

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144406.5 - phase: 0
(47 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
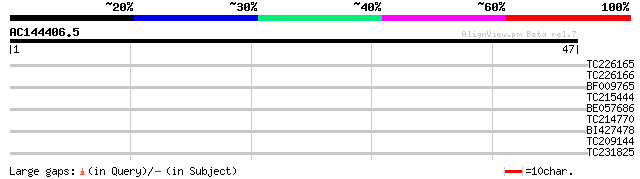
Score E
Sequences producing significant alignments: (bits) Value
TC226165 GAI1 [Glycine max] 32 0.071
TC226166 similar to UP|O23724 (O23724) GAI protein, partial (70%) 31 0.16
BF009765 similar to PIR|E96661|E96 GMP synthase 61700-64653 [im... 27 1.8
TC215444 homologue to UP|Q39013 (Q39013) ATAF1 protein, partial ... 27 2.3
BE057686 27 3.0
TC214770 27 3.0
BI427478 homologue to GP|24078517|gb friend of GATA-1 {Homo sapi... 25 6.7
TC209144 similar to UP|Q8LBT7 (Q8LBT7) Rev interacting protein m... 25 6.7
TC231825 25 6.7
>TC226165 GAI1 [Glycine max]
Length = 1313
Score = 32.0 bits (71), Expect = 0.071
Identities = 24/61 (39%), Positives = 27/61 (43%), Gaps = 17/61 (27%)
Frame = -3
Query: 2 GGRSRC-------PGRTRLLLTLEDHWKRP----------GTRACRERSTPGGRASTVSP 44
GGR C P RTRLL + W+R GTR R S P RAST+ P
Sbjct: 1176 GGRGACETRRRSSPPRTRLLG--DPAWRRRDRRGCLRRSRGTRTARGSSPPAARASTLPP 1003
Query: 45 G 45
G
Sbjct: 1002 G 1000
>TC226166 similar to UP|O23724 (O23724) GAI protein, partial (70%)
Length = 1613
Score = 30.8 bits (68), Expect = 0.16
Identities = 23/61 (37%), Positives = 26/61 (41%), Gaps = 17/61 (27%)
Frame = -2
Query: 2 GGRSRC-------PGRTRLLLTLEDHWKRPGTRACRERS----------TPGGRASTVSP 44
GGR C P RTRLL + W+R R C RS P RAST+ P
Sbjct: 721 GGRGACETRRRSSPPRTRLLG--DRAWRRRDRRGCSRRSHGTRTAR*SPPPAARASTLPP 548
Query: 45 G 45
G
Sbjct: 547 G 545
>BF009765 similar to PIR|E96661|E96 GMP synthase 61700-64653 [imported] -
Arabidopsis thaliana, partial (20%)
Length = 412
Score = 27.3 bits (59), Expect = 1.8
Identities = 18/40 (45%), Positives = 20/40 (50%), Gaps = 2/40 (5%)
Frame = -3
Query: 2 GGRSRCPGRTRLLLTLEDHWKRP--GTRACRERSTPGGRA 39
GGRSRC GR R HW P G+R R S GR+
Sbjct: 275 GGRSRCRGR*R-------HWNGPTGGSRRGRRGSPGCGRS 177
>TC215444 homologue to UP|Q39013 (Q39013) ATAF1 protein, partial (59%)
Length = 1198
Score = 26.9 bits (58), Expect = 2.3
Identities = 16/42 (38%), Positives = 21/42 (49%)
Frame = +1
Query: 5 SRCPGRTRLLLTLEDHWKRPGTRACRERSTPGGRASTVSPGH 46
+RCPG TR W R RA R R++ G A+T + H
Sbjct: 415 TRCPGCTRRTRAAPSRWCRRSLRA-RCRASRRGAATTTTSLH 537
>BE057686
Length = 421
Score = 26.6 bits (57), Expect = 3.0
Identities = 13/35 (37%), Positives = 16/35 (45%)
Frame = +3
Query: 12 RLLLTLEDHWKRPGTRACRERSTPGGRASTVSPGH 46
RLLL H +RP ER+ P AS+ H
Sbjct: 123 RLLLRQAQHQRRPRRARSNERTPPSSHASSARCSH 227
>TC214770
Length = 978
Score = 26.6 bits (57), Expect = 3.0
Identities = 11/23 (47%), Positives = 15/23 (64%)
Frame = -1
Query: 24 PGTRACRERSTPGGRASTVSPGH 46
PGTR+ R R+ PG + +T GH
Sbjct: 264 PGTRSRRWRA*PGAKVTTTRHGH 196
>BI427478 homologue to GP|24078517|gb friend of GATA-1 {Homo sapiens},
partial (1%)
Length = 422
Score = 25.4 bits (54), Expect = 6.7
Identities = 13/40 (32%), Positives = 20/40 (49%), Gaps = 1/40 (2%)
Frame = +2
Query: 4 RSRCPGRTRLLLTLEDHWKRPGTRAC-RERSTPGGRASTV 42
+ R GRTR T WK P + C + ++TP +T+
Sbjct: 50 KRRRSGRTRSTCTSSTPWKPPSSAQCSKTKATPSPPRATL 169
>TC209144 similar to UP|Q8LBT7 (Q8LBT7) Rev interacting protein mis3-like,
partial (45%)
Length = 646
Score = 25.4 bits (54), Expect = 6.7
Identities = 10/16 (62%), Positives = 11/16 (68%)
Frame = -2
Query: 21 WKRPGTRACRERSTPG 36
W+R RACR RST G
Sbjct: 276 WRRS*LRACRRRSTKG 229
>TC231825
Length = 700
Score = 25.4 bits (54), Expect = 6.7
Identities = 13/40 (32%), Positives = 20/40 (49%), Gaps = 1/40 (2%)
Frame = +3
Query: 4 RSRCPGRTRLLLTLEDHWKRPGTRAC-RERSTPGGRASTV 42
+ R GRTR T WK P + C + ++TP +T+
Sbjct: 135 KRRRSGRTRSTCTSSTPWKPPSSAQCSKTKATPSPPRATL 254
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.318 0.135 0.445
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,677,321
Number of Sequences: 63676
Number of extensions: 28514
Number of successful extensions: 206
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 206
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 206
length of query: 47
length of database: 12,639,632
effective HSP length: 23
effective length of query: 24
effective length of database: 11,175,084
effective search space: 268202016
effective search space used: 268202016
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 53 (25.0 bits)
Medicago: description of AC144406.5


