

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
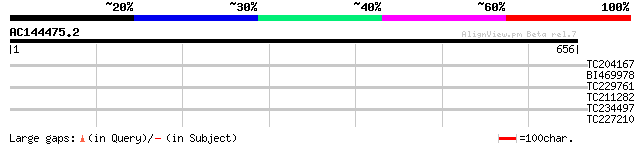
Query= AC144475.2 - phase: 0 /pseudo
(656 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC204167 weakly similar to UP|O22209 (O22209) Expressed protein,... 35 0.11
BI469978 homologue to GP|27808562|gb| At3g52790 {Arabidopsis tha... 31 1.5
TC229761 similar to UP|Q9C566 (Q9C566) Cyclophilin-40 (Expressed... 30 2.6
TC211282 UP|C7D9_SOYBN (O81971) Cytochrome P450 71D9 (P450 CP3)... 30 3.4
TC234497 similar to UP|Q94G16 (Q94G16) HCF106, partial (40%) 30 4.4
TC227210 similar to UP|Q8VYV8 (Q8VYV8) At1g12570/T12C24_9, parti... 28 9.9
>TC204167 weakly similar to UP|O22209 (O22209) Expressed protein, partial
(56%)
Length = 1770
Score = 35.0 bits (79), Expect = 0.11
Identities = 31/99 (31%), Positives = 52/99 (52%), Gaps = 2/99 (2%)
Frame = -1
Query: 318 FLVLLEIGTLISS*ITCQLTLLIKF-LRFQSLMMMMVQTLLGGVEPIP-TISQFKVLILC 375
F +E+ +SS QL L+K R +S + TLLGG+ +P +++ FK LIL
Sbjct: 963 FFSSVEVWDRVSS--ESQLLELLKMRARARSYSTFLSSTLLGGLLFLPASLASFK-LILL 793
Query: 376 NIKIARFWKVTGKVFGSGMAHIASKPLFGWLRTSAFLQI 414
+ + + +G F + +A I+ K G+L ++ FL I
Sbjct: 792 GLGLMAIFIFSGLSFSNCLAGISDKRTMGYLLSAIFLTI 676
>BI469978 homologue to GP|27808562|gb| At3g52790 {Arabidopsis thaliana},
partial (25%)
Length = 421
Score = 31.2 bits (69), Expect = 1.5
Identities = 13/41 (31%), Positives = 25/41 (60%)
Frame = +2
Query: 70 IKLVYISPRMLIPNFARIFYITQGLIRLIAWACILGLILLL 110
+ LVYI+P++++ N YI + I+W C+L + ++L
Sbjct: 128 LPLVYITPQVIMANGMGFKYI**SMAEKISWCCVLFMAIIL 250
>TC229761 similar to UP|Q9C566 (Q9C566) Cyclophilin-40 (Expressed protein),
partial (23%)
Length = 523
Score = 30.4 bits (67), Expect = 2.6
Identities = 12/43 (27%), Positives = 24/43 (54%)
Frame = -3
Query: 18 FLCAQAGMVHKYLIYFLLMIYFFLQRPQLNRLIVFCIALIFFV 60
++ Q+G +HK Y+ +++Y F R +L+ + A FF+
Sbjct: 296 YMLQQSGCLHKPECYWNILLYAFFSRSRLSAIFFLAAASSFFI 168
>TC211282 UP|C7D9_SOYBN (O81971) Cytochrome P450 71D9 (P450 CP3) , complete
Length = 1573
Score = 30.0 bits (66), Expect = 3.4
Identities = 21/63 (33%), Positives = 34/63 (53%)
Frame = -2
Query: 326 TLISS*ITCQLTLLIKFLRFQSLMMMMVQTLLGGVEPIPTISQFKVLILCNIKIARFWKV 385
TL++S C ++++ FL ++ +MV + P P ISQ LIL ++K F+K
Sbjct: 1014 TLLTS--VCTFSIVLGFLIISAIAHVMVADDVS--LPPPKISQITALILSSLKPNSFFKS 847
Query: 386 TGK 388
T K
Sbjct: 846 TSK 838
>TC234497 similar to UP|Q94G16 (Q94G16) HCF106, partial (40%)
Length = 590
Score = 29.6 bits (65), Expect = 4.4
Identities = 20/71 (28%), Positives = 33/71 (46%), Gaps = 13/71 (18%)
Frame = -3
Query: 348 LMMMMVQTLLGGVEPIPTISQFKVLILCNIKI-------ARFWKVTGKVFGS------GM 394
L + ++ T LG P+ S+ L+LCN+KI +R W+ GS G+
Sbjct: 267 LDLRLIVTFLGRCLPLMLCSRCLQLLLCNLKILLRAVCFSRIWRAIRVNSGSG**SWCGI 88
Query: 395 AHIASKPLFGW 405
A++ + GW
Sbjct: 87 AYVGIVCVLGW 55
>TC227210 similar to UP|Q8VYV8 (Q8VYV8) At1g12570/T12C24_9, partial (72%)
Length = 1574
Score = 28.5 bits (62), Expect = 9.9
Identities = 19/55 (34%), Positives = 27/55 (48%), Gaps = 10/55 (18%)
Frame = -2
Query: 382 FWKVTGKVFGSGMAHIASK----------PLFGWLRTSAFLQISKGVNGVVEYLL 426
FW+ TGKVF +G ++A+K L+G R GV+GVV + L
Sbjct: 1186 FWRTTGKVFTTGSFNVAAKTGDAHHLNQRDLYGDRRRD-----KDGVHGVVSHSL 1037
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.356 0.161 0.537
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 28,470,760
Number of Sequences: 63676
Number of extensions: 412972
Number of successful extensions: 5160
Number of sequences better than 10.0: 13
Number of HSP's better than 10.0 without gapping: 2901
Number of HSP's successfully gapped in prelim test: 194
Number of HSP's that attempted gapping in prelim test: 2133
Number of HSP's gapped (non-prelim): 3253
length of query: 656
length of database: 12,639,632
effective HSP length: 103
effective length of query: 553
effective length of database: 6,081,004
effective search space: 3362795212
effective search space used: 3362795212
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.6 bits)
S2: 62 (28.5 bits)
Medicago: description of AC144475.2


