

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
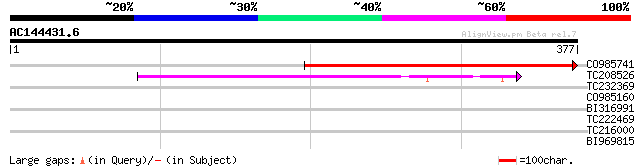
Query= AC144431.6 - phase: 0
(377 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
CO985741 166 2e-41
TC208526 similar to UP|Q6YW97 (Q6YW97) MAP3K protein kinase-like... 62 4e-10
TC232369 weakly similar to UP|Q948Y6 (Q948Y6) VMP4 protein, part... 31 1.0
CO985160 31 1.0
BI316991 weakly similar to GP|28144147|gb| PDZ-LIM protein cyphe... 30 1.8
TC222469 29 3.0
TC216000 similar to UP|O22253 (O22253) Photolyase/blue-light rec... 28 6.8
BI969815 weakly similar to GP|9757860|dbj exonuclease-like prote... 28 8.9
>CO985741
Length = 761
Score = 166 bits (420), Expect = 2e-41
Identities = 103/181 (56%), Positives = 119/181 (64%)
Frame = -2
Query: 197 KAFLSRGNTS*KTNMEMKVCSRVLVRIEMNRHQ*TQNLGH*F**TFFILHRIVLRPVATI 256
+A L G+T *KTNMEM VLV+ N Q*TQ+L H F T FIL +I LRP TI
Sbjct: 760 RALLISGHT**KTNMEMMA*KLVLVQAVQNHPQ*TQSLDHWFWSTTFILLQIDLRPALTI 581
Query: 257 QLHYSAC*RLAMKRPATDGQISLLLIII*GVMVEEYHKLLMQQMVVSLVDVTALLIARQM 316
+LHY *RL K+PAT G ISLLLII GVM EE+H LM+QMV+SL VT L IAR+
Sbjct: 580 RLHYLT**RLVTKQPATVGPISLLLIITRGVMAEEHH*PLMKQMVISLAGVTTLPIARKT 401
Query: 317 VRLVHVMSLQFLHLHLLQKLHLMETRNLLIIVKQAMHIVVEQL**CNWLC*LWLPQLSWH 376
+ L HVMSL FLHLHLLQK ET I + Q M I+VE+ *CN L WL +LSWH
Sbjct: 400 LSLAHVMSLPFLHLHLLQKPPPPETSRPKITITQTMLILVERPS*CNRLLEFWLRRLSWH 221
Query: 377 G 377
G
Sbjct: 220 G 218
>TC208526 similar to UP|Q6YW97 (Q6YW97) MAP3K protein kinase-like protein,
partial (59%)
Length = 1214
Score = 62.0 bits (149), Expect = 4e-10
Identities = 84/262 (32%), Positives = 114/262 (43%), Gaps = 7/262 (2%)
Frame = +3
Query: 86 CMIFKTMYGCVILLEANVSTSVHSYLQ*TRLGTCVHFWTQIHLK*SPSLLRIMLELLQP* 145
C + +YG V N +TS+H Q H +I L+ SPS L+ M L + *
Sbjct: 30 CTTSRMIYGSVTHFAGNATTSLHFNQQLIL*KKWRHS*RKIQLRLSPS*LKTMCTLQRG* 209
Query: 146 LRSSKLQGLTSTCSRLVGCPRTVAIGLLLMT*S*TINDLSHSPRDLPRKQLKAFLSRGNT 205
L S++ S C + V IG L I DL S RKQ K L G+T
Sbjct: 210 LMCSQVLDWISIGFLCPRCRKRVTIGPL*RRWFKQITDLLFSLPMPQRKQGKE*LISGST 389
Query: 206 S*KTNMEMKVCSRVLVRIEMNRHQ*TQNLGH*F**TFFILHRIVLRPVATIQLHYSAC*R 265
K ++E+ C+RVLVR E N+ * + H ** F ++ L +IQLH
Sbjct: 390 WLKMSLEILECNRVLVRTEKNQRH*ILKVIHFS**ITFRHIQLKLTLAKSIQLHL----- 554
Query: 266 LAMKRPATDGQ-----ISLLLIII*GVMVEEYHKLLMQQMVVSLVDVTALLIARQMVRLV 320
L P T Q LLI *GVM + L ++ M + V V Q R V
Sbjct: 555 LRWSIPVTKLQGI*CPTL*LLIFT*GVMEVVFLILWIK*MAIHCVGVA----LSQPARWV 722
Query: 321 HVMSLQ--FLHLHLLQKLHLME 340
+ + L FL+L +Q L L+E
Sbjct: 723 YPLGLARIFLYLVRVQ*LILLE 788
>TC232369 weakly similar to UP|Q948Y6 (Q948Y6) VMP4 protein, partial (3%)
Length = 1311
Score = 30.8 bits (68), Expect = 1.0
Identities = 14/30 (46%), Positives = 20/30 (66%)
Frame = -2
Query: 319 LVHVMSLQFLHLHLLQKLHLMETRNLLIIV 348
L+ LQ LHL +L HL+E RNLL+++
Sbjct: 557 LLQTQILQLLHLFVLVLKHLLELRNLLLLL 468
>CO985160
Length = 777
Score = 30.8 bits (68), Expect = 1.0
Identities = 21/78 (26%), Positives = 45/78 (56%)
Frame = +3
Query: 279 LLLIII*GVMVEEYHKLLMQQMVVSLVDVTALLIARQMVRLVHVMSLQFLHLHLLQKLHL 338
LLL+ + +++ + KLL+ +++ L+ + LL+ +R + ++ LQ L LL L
Sbjct: 411 LLLLDLRKLLLLQLRKLLLLDLLLLLLQLRKLLLLLLQLRKLLLLLLQLRKLLLL----L 578
Query: 339 METRNLLIIVKQAMHIVV 356
+E R LL+++ Q +++
Sbjct: 579 LELRKLLLLLLQLRKLLL 632
>BI316991 weakly similar to GP|28144147|gb| PDZ-LIM protein cypher3s {Mus
musculus}, partial (5%)
Length = 375
Score = 30.0 bits (66), Expect = 1.8
Identities = 12/37 (32%), Positives = 25/37 (67%)
Frame = -3
Query: 317 VRLVHVMSLQFLHLHLLQKLHLMETRNLLIIVKQAMH 353
VR +HV++L+ LH+ ++ LH + +R L ++ + +H
Sbjct: 352 VRGLHVLALRGLHVLTIRGLHALTSRGLHVLALRGLH 242
Score = 28.1 bits (61), Expect = 6.8
Identities = 11/37 (29%), Positives = 24/37 (64%)
Frame = -1
Query: 317 VRLVHVMSLQFLHLHLLQKLHLMETRNLLIIVKQAMH 353
+R +HV++L+ LH+ + LH + +R L ++ + +H
Sbjct: 210 IRGLHVLALRGLHVLTTRGLHALTSRGL*VLALRGLH 100
>TC222469
Length = 438
Score = 29.3 bits (64), Expect = 3.0
Identities = 12/39 (30%), Positives = 26/39 (65%)
Frame = +1
Query: 317 VRLVHVMSLQFLHLHLLQKLHLMETRNLLIIVKQAMHIV 355
VR +HV++L+ LH+ ++ LH++ R L + + +H++
Sbjct: 172 VRGLHVLALRGLHVLAIRGLHVLTFRGLHALAIRGLHVL 288
>TC216000 similar to UP|O22253 (O22253) Photolyase/blue-light receptor
(Photolyase/blue light photoreceptor PHR2), partial
(84%)
Length = 1641
Score = 28.1 bits (61), Expect = 6.8
Identities = 15/47 (31%), Positives = 28/47 (58%), Gaps = 3/47 (6%)
Frame = -3
Query: 303 SLVDVTALLIARQMVRLVHVMSLQFLHLHLL---QKLHLMETRNLLI 346
S++ L + RQ+V ++ S + LHLH+L ++LHL+ L++
Sbjct: 715 SVIGRHVLQLKRQIVHVIQRASPEVLHLHILLLHRRLHLLLRLRLVV 575
>BI969815 weakly similar to GP|9757860|dbj exonuclease-like protein
{Arabidopsis thaliana}, partial (20%)
Length = 726
Score = 27.7 bits (60), Expect = 8.9
Identities = 15/39 (38%), Positives = 21/39 (53%)
Frame = +1
Query: 39 IHG*RHIIHLLWLVQDQPQVQLSLHL*TRMTLSLINSKM 77
+HG RH H + LV DQPQ + + TLSL + +
Sbjct: 484 LHGHRHQEHYIDLVMDQPQNSIVAGQPYQKTLSLFQTAL 600
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.358 0.158 0.540
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,838,410
Number of Sequences: 63676
Number of extensions: 242967
Number of successful extensions: 2682
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 2631
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2675
length of query: 377
length of database: 12,639,632
effective HSP length: 99
effective length of query: 278
effective length of database: 6,335,708
effective search space: 1761326824
effective search space used: 1761326824
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 60 (27.7 bits)
Medicago: description of AC144431.6


