroxidase 1 , ... 26 6.4
TC205710 similar to UP|Q8L5X1 (Q8L5X1) Fimbrin-like protein (Act... 26 8.4
>TC218302 similar to UP|Q6K1Q5 (Q6K1Q5) Glycolipid transfer protein-like,
complete
Length = 948
Score = 215 bits (548), Expect(2) = 3e-65
Identities = 105/118 (88%), Positives = 111/118 (93%)
Frame = +2
Query: 1 MAIVKSDIGGNISRLESKYLSNPAKFNCLYSLVQVEVETKTAKASSSCTTGLLWLTRAMD 60
MA+VKSDIGGNISRLESKY SNP KFN LYSLVQVEVETKTAK+SSSCT GLLWLTRAMD
Sbjct: 224 MALVKSDIGGNISRLESKYSSNPTKFNYLYSLVQVEVETKTAKSSSSCTNGLLWLTRAMD 403
Query: 61 FLVAVFRNLIEHADWSMSQACTDSYYKTLKKWHGWLASSTVTVAMMLAPDRKKFMEVV 118
FLVA+F+NLIEHADWSMSQACTDSY KTLKKWHGWLASS+ TVAM LAPDRKKFMEV+
Sbjct: 404 FLVALFQNLIEHADWSMSQACTDSYNKTLKKWHGWLASSSFTVAMKLAPDRKKFMEVI 577
Score = 49.7 bits (117), Expect(2) = 3e-65
Identities = 25/38 (65%), Positives = 29/38 (75%)
Frame = +3
Query: 114 FMEVVISMLTLSNFVLAFLLSLKRITSFWLDLDWMN*R 151
+ +VIS+L NFVL FLLSLKRITSFWL + WM *R
Sbjct: 576 YKALVISVLIYRNFVLTFLLSLKRITSFWLVVAWMI*R 689
>TC220993 similar to UP|Q6K1Q5 (Q6K1Q5) Glycolipid transfer protein-like,
complete
Length = 721
Score = 203 bits (516), Expect(2) = 7e-59
Identities = 98/118 (83%), Positives = 106/118 (89%)
Frame = +2
Query: 1 MAIVKSDIGGNISRLESKYLSNPAKFNCLYSLVQVEVETKTAKASSSCTTGLLWLTRAMD 60
MA+VKSDIGGNISR E+ Y SNP++FN LYSLVQVEVETKTAK+SSSCT GLLWLTRAMD
Sbjct: 134 MALVKSDIGGNISRSETMYSSNPSRFNYLYSLVQVEVETKTAKSSSSCTNGLLWLTRAMD 313
Query: 61 FLVAVFRNLIEHADWSMSQACTDSYYKTLKKWHGWLASSTVTVAMMLAPDRKKFMEVV 118
FLVA+FRNLIEH DW MSQACTDSY KTLKKWHGWLASS+ TV M LAPDRKKFM V+
Sbjct: 314 FLVALFRNLIEHEDWPMSQACTDSYNKTLKKWHGWLASSSFTVVMKLAPDRKKFMNVI 487
Score = 40.4 bits (93), Expect(2) = 7e-59
Identities = 22/35 (62%), Positives = 23/35 (64%)
Frame = +3
Query: 117 VVISMLTLSNFVLAFLLSLKRITSFWLDLDWMN*R 151
+ ISML NFV FLLS KR TSFWLD M *R
Sbjct: 495 LAISMLISRNFVQPFLLSFKRFTSFWLDFA*MI*R 599
>TC219404 weakly similar to UP|Q9LU33 (Q9LU33) Arabidopsis thaliana genomic
DNA, chromosome 3, P1 clone: MXL8, partial (48%)
Length = 540
Score = 75.1 bits (183), Expect = 1e-14
Identities = 42/131 (32%), Positives = 65/131 (49%)
Frame = +3
Query: 1 MAIVKSDIGGNISRLESKYLSNPAKFNCLYSLVQVEVETKTAKASSSCTTGLLWLTRAMD 60
MA+++ D+ NI RLE + NP+ + L +++ E ++ SSC+ LWLTR++D
Sbjct: 63 MAVLRQDVSQNIKRLEVMHELNPSMNSNLVEILKSEASKGKSRKRSSCSKAFLWLTRSLD 242
Query: 61 FLVAVFRNLIEHADWSMSQACTDSYYKTLKKWHGWLASSTVTVAMMLAPDRKKFMEVVIS 120
F A+ ++L M Q + Y TL WHGW++S+ V + S
Sbjct: 243 FSSALLKSLENDPKKDMEQIVQECYDVTLSPWHGWISSAAFRVI------------TICS 386
Query: 121 MLTLSNFVLAF 131
L L N V AF
Sbjct: 387 FLHLFNHVYAF 419
>TC219403 weakly similar to UP|Q7FB31 (Q7FB31) OJ000114_01.1 protein, partial
(39%)
Length = 607
Score = 69.3 bits (168), Expect = 7e-13
Identities = 32/90 (35%), Positives = 50/90 (55%)
Frame = +1
Query: 29 LYSLVQVEVETKTAKASSSCTTGLLWLTRAMDFLVAVFRNLIEHADWSMSQACTDSYYKT 88
L +++ E ++ SSC+ LWLTR++DF A+ ++L M Q + Y T
Sbjct: 4 LVEILKSEASKGKSRKRSSCSKAFLWLTRSLDFSSALLKSLENDPKKDMEQIVQECYDVT 183
Query: 89 LKKWHGWLASSTVTVAMMLAPDRKKFMEVV 118
L WHGW++S+ VA L PD K FM+++
Sbjct: 184 LSPWHGWISSAAFRVAKKLVPDSKTFMDLL 273
>TC211680 weakly similar to UP|Q01494 (Q01494) HET-C2, partial (91%)
Length = 832
Score = 61.6 bits (148), Expect = 1e-10
Identities = 34/113 (30%), Positives = 59/113 (52%)
Frame = +3
Query: 2 AIVKSDIGGNISRLESKYLSNPAKFNCLYSLVQVEVETKTAKASSSCTTGLLWLTRAMDF 61
++VK+D+ GNI++++ +Y A+ L LV E+++K A T GLLWLTR ++F
Sbjct: 225 SLVKNDMNGNITKIQQRYTEATAESTTLQELVVNELKSKKHVA----TEGLLWLTRGLEF 392
Query: 62 LVAVFRNLIEHADWSMSQACTDSYYKTLKKWHGWLASSTVTVAMMLAPDRKKF 114
+ I + ++ + ++Y TLK H ++ + AM P R F
Sbjct: 393 MCLALNANINNKTEELATSFRNAYGNTLKPHHSFVVKPLFSAAMSACPARNDF 551
>TC218301
Length = 327
Score = 48.9 bits (115), Expect = 9e-07
Identities = 25/35 (71%), Positives = 28/35 (79%)
Frame = +2
Query: 117 VVISMLTLSNFVLAFLLSLKRITSFWLDLDWMN*R 151
+VIS+L NFVL FLLSLKRITSFWL + WM *R
Sbjct: 8 LVISVLIYRNFVLTFLLSLKRITSFWLVVVWMI*R 112
>CD415388 similar to GP|22022510|gb| AT4g39670/T19P19_60 {Arabidopsis
thaliana}, partial (41%)
Length = 510
Score = 29.3 bits (64), Expect = 0.76
Identities = 12/52 (23%), Positives = 23/52 (44%)
Frame = -1
Query: 64 AVFRNLIEHADWSMSQACTDSYYKTLKKWHGWLASSTVTVAMMLAPDRKKFM 115
A+F L+ D S+ + + +Y + +H W + V M P R + +
Sbjct: 507 AIFEQLLSTDDSSLKEVASTAYGQVCAPYHTWAVRTAVYAGMYTLPTRDQLL 352
>BQ296989
Length = 428
Score = 29.3 bits (64), Expect = 0.76
Identities = 14/38 (36%), Positives = 23/38 (59%)
Frame = -2
Query: 14 RLESKYLSNPAKFNCLYSLVQVEVETKTAKASSSCTTG 51
R S L+ P+K+ S VQ ++ +TA+A+ +CT G
Sbjct: 226 RSTSSLLTIPSKYGVKISGVQFSLKLRTAEAACACTRG 113
>TC214184 similar to UP|Q6NLW5 (Q6NLW5) At5g48720, partial (29%)
Length = 791
Score = 28.9 bits (63), Expect = 0.99
Identities = 17/47 (36%), Positives = 21/47 (44%), Gaps = 3/47 (6%)
Frame = +2
Query: 84 SYYKTLKKWHGWLASSTVTVAMMLAPDRKKFM---EVVISMLTLSNF 127
S Y+ L+ GWLA M L PD F EV + + L NF
Sbjct: 23 SNYEDLESAEGWLADCFKDAEMQLCPDDLNFSGADEVQVDVADLGNF 163
>TC207222
Length = 2191
Score = 28.5 bits (62), Expect = 1.3
Identities = 14/54 (25%), Positives = 29/54 (52%), Gaps = 2/54 (3%)
Frame = -3
Query: 92 WHGWL--ASSTVTVAMMLAPDRKKFMEVVISMLTLSNFVLAFLLSLKRITSFWL 143
WH + A + + + +++ P + +V+ + T+S+ VL + KRIT W+
Sbjct: 440 WHMCISTAKTXIIIPLLITP-----VPIVVMVRTISSLVLTTITISKRITLLWM 294
>TC219611 similar to UP|Q8DLK1 (Q8DLK1) ABC transporter subunit, partial
(38%)
Length = 776
Score = 27.7 bits (60), Expect = 2.2
Identities = 16/41 (39%), Positives = 23/41 (56%)
Frame = +2
Query: 13 SRLESKYLSNPAKFNCLYSLVQVEVETKTAKASSSCTTGLL 53
SR+ SK +S NC LVQV + + A+ SS C + L+
Sbjct: 194 SRIISKGISVGHSRNCYRGLVQVLSKAENARNSSQCDSMLI 316
>TC219822 homologue to UP|CENB_HUMAN (P07199) Major centromere autoantigen B
(Centromere protein B) (CENP-B), partial (5%)
Length = 710
Score = 27.3 bits (59), Expect = 2.9
Identities = 13/47 (27%), Positives = 25/47 (52%)
Frame = +1
Query: 1 MAIVKSDIGGNISRLESKYLSNPAKFNCLYSLVQVEVETKTAKASSS 47
M +V + G + ++ + + CL S +Q+ ++T TA +SSS
Sbjct: 553 MLVVIRSLKGTLETIKVDVQAIKERVKCLQSFLQIYLKTATASSSSS 693
>TC207166 similar to UP|O23207 (O23207) Minor allergen, partial (79%)
Length = 1187
Score = 26.9 bits (58), Expect = 3.8
Identities = 13/26 (50%), Positives = 17/26 (65%)
Frame = -1
Query: 12 ISRLESKYLSNPAKFNCLYSLVQVEV 37
+S LES +L +P KFNC+ L EV
Sbjct: 263 LSCLESNHLLSPLKFNCVNILSNNEV 186
>TC213980
Length = 516
Score = 26.9 bits (58), Expect = 3.8
Identities = 17/57 (29%), Positives = 30/57 (51%)
Frame = -3
Query: 46 SSCTTGLLWLTRAMDFLVAVFRNLIEHADWSMSQACTDSYYKTLKKWHGWLASSTVT 102
+SC+ L+W +R++ F R+ A ++S ++S TL+ LA+ST T
Sbjct: 250 ASCSILLIWRSRSLTFA----RSSFSSASLTISSTISNSVLLTLQSQPA*LAASTKT 92
>TC226103 weakly similar to UP|Q9FGQ2 (Q9FGQ2) MtN3-like protein, partial (67%)
Length = 1423
Score = 26.6 bits (57), Expect = 4.9
Identities = 13/28 (46%), Positives = 17/28 (60%)
Frame = -1
Query: 120 SMLTLSNFVLAFLLSLKRITSFWLDLDW 147
S LTLS+ +L FLL L ++S W W
Sbjct: 1051 SQLTLSSVLLLFLLMLFLLSSVWS*ATW 968
>TC231832 weakly similar to UP|Q8TVZ4 (Q8TVZ4) Uncharacterized protein
conserved in archaea, partial (14%)
Length = 679
Score = 26.2 bits (56), Expect = 6.4
Identities = 15/51 (29%), Positives = 29/51 (56%)
Frame = +2
Query: 71 EHADWSMSQACTDSYYKTLKKWHGWLASSTVTVAMMLAPDRKKFMEVVISM 121
E ++W +++ S+ KTL WH L +S +T A+ + R K +++ S+
Sbjct: 98 EISNWGLNK----SFGKTLLGWH*ELLASEITKAV*IWVSRSKARKLMYSL 238
>BF070176 GP|310561|gb|A ascorbate peroxidase {Glycine max}, partial (38%)
Length = 330
Score = 26.2 bits (56), Expect = 6.4
Identities = 11/26 (42%), Positives = 15/26 (57%)
Frame = +2
Query: 47 SCTTGLLWLTRAMDFLVAVFRNLIEH 72
SCT G W+ ++D + FR LI H
Sbjct: 230 SCTQGAFWI*GSLDL*SSYFRQLILH 307
>TC204257 UP|Q76LA8 (Q76LA8) Cytosolic ascorbate peroxidase 1 , complete
Length = 1102
Score = 26.2 bits (56), Expect = 6.4
Identities = 11/26 (42%), Positives = 15/26 (57%)
Frame = +2
Query: 47 SCTTGLLWLTRAMDFLVAVFRNLIEH 72
SCT G W+ ++D + FR LI H
Sbjct: 575 SCTQGAFWI*GSLDL*SSYFRQLILH 652
>TC205710 similar to UP|Q8L5X1 (Q8L5X1) Fimbrin-like protein (Actin binding
motif), partial (11%)
Length = 714
Score = 25.8 bits (55), Expect = 8.4
Identities = 12/33 (36%), Positives = 21/33 (63%)
Frame = +2
Query: 43 KASSSCTTGLLWLTRAMDFLVAVFRNLIEHADW 75
++SSS +G ++ + FLV VF + EH+D+
Sbjct: 104 QSSSSSFSGTFDMSNPITFLVRVFEFVSEHSDF 202
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.327 0.134 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,450,111
Number of Sequences: 63676
Number of extensions: 94712
Number of successful extensions: 710
Number of sequences better than 10.0: 39
Number of HSP's better than 10.0 without gapping: 708
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 710
length of query: 152
length of database: 12,639,632
effective HSP length: 89
effective length of query: 63
effective length of database: 6,972,468
effective search space: 439265484
effective search space used: 439265484
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 54 (25.4 bits)
Medicago: description of AC144481.4


