

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
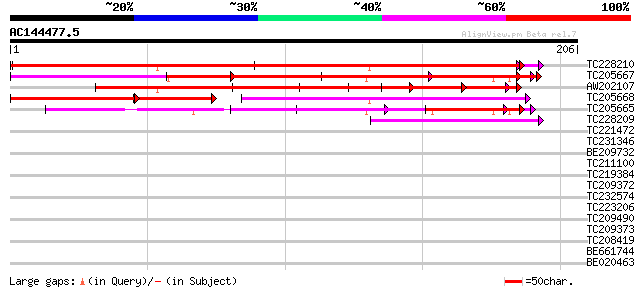
Query= AC144477.5 - phase: 1 /pseudo
(206 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC228210 192 9e-50
TC205667 181 3e-48
AW202107 122 9e-29
TC205668 weakly similar to UP|Q6K620 (Q6K620) Leucine-rich repea... 83 2e-20
TC205665 68 3e-12
TC228209 41 3e-04
TC221472 35 0.024
TC231346 weakly similar to UP|Q6QM04 (Q6QM04) LRR protein WM1.2,... 34 0.041
BE209732 34 0.053
TC211100 similar to GB|AAM91352.1|22137014|AY133522 At5g23340/MK... 32 0.26
TC219384 30 1.0
TC209372 weakly similar to UP|Q96477 (Q96477) LRR protein, parti... 29 1.3
TC232574 weakly similar to UP|Q6QM04 (Q6QM04) LRR protein WM1.2,... 29 1.3
TC223206 similar to UP|Q38695 (Q38695) Polygalacturonase inhibit... 29 1.3
TC209490 UP|Q8GSN9 (Q8GSN9) LRR receptor-like kinase (Fragment),... 28 2.2
TC209373 weakly similar to UP|Q96477 (Q96477) LRR protein, parti... 28 2.2
TC208419 weakly similar to UP|Q6X1D9 (Q6X1D9) Resistance protein... 28 2.2
BE661744 28 2.2
BE020463 28 3.8
>TC228210
Length = 1232
Score = 192 bits (488), Expect = 9e-50
Identities = 107/191 (56%), Positives = 141/191 (73%), Gaps = 4/191 (2%)
Frame = +1
Query: 1 LQKLSTLNVEGCSITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHM----FPFL 56
LQKL+ LN+EGC +TAAC + ++ L AL+ LNLNRC LSD+G +KFS L+++ F
Sbjct: 10 LQKLALLNLEGCLVTAACLDSLAELPALSNLNLNRCLLSDNGCKKFSRLENLKILNLGFN 189
Query: 57 QVLQA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISG 116
+ A + LT+L++ KIGDEGLVNL G L L LSDTEVG++G+ ++SG
Sbjct: 190 VITDACLVHLKGLTKLESLNLDSCKIGDEGLVNLAGHKQLNCLELSDTEVGSNGLHHLSG 369
Query: 117 LNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGA 176
L+ L+ +NLSFT ++D+ L++L GL++LKSLNLDA QITDAGLANLTSL+GL LDLFGA
Sbjct: 370 LSSLQKINLSFTMISDSSLRKLCGLSSLKSLNLDAYQITDAGLANLTSLTGLTDLDLFGA 549
Query: 177 RITDSGTTYLR 187
RITD GT YL+
Sbjct: 550 RITDFGTNYLK 582
Score = 67.4 bits (163), Expect = 4e-12
Identities = 58/198 (29%), Positives = 97/198 (48%), Gaps = 5/198 (2%)
Frame = +1
Query: 2 QKLSTLNVEGCSITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHM----FPFLQ 57
++L+ L + + + ++S L++L +NL+ +SD K L + Q
Sbjct: 301 KQLNCLELSDTEVGSNGLHHLSGLSSLQKINLSFTMISDSSLRKLCGLSSLKSLNLDAYQ 480
Query: 58 VLQA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGL 117
+ A + LT L F +I D G L L+SL + + ++G++ I L
Sbjct: 481 ITDAGLANLTSLTGLTDLDLFGARITDFGTNYLKKFKNLRSLEICGGGLTDTGVKNIKEL 660
Query: 118 NKLEDLNLSFTS-VTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGA 176
+ L LNLS S +TD L+ + GLT L SLN+ +IT+AGL +L +L L +L L
Sbjct: 661 SSLVCLNLSQNSNLTDKTLELISGLTGLVSLNVSNSRITNAGLQHLKTLKNLRSLTLESC 840
Query: 177 RITDSGTTYLRCMYIYQL 194
++T + L+ MY+ L
Sbjct: 841 KVTANDIKKLKSMYLPNL 894
Score = 51.6 bits (122), Expect = 2e-07
Identities = 37/97 (38%), Positives = 51/97 (52%)
Frame = +1
Query: 90 LTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNL 149
L GL L L L V + + ++ L L +LNL+ ++DNG K+ L NLK LNL
Sbjct: 1 LKGLQKLALLNLEGCLVTAACLDSLAELPALSNLNLNRCLLSDNGCKKFSRLENLKILNL 180
Query: 150 DARQITDAGLANLTSLSGLITLDLFGARITDSGTTYL 186
ITDA L +L L+ L +L+L +I D G L
Sbjct: 181 GFNVITDACLVHLKGLTKLESLNLDSCKIGDEGLVNL 291
>TC205667
Length = 633
Score = 181 bits (460), Expect(2) = 3e-48
Identities = 93/113 (82%), Positives = 103/113 (90%)
Frame = +1
Query: 81 KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLG 140
+IGD GL NLTGLTLLKSLVLSDT++GNSG+R+ISGL KLEDLN+SFT+VTDNGLKRL G
Sbjct: 121 RIGDGGLANLTGLTLLKSLVLSDTDIGNSGLRHISGLKKLEDLNVSFTTVTDNGLKRLSG 300
Query: 141 LTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGARITDSGTTYLRCMYIYQ 193
LT LKSLNLDARQITDAGLANLTSLSGLI LDLFGARI+D+GTT+LR I Q
Sbjct: 301 LTQLKSLNLDARQITDAGLANLTSLSGLIALDLFGARISDNGTTFLRSFKILQ 459
Score = 66.2 bits (160), Expect = 1e-11
Identities = 57/202 (28%), Positives = 95/202 (46%), Gaps = 11/202 (5%)
Frame = +1
Query: 1 LQKLSTLNVEGCSITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHMFPFL---- 56
L+ L L++ IT AC ++ L L LNL+ C + D G + L + +
Sbjct: 13 LKNLKRLSLAFNRITDACLVHLKDLTNLEYLNLDSCRIGDGGLANLTGLTLLKSLVLSDT 192
Query: 57 QVLQA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISG 116
+ + + L +L+ + D GL L+GLT LKSL L ++ ++G+ ++
Sbjct: 193 DIGNSGLRHISGLKKLEDLNVSFTTVTDNGLKRLSGLTQLKSLNLDARQITDAGLANLTS 372
Query: 117 LNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLF-G 175
L+ L L+L ++DNG L L+SL + +TDAG+ N+ + L L+L
Sbjct: 373 LSGLIALDLFGARISDNGTTFLRSFKILQSLEICGGGLTDAGVKNIREIVSLTQLNLSQN 552
Query: 176 ARITD------SGTTYLRCMYI 191
+TD SG T LR + +
Sbjct: 553 CNLTDKTLELISGMTALRSLNV 618
Score = 58.5 bits (140), Expect = 2e-09
Identities = 35/73 (47%), Positives = 47/73 (63%)
Frame = +1
Query: 114 ISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDL 173
ISGL L+ L+L+F +TD L L LTNL+ LNLD+ +I D GLANLT L+ L +L L
Sbjct: 4 ISGLKNLKRLSLAFNRITDACLVHLKDLTNLEYLNLDSCRIGDGGLANLTGLTLLKSLVL 183
Query: 174 FGARITDSGTTYL 186
I +SG ++
Sbjct: 184 SDTDIGNSGLRHI 222
Score = 47.8 bits (112), Expect = 4e-06
Identities = 42/155 (27%), Positives = 71/155 (45%), Gaps = 1/155 (0%)
Frame = +1
Query: 1 LQKLSTLNVEGCSITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHMFPFLQVLQ 60
L+KL LNV ++T + +S L L LNL+ ++D G + L + L +
Sbjct: 229 LKKLEDLNVSFTTVTDNGLKRLSGLTQLKSLNLDARQITDAGLANLTSLSGLIA-LDLFG 405
Query: 61 A*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKL 120
A +I D G L +L+SL + + ++G++ I + L
Sbjct: 406 A-------------------RISDNGTTFLRSFKILQSLEICGGGLTDAGVKNIREIVSL 528
Query: 121 EDLNLSFT-SVTDNGLKRLLGLTNLKSLNLDARQI 154
LNLS ++TD L+ + G+T L+SLN+ +I
Sbjct: 529 TQLNLSQNCNLTDKTLELISGMTALRSLNVSNSRI 633
Score = 37.4 bits (85), Expect = 0.005
Identities = 22/50 (44%), Positives = 29/50 (58%)
Frame = +1
Query: 137 RLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGARITDSGTTYL 186
++ GL NLK L+L +ITDA L +L L+ L L+L RI D G L
Sbjct: 1 KISGLKNLKRLSLAFNRITDACLVHLKDLTNLEYLNLDSCRIGDGGLANL 150
Score = 27.3 bits (59), Expect(2) = 3e-48
Identities = 17/25 (68%), Positives = 18/25 (72%)
Frame = +2
Query: 58 VLQA*RG*VLHLTRLQTHA*FI*KI 82
VL+ *RG* LT Q HA*FI*KI
Sbjct: 11 VLKT*RG*AWLLTGSQMHA*FI*KI 85
>AW202107
Length = 420
Score = 122 bits (307), Expect = 9e-29
Identities = 70/120 (58%), Positives = 87/120 (72%), Gaps = 4/120 (3%)
Frame = +2
Query: 32 NLNRCGLSDDGFEKFSVLQHM----FPFLQVLQA*RG*VLHLTRLQTHA*FI*KIGDEGL 87
+LNRCGLSDDGFEK S L+++ F ++ A + LT L+ +IGD+GL
Sbjct: 5 DLNRCGLSDDGFEKISGLKNLKRLSLAFNRITDACLVHLKGLTNLEYLNLDYCRIGDDGL 184
Query: 88 VNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSL 147
NLTGLTLLKSLVLSDT++GNSG+R+ISGL KLEDLNLSFT+VTD+GLK G T+L L
Sbjct: 185 ANLTGLTLLKSLVLSDTDIGNSGLRHISGLKKLEDLNLSFTTVTDHGLKGCQGWTHLNHL 364
Score = 65.1 bits (157), Expect = 2e-11
Identities = 37/101 (36%), Positives = 58/101 (56%)
Frame = +2
Query: 82 IGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGL 141
+ D+G ++GL LK L L+ + ++ + ++ GL LE LNL + + D+GL L GL
Sbjct: 23 LSDDGFEKISGLKNLKRLSLAFNRITDACLVHLKGLTNLEYLNLDYCRIGDDGLANLTGL 202
Query: 142 TNLKSLNLDARQITDAGLANLTSLSGLITLDLFGARITDSG 182
T LKSL L I ++GL +++ L L L+L +TD G
Sbjct: 203 TLLKSLVLSDTDIGNSGLRHISGLKKLEDLNLSFTTVTDHG 325
Score = 61.6 bits (148), Expect = 2e-10
Identities = 37/81 (45%), Positives = 50/81 (61%)
Frame = +2
Query: 106 VGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSL 165
+ + G ISGL L+ L+L+F +TD L L GLTNL+ LNLD +I D GLANLT L
Sbjct: 23 LSDDGFEKISGLKNLKRLSLAFNRITDACLVHLKGLTNLEYLNLDYCRIGDDGLANLTGL 202
Query: 166 SGLITLDLFGARITDSGTTYL 186
+ L +L L I +SG ++
Sbjct: 203 TLLKSLVLSDTDIGNSGLRHI 265
Score = 48.5 bits (114), Expect = 2e-06
Identities = 25/31 (80%), Positives = 27/31 (86%)
Frame = +1
Query: 136 KRLLGLTNLKSLNLDARQITDAGLANLTSLS 166
+RL + LKSLNLDARQITDAGLANLTSLS
Sbjct: 328 ERLSRMDTLKSLNLDARQITDAGLANLTSLS 420
Score = 43.1 bits (100), Expect = 9e-05
Identities = 25/63 (39%), Positives = 38/63 (59%)
Frame = +2
Query: 124 NLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGARITDSGT 183
+L+ ++D+G +++ GL NLK L+L +ITDA L +L L+ L L+L RI D G
Sbjct: 5 DLNRCGLSDDGFEKISGLKNLKRLSLAFNRITDACLVHLKGLTNLEYLNLDYCRIGDDGL 184
Query: 184 TYL 186
L
Sbjct: 185 ANL 193
>TC205668 weakly similar to UP|Q6K620 (Q6K620) Leucine-rich repeat-like
protein, partial (40%)
Length = 1052
Score = 83.2 bits (204), Expect(2) = 2e-20
Identities = 38/47 (80%), Positives = 43/47 (90%)
Frame = +3
Query: 1 LQKLSTLNVEGCSITAACFEYISALAALACLNLNRCGLSDDGFEKFS 47
L+KL+TLNVEGC+ITAAC E+I AL +LACLNLNRCGLSDDGFEK S
Sbjct: 732 LKKLTTLNVEGCNITAACLEFIHALTSLACLNLNRCGLSDDGFEKIS 872
Score = 52.4 bits (124), Expect = 1e-07
Identities = 34/106 (32%), Positives = 53/106 (49%), Gaps = 1/106 (0%)
Frame = +3
Query: 85 EGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTS-VTDNGLKRLLGLTN 143
+G+ + L L+ L L + G ++ GL KLE LN+ VTD+ +K + L N
Sbjct: 489 DGMRAFSNLFNLEKLDLERCSEIHGGFVHLKGLKKLEYLNIGCCKCVTDSDIKSISELIN 668
Query: 144 LKSLNLDARQITDAGLANLTSLSGLITLDLFGARITDSGTTYLRCM 189
LK L + ITD G+ L L L TL++ G IT + ++ +
Sbjct: 669 LKELQISNSSITDIGITYLRGLKKLTTLNVEGCNITAACLEFIHAL 806
Score = 35.4 bits (80), Expect = 0.018
Identities = 34/141 (24%), Positives = 60/141 (42%), Gaps = 1/141 (0%)
Frame = +3
Query: 1 LQKLSTLNVEGCSITAACFEYISALAALACLNLNRCG-LSDDGFEKFSVLQHMFPFLQVL 59
L L L++E CS F ++ L L LN+ C ++D + S L ++ LQ+
Sbjct: 513 LFNLEKLDLERCSEIHGGFVHLKGLKKLEYLNIGCCKCVTDSDIKSISELINLKE-LQIS 689
Query: 60 QA*RG*VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNK 119
+ I D G+ L GL L +L + + + + +I L
Sbjct: 690 NS-------------------SITDIGITYLRGLKKLTTLNVEGCNITAACLEFIHALTS 812
Query: 120 LEDLNLSFTSVTDNGLKRLLG 140
L LNL+ ++D+G +++ G
Sbjct: 813 LACLNLNRCGLSDDGFEKISG 875
Score = 32.3 bits (72), Expect(2) = 2e-20
Identities = 18/30 (60%), Positives = 20/30 (66%)
Frame = +2
Query: 46 FSVLQHMFPFLQVLQA*RG*VLHLTRLQTH 75
F VL H PFLQ L+ *+G* LTR QTH
Sbjct: 959 FLVL*HALPFLQDLKT*KG*A*LLTRSQTH 1048
Score = 29.3 bits (64), Expect = 1.3
Identities = 23/81 (28%), Positives = 43/81 (52%), Gaps = 1/81 (1%)
Frame = +3
Query: 87 LVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTS-VTDNGLKRLLGLTNLK 145
+++ GL+LL S+ ++ ++V + G+R + + L+ L LS+ ++ GLK +
Sbjct: 276 VISSQGLSLL-SVDVAGSQVTDDGLRLLKDCSSLQALTLSYCDQFSEYGLKHI------- 431
Query: 146 SLNLDARQITDAGLANLTSLS 166
+GL+NLTSLS
Sbjct: 432 -----------SGLSNLTSLS 461
>TC205665
Length = 796
Score = 67.8 bits (164), Expect = 3e-12
Identities = 33/36 (91%), Positives = 36/36 (99%)
Frame = +1
Query: 152 RQITDAGLANLTSLSGLITLDLFGARITDSGTTYLR 187
RQITDAGLANLTSLSGLITLDLFGARI+D+GTT+LR
Sbjct: 34 RQITDAGLANLTSLSGLITLDLFGARISDNGTTFLR 141
Score = 58.2 bits (139), Expect = 3e-09
Identities = 33/108 (30%), Positives = 59/108 (54%), Gaps = 1/108 (0%)
Frame = +1
Query: 81 KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLG 140
+I D GL NLT L+ L +L L + ++G ++ L+ L + +TD G+K +
Sbjct: 37 QITDAGLANLTSLSGLITLDLFGARISDNGTTFLRSFKNLQSLEICGGGLTDAGVKNIRE 216
Query: 141 LTNLKSLNLDAR-QITDAGLANLTSLSGLITLDLFGARITDSGTTYLR 187
+ +L LNL +TD L ++ ++ L +L++ +RIT+ G YL+
Sbjct: 217 IVSLTQLNLSQNCNLTDKTLELISGMTALRSLNVSNSRITNEGLRYLK 360
Score = 52.4 bits (124), Expect = 1e-07
Identities = 35/102 (34%), Positives = 55/102 (53%), Gaps = 1/102 (0%)
Frame = +1
Query: 81 KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFT-SVTDNGLKRLL 139
+I D G L L+SL + + ++G++ I + L LNLS ++TD L+ +
Sbjct: 109 RISDNGTTFLRSFKNLQSLEICGGGLTDAGVKNIREIVSLTQLNLSQNCNLTDKTLELIS 288
Query: 140 GLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGARITDS 181
G+T L+SLN+ +IT+ GL L L L TL L ++T S
Sbjct: 289 GMTALRSLNVSNSRITNEGLRYLKPLKNLRTLTLESCKVTAS 414
Score = 42.4 bits (98), Expect = 1e-04
Identities = 38/136 (27%), Positives = 65/136 (46%), Gaps = 11/136 (8%)
Frame = +1
Query: 14 ITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHMFPFLQVLQA*RG*-------- 65
IT A +++L+ L L+L +SD+G + F LQ L+ G
Sbjct: 40 ITDAGLANLTSLSGLITLDLFGARISDNG----TTFLRSFKNLQSLEICGGGLTDAGVKN 207
Query: 66 ---VLHLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLED 122
++ LT+L + D+ L ++G+T L+SL +S++ + N G+RY+ L L
Sbjct: 208 IREIVSLTQLNLSQNC--NLTDKTLELISGMTALRSLNVSNSRITNEGLRYLKPLKNLRT 381
Query: 123 LNLSFTSVTDNGLKRL 138
L L VT + +K+L
Sbjct: 382 LTLESCKVTASEIKKL 429
Score = 42.0 bits (97), Expect = 2e-04
Identities = 28/94 (29%), Positives = 49/94 (51%), Gaps = 7/94 (7%)
Frame = +1
Query: 105 EVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTS 164
++ ++G+ ++ L+ L L+L ++DNG L NL+SL + +TDAG+ N+
Sbjct: 37 QITDAGLANLTSLSGLITLDLFGARISDNGTTFLRSFKNLQSLEICGGGLTDAGVKNIRE 216
Query: 165 LSGLITLDLF-GARITD------SGTTYLRCMYI 191
+ L L+L +TD SG T LR + +
Sbjct: 217 IVSLTQLNLSQNCNLTDKTLELISGMTALRSLNV 318
>TC228209
Length = 466
Score = 41.2 bits (95), Expect = 3e-04
Identities = 23/63 (36%), Positives = 37/63 (58%)
Frame = +3
Query: 132 DNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGARITDSGTTYLRCMYI 191
D L+ + GLT L SLN+ +IT+AGL +L +L L +L L ++T + L+ +Y+
Sbjct: 12 DKTLELISGLTGLVSLNVSNSRITNAGLQHLKTLKNLRSLTLESCKVTANDIKKLKSIYL 191
Query: 192 YQL 194
L
Sbjct: 192 PNL 200
Score = 39.3 bits (90), Expect = 0.001
Identities = 24/65 (36%), Positives = 39/65 (59%), Gaps = 2/65 (3%)
Frame = +3
Query: 84 DEGLVNLTGLTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRL--LGL 141
D+ L ++GLT L SL +S++ + N+G++++ L L L L VT N +K+L + L
Sbjct: 12 DKTLELISGLTGLVSLNVSNSRITNAGLQHLKTLKNLRSLTLESCKVTANDIKKLKSIYL 191
Query: 142 TNLKS 146
NL S
Sbjct: 192 PNLVS 206
>TC221472
Length = 640
Score = 35.0 bits (79), Expect = 0.024
Identities = 28/94 (29%), Positives = 47/94 (49%), Gaps = 5/94 (5%)
Frame = +2
Query: 101 LSDTEVGNSGIR---YISGLNKLEDLNLSFTSVTDNGLKRLLGLT-NLKSLNLDARQITD 156
LS +V NS ++S + +E LNLS + D+ ++ + NLKSLNL +++
Sbjct: 140 LSFLDVANSSFHRFFFLSKMKVIEHLNLSSCMMGDDSVEMVACAGGNLKSLNLSGTRVSS 319
Query: 157 AGLANLTS-LSGLITLDLFGARITDSGTTYLRCM 189
AGL L + L L L + D+ +++ M
Sbjct: 320 AGLGILAGHVPHLEILSLSQTPVDDTAISFISMM 421
Score = 31.6 bits (70), Expect = 0.26
Identities = 42/168 (25%), Positives = 70/168 (41%), Gaps = 17/168 (10%)
Frame = +2
Query: 4 LSTLNVEGCSITAACFEYISALAALACLNLNRCGLSDDGFEKFSVLQHMFPFL-----QV 58
LS L+V S F ++S + + LNL+ C + DD E + L +V
Sbjct: 140 LSFLDVANSSFHR--FFFLSKMKVIEHLNLSSCMMGDDSVEMVACAGGNLKSLNLSGTRV 313
Query: 59 LQA*RG*VL-HLTRLQTHA*FI*KIGDEGLVNLTGLTLLKSLVLSDTEV------GNSGI 111
A G + H+ L+ + + D + ++ + LK + LS+T + G + +
Sbjct: 314 SSAGLGILAGHVPHLEILSLSQTPVDDTAISFISMMPSLKDVDLSNTNIKGFLHQGRTDV 493
Query: 112 RYISGLN-----KLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQI 154
+ L KLE LNL T V D L L L+ L+L + +
Sbjct: 494 NSLLSLMALQNLKLERLNLEHTQVRDEALYPLSSFQELRYLSLKSASL 637
>TC231346 weakly similar to UP|Q6QM04 (Q6QM04) LRR protein WM1.2, partial
(14%)
Length = 1790
Score = 34.3 bits (77), Expect = 0.041
Identities = 22/59 (37%), Positives = 30/59 (50%)
Frame = +1
Query: 115 SGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDL 173
+ L+ L LNL+ + K L NL+ LNL A +T L +LS L+TLDL
Sbjct: 190 ANLSSLRTLNLAHNRLNGTIPKSFEFLKNLQVLNLGANSLTGDVPVTLGTLSNLVTLDL 366
>BE209732
Length = 441
Score = 33.9 bits (76), Expect = 0.053
Identities = 20/55 (36%), Positives = 34/55 (61%), Gaps = 1/55 (1%)
Frame = +3
Query: 120 LEDLNLSFTSVTDNGLKRLLGLTNLKSLNLD-ARQITDAGLANLTSLSGLITLDL 173
L +++S + VTDNGL L +NL++L L+ Q ++ GL ++ LS L +L +
Sbjct: 12 LLSVDVSGSQVTDNGLPFLKDCSNLQALTLNFGDQFSEYGLKHIIGLSNLTSLSI 176
>TC211100 similar to GB|AAM91352.1|22137014|AY133522 At5g23340/MKD15_20
{Arabidopsis thaliana;} , partial (61%)
Length = 815
Score = 31.6 bits (70), Expect = 0.26
Identities = 44/161 (27%), Positives = 72/161 (44%), Gaps = 10/161 (6%)
Frame = +1
Query: 23 SALAALACLNLNRC-GLSDDGFEKFSVLQHMFPFLQVLQA*RG*VLHLTRLQT-HA*FI* 80
+A L LNL+ C G++D G + HL+ LQ+ +
Sbjct: 343 TAFTCLKILNLHNCKGITDAGMKAIGE-------------------HLSLLQSLDVSYCR 465
Query: 81 KIGDEGLVNLT-GLTLLKSLVLSDTEVGNSGIRYISGLN--KLEDLNL-SFTSVTDNGLK 136
K+ D+GL + G L+ L ++ G+ N LE+L L TS+TDNGL
Sbjct: 466 KLTDKGLSAVAKGCCDLRILHMAGCRFVTDGVLEALSKNCGNLEELGLHGCTSITDNGLI 645
Query: 137 RLL-GLTNLKSLNLD-ARQITDAGLANLTSL--SGLITLDL 173
L G ++ L+++ +TD G+++++ S L TL L
Sbjct: 646 NLASGCRRIRFLDINKCSNVTDVGVSSVSRACSSSLKTLKL 768
>TC219384
Length = 1313
Score = 29.6 bits (65), Expect = 1.0
Identities = 32/110 (29%), Positives = 48/110 (43%), Gaps = 30/110 (27%)
Frame = +2
Query: 81 KIGDEGLVNLTGLTLLKSLVLS---------------------------DTEVGNSGIRY 113
+I DEGL++++ + L SL + T V + GI
Sbjct: 146 EIDDEGLMSISSCSRLSSLKIGICLNITDRGLTYVGMHCSKLKELDLYRSTGVDDLGISA 325
Query: 114 IS-GLNKLEDLNLSF-TSVTDNGLKRLLGLTNLKSLNL-DARQITDAGLA 160
I+ G LE +N S+ TS+TD L L +NLK+L + +T GLA
Sbjct: 326 IARGCPGLEMINTSYCTSITDRALITLSKCSNLKTLEIRGCLLVTSIGLA 475
Score = 29.3 bits (64), Expect = 1.3
Identities = 19/52 (36%), Positives = 28/52 (53%), Gaps = 1/52 (1%)
Frame = +2
Query: 120 LEDLNLSFTSVTDNGLKRLLGLTNLKSLN-LDARQITDAGLANLTSLSGLIT 170
L +NLS++SVTD GL L ++ L+S L + + GLA G +T
Sbjct: 578 LRQINLSYSSVTDVGLLSLANISCLQSFTLLHLQGLVPGGLAAALLACGGLT 733
>TC209372 weakly similar to UP|Q96477 (Q96477) LRR protein, partial (71%)
Length = 690
Score = 29.3 bits (64), Expect = 1.3
Identities = 21/69 (30%), Positives = 32/69 (45%)
Frame = +2
Query: 117 LNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGA 176
L L L L ++T + L L+N+K L L++ ++T LT L L LDL
Sbjct: 365 LKNLLSLGLYQNNLTGSIPATLSNLSNIKFLRLNSNKLTGRIPRELTKLGNLKILDLSNN 544
Query: 177 RITDSGTTY 185
+ + TY
Sbjct: 545 DLCGTXPTY 571
>TC232574 weakly similar to UP|Q6QM04 (Q6QM04) LRR protein WM1.2, partial
(10%)
Length = 852
Score = 29.3 bits (64), Expect = 1.3
Identities = 23/78 (29%), Positives = 39/78 (49%)
Frame = +3
Query: 96 LKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQIT 155
LKS+ LS + + + L L LNLS +++ R+ L +L+SL+L I+
Sbjct: 27 LKSIDLSSNNLTGEIPKEVGYLLGLVSLNLSRNNLSGEIPSRIGNLRSLESLDLSRNHIS 206
Query: 156 DAGLANLTSLSGLITLDL 173
++L+ + L LDL
Sbjct: 207 RRIPSSLSEIDDLQKLDL 260
>TC223206 similar to UP|Q38695 (Q38695) Polygalacturonase inhibitor
precursor, partial (9%)
Length = 458
Score = 29.3 bits (64), Expect = 1.3
Identities = 15/35 (42%), Positives = 19/35 (53%)
Frame = +1
Query: 1 LQKLSTLNVEGCSITAACFEYISALAALACLNLNR 35
LQ LS L++ GC E + L L CLNL+R
Sbjct: 349 LQALSRLDISGCDSLLRVPEGLQNLKKLQCLNLSR 453
>TC209490 UP|Q8GSN9 (Q8GSN9) LRR receptor-like kinase (Fragment), complete
Length = 3298
Score = 28.5 bits (62), Expect = 2.2
Identities = 25/86 (29%), Positives = 41/86 (47%), Gaps = 1/86 (1%)
Frame = +1
Query: 89 NLTGLTLLKSLVLSDTEVGNSGIRYISG-LNKLEDLNLSFTSVTDNGLKRLLGLTNLKSL 147
+L+ L L+ L L GI G + L L+LS +++ L LTNL +L
Sbjct: 727 SLSKLKTLRYLKLGYNNAYEGGIPPEFGSMKSLRYLDLSSCNLSGEIPPSLANLTNLDTL 906
Query: 148 NLDARQITDAGLANLTSLSGLITLDL 173
L +T + L+++ L++LDL
Sbjct: 907 FLQINNLTGTIPSELSAMVSLMSLDL 984
>TC209373 weakly similar to UP|Q96477 (Q96477) LRR protein, partial (51%)
Length = 654
Score = 28.5 bits (62), Expect = 2.2
Identities = 21/69 (30%), Positives = 32/69 (45%)
Frame = +1
Query: 117 LNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQITDAGLANLTSLSGLITLDLFGA 176
L L L L ++T + L L+N+K L L++ ++T LT L L LDL
Sbjct: 178 LKNLLSLGLYQNNLTGSIPATLSNLSNIKFLRLNSNKLTGRIPRELTKLGNLKILDLSNN 357
Query: 177 RITDSGTTY 185
+ + TY
Sbjct: 358 DLCGTFPTY 384
>TC208419 weakly similar to UP|Q6X1D9 (Q6X1D9) Resistance protein SlVe1
precursor, partial (9%)
Length = 1140
Score = 28.5 bits (62), Expect = 2.2
Identities = 23/79 (29%), Positives = 40/79 (50%)
Frame = +1
Query: 95 LLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDARQI 154
LLK+L S + + L +L LNLS ++ ++ G+ NL+SL+L +
Sbjct: 367 LLKNLDFSTNNLSGEIPPELFSLTELLFLNLSRNNLMGKIPSKIGGMKNLESLDLSNNHL 546
Query: 155 TDAGLANLTSLSGLITLDL 173
+ A +++LS L L+L
Sbjct: 547 SGEIPAAISNLSFLSYLNL 603
>BE661744
Length = 743
Score = 28.5 bits (62), Expect = 2.2
Identities = 15/28 (53%), Positives = 19/28 (67%)
Frame = +3
Query: 1 LQKLSTLNVEGCSITAACFEYISALAAL 28
L+KL+ LNVEG +ITA I AL +L
Sbjct: 657 LEKLTXLNVEGWNITADGLXLIHALTSL 740
Score = 27.7 bits (60), Expect = 3.8
Identities = 19/46 (41%), Positives = 29/46 (62%), Gaps = 7/46 (15%)
Frame = +3
Query: 128 TSVTDNGLKRLLGLTNLKSLNLD-ARQITD------AGLANLTSLS 166
T VTD+GL+ L ++L++L L Q ++ +GL+NLTSLS
Sbjct: 66 TWVTDDGLRLLKDCSSLQALTLSYCDQFSEYGLKHISGLSNLTSLS 203
>BE020463
Length = 422
Score = 27.7 bits (60), Expect = 3.8
Identities = 19/73 (26%), Positives = 38/73 (52%)
Frame = +2
Query: 93 LTLLKSLVLSDTEVGNSGIRYISGLNKLEDLNLSFTSVTDNGLKRLLGLTNLKSLNLDAR 152
L L SL L + + S + L DLNL++ S++ N + + +++L SLN+
Sbjct: 80 LKQLSSLHLEENSLTGSIPAELGHCAMLVDLNLAWNSLSGNIPQSVSLMSSLNSLNISGN 259
Query: 153 QITDAGLANLTSL 165
+++ + NL ++
Sbjct: 260 KLSGSIPENLEAI 298
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.335 0.148 0.437
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,999,426
Number of Sequences: 63676
Number of extensions: 92672
Number of successful extensions: 768
Number of sequences better than 10.0: 44
Number of HSP's better than 10.0 without gapping: 715
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 753
length of query: 206
length of database: 12,639,632
effective HSP length: 93
effective length of query: 113
effective length of database: 6,717,764
effective search space: 759107332
effective search space used: 759107332
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 56 (26.2 bits)
Medicago: description of AC144477.5


