

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144345.5 - phase: 0
(131 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
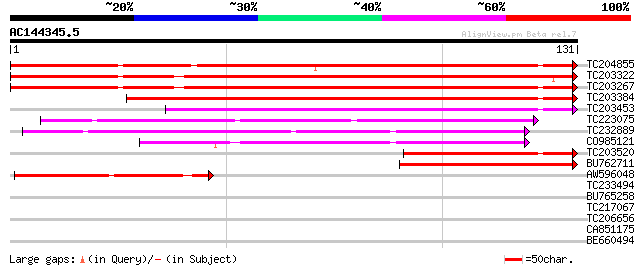
Sequences producing significant alignments: (bits) Value
TC204855 UP|Q93Z89 (Q93Z89) Matrix metalloproteinase MMP2, complete 127 1e-30
TC203322 similar to UP|Q93Z89 (Q93Z89) Matrix metalloproteinase ... 113 3e-26
TC203267 similar to UP|Q93Z89 (Q93Z89) Matrix metalloproteinase ... 113 3e-26
TC203384 similar to UP|Q93Z89 (Q93Z89) Matrix metalloproteinase ... 86 7e-18
TC203453 similar to UP|Q93Z89 (Q93Z89) Matrix metalloproteinase ... 72 7e-14
TC223075 UP|MEP1_SOYBN (P29136) Metalloendoproteinase 1 precurso... 71 1e-13
TC232889 weakly similar to UP|O65340 (O65340) Metalloproteinase ... 71 2e-13
CO985121 54 2e-08
TC203520 similar to UP|Q93Z89 (Q93Z89) Matrix metalloproteinase ... 52 1e-07
BU762711 48 1e-06
AW596048 weakly similar to GP|16901508|gb| matrix metalloprotein... 45 1e-05
TC233494 similar to GB|AAM62930.1|21553837|AY085712 phytochelati... 29 0.73
BU765258 weakly similar to PIR|T14319|T14 protein AX110P - carro... 27 2.1
TC217067 similar to UP|PE21_ARATH (Q42580) Peroxidase 21 precurs... 27 2.8
TC206656 27 3.6
CA851175 26 6.2
BE660494 similar to GP|18252419|gb cytochrome b {Ostrinia nubila... 25 8.0
>TC204855 UP|Q93Z89 (Q93Z89) Matrix metalloproteinase MMP2, complete
Length = 1307
Score = 127 bits (320), Expect = 1e-30
Identities = 67/133 (50%), Positives = 93/133 (69%), Gaps = 2/133 (1%)
Frame = +1
Query: 1 MTNVFRNAFTRWSQTTRVLNFSETTSYDDANIKIGFYNINYIDGVDDVVVGDTVIKLGSN 60
MT VFR++F RW+Q + VLN +ETT YD+A+I++GFYN Y+ G+D V G ++I L +
Sbjct: 592 MTKVFRDSFARWAQASGVLNLTETT-YDNADIQVGFYNFTYL-GIDIEVYGGSLIFLQPD 765
Query: 61 VNSGFIRLI--ASKYWVLPTDNYMWSWQNGEFDLETAAMHQIGHLLGLDHSFDKEYVMYP 118
+ L+ +K W LP++N SW+ G DLE+AAMH+IGHLLGLDHS ++ VMYP
Sbjct: 766 STKKGVILLDGTNKLWALPSENGRLSWEEGVLDLESAAMHEIGHLLGLDHSNKEDSVMYP 945
Query: 119 AILPLQQQRKLQI 131
ILP QRK+Q+
Sbjct: 946 CILP-SHQRKVQL 981
>TC203322 similar to UP|Q93Z89 (Q93Z89) Matrix metalloproteinase MMP2,
partial (74%)
Length = 1664
Score = 113 bits (282), Expect = 3e-26
Identities = 61/132 (46%), Positives = 86/132 (64%), Gaps = 1/132 (0%)
Frame = +2
Query: 1 MTNVFRNAFTRWSQTTRVLNFSETTSYDDANIKIGFYNINYIDGVDDVVVGDTVIKLGSN 60
MT VFR++F RW+Q + L+ ETT YD+A+I++GFYN + + +V G T+ +
Sbjct: 737 MTKVFRDSFARWAQASGTLSLRETT-YDNADIQVGFYN--FTNSRIEVYGGSTIFLQPDS 907
Query: 61 VNSGFIRLIASKYWVLPTDNYMWSWQNGEFDLETAAMHQIGHLLGLDHSFDKEYVMYPAI 120
G + L + W+LP++N S +G DLET AMHQIGHLLGLDHS ++ VMYP I
Sbjct: 908 SKKGVVLLDGNMGWLLPSENATLSKDDGVLDLETVAMHQIGHLLGLDHSHKEDSVMYPYI 1087
Query: 121 LPLQ-QQRKLQI 131
L Q +QRK+Q+
Sbjct: 1088LSSQSEQRKVQL 1123
>TC203267 similar to UP|Q93Z89 (Q93Z89) Matrix metalloproteinase MMP2,
partial (61%)
Length = 1338
Score = 113 bits (282), Expect = 3e-26
Identities = 61/131 (46%), Positives = 86/131 (65%)
Frame = +3
Query: 1 MTNVFRNAFTRWSQTTRVLNFSETTSYDDANIKIGFYNINYIDGVDDVVVGDTVIKLGSN 60
MT VFR++F RW+Q + L+ +ETT YD+A+I++GFYN + + +V G + +
Sbjct: 618 MTKVFRDSFARWAQASGTLSLTETT-YDNADIQVGFYN--FTNSRIEVYGGSLIFLQPDS 788
Query: 61 VNSGFIRLIASKYWVLPTDNYMWSWQNGEFDLETAAMHQIGHLLGLDHSFDKEYVMYPAI 120
G + L + W+LP++N S +G DLETAAMHQIGHLLGLDHS ++ VMYP I
Sbjct: 789 SKKGVVLLNGNMGWLLPSENATLSKDDGVLDLETAAMHQIGHLLGLDHSHKEDSVMYPYI 968
Query: 121 LPLQQQRKLQI 131
L QQRK+Q+
Sbjct: 969 LS-SQQRKVQL 998
>TC203384 similar to UP|Q93Z89 (Q93Z89) Matrix metalloproteinase MMP2,
partial (32%)
Length = 643
Score = 85.5 bits (210), Expect = 7e-18
Identities = 44/104 (42%), Positives = 64/104 (61%)
Frame = +1
Query: 28 DDANIKIGFYNINYIDGVDDVVVGDTVIKLGSNVNSGFIRLIASKYWVLPTDNYMWSWQN 87
D+A+I++GFYN + +V G + + G + + + W+LP++N S +
Sbjct: 1 DNADIQVGFYNFTALSIEVEVYGGSLIFLQPDSSKKGMVLMDGNMGWLLPSENATLSKDD 180
Query: 88 GEFDLETAAMHQIGHLLGLDHSFDKEYVMYPAILPLQQQRKLQI 131
G DLET AMHQIGHLLGLDHS ++ VMYP IL QQRK+++
Sbjct: 181 GVLDLETVAMHQIGHLLGLDHSHKEDSVMYPYILS-SQQRKVKL 309
>TC203453 similar to UP|Q93Z89 (Q93Z89) Matrix metalloproteinase MMP2,
partial (29%)
Length = 622
Score = 72.0 bits (175), Expect = 7e-14
Identities = 39/95 (41%), Positives = 55/95 (57%)
Frame = +1
Query: 37 YNINYIDGVDDVVVGDTVIKLGSNVNSGFIRLIASKYWVLPTDNYMWSWQNGEFDLETAA 96
YN + +V G + + G + + + W+LP++N S +G DLET A
Sbjct: 1 YNFTALRIEVEVYGGSLIFLQPDSSKKGVVLMDGNMGWLLPSENATLSKDDGVLDLETVA 180
Query: 97 MHQIGHLLGLDHSFDKEYVMYPAILPLQQQRKLQI 131
MHQIGHLLGLDHS ++ VMYP IL QQRK+++
Sbjct: 181 MHQIGHLLGLDHSHKEDSVMYPYILS-SQQRKVKL 282
>TC223075 UP|MEP1_SOYBN (P29136) Metalloendoproteinase 1 precursor (SMEP1) ,
complete
Length = 1198
Score = 71.2 bits (173), Expect = 1e-13
Identities = 46/115 (40%), Positives = 61/115 (53%)
Frame = +3
Query: 8 AFTRWSQTTRVLNFSETTSYDDANIKIGFYNINYIDGVDDVVVGDTVIKLGSNVNSGFIR 67
AF++W+ + F ETTSY+ ANIKI F + N+ D G ++ G
Sbjct: 555 AFSKWTPVVNIA-FQETTSYETANIKILFASKNHGDPYPFDGPGG-ILGHAFAPTDGRCH 728
Query: 68 LIASKYWVLPTDNYMWSWQNGEFDLETAAMHQIGHLLGLDHSFDKEYVMYPAILP 122
A +YWV D S FDLE+ A+H+IGHLLGL HS D +MYP+I P
Sbjct: 729 FDADEYWVASGD-VTKSPVTSAFDLESVAVHEIGHLLGLGHSSDLRAIMYPSIPP 890
>TC232889 weakly similar to UP|O65340 (O65340) Metalloproteinase , partial
(38%)
Length = 734
Score = 70.9 bits (172), Expect = 2e-13
Identities = 48/117 (41%), Positives = 66/117 (56%)
Frame = +1
Query: 4 VFRNAFTRWSQTTRVLNFSETTSYDDANIKIGFYNINYIDGVDDVVVGDTVIKLGSNVNS 63
VF AF RW++ T L F ET SY A+I+IGF++ ++ DG T+ S N
Sbjct: 109 VFAAAFARWAEVTS-LKFRETASYFGADIRIGFFSGDHGDGEPFDGSLGTLAHAFSPTNG 285
Query: 64 GFIRLIASKYWVLPTDNYMWSWQNGEFDLETAAMHQIGHLLGLDHSFDKEYVMYPAI 120
F L A++ WV+ D + DLE+ A+H+IGHLLGL HS +E VM+P I
Sbjct: 286 RF-HLDAAEDWVVSGDVTRSALPTA-VDLESVAVHEIGHLLGLGHSSVEEAVMFPTI 450
>CO985121
Length = 535
Score = 54.3 bits (129), Expect = 2e-08
Identities = 35/91 (38%), Positives = 51/91 (55%), Gaps = 1/91 (1%)
Frame = -2
Query: 31 NIKIGFYNINYIDGVD-DVVVGDTVIKLGSNVNSGFIRLIASKYWVLPTDNYMWSWQNGE 89
+I+IGFY ++ DG D V+G + +G L +++ WV D S N
Sbjct: 534 DIRIGFYGGDHGDGEPFDGVLG--TLAHAXXPTNGMFHLDSAEDWVASGDVTKASLSNA- 364
Query: 90 FDLETAAMHQIGHLLGLDHSFDKEYVMYPAI 120
DLE+ A+H+IGHLLGL HS ++ +MYP I
Sbjct: 363 VDLESVAVHEIGHLLGLGHSSVEDAIMYPTI 271
>TC203520 similar to UP|Q93Z89 (Q93Z89) Matrix metalloproteinase MMP2,
partial (13%)
Length = 434
Score = 51.6 bits (122), Expect = 1e-07
Identities = 26/40 (65%), Positives = 32/40 (80%)
Frame = +2
Query: 92 LETAAMHQIGHLLGLDHSFDKEYVMYPAILPLQQQRKLQI 131
LET AMHQIGHLLGL+HS ++ VMYP IL QQRK+++
Sbjct: 2 LETVAMHQIGHLLGLEHSPKEDSVMYPYILS-SQQRKVKL 118
>BU762711
Length = 428
Score = 48.1 bits (113), Expect = 1e-06
Identities = 22/41 (53%), Positives = 31/41 (74%)
Frame = +1
Query: 91 DLETAAMHQIGHLLGLDHSFDKEYVMYPAILPLQQQRKLQI 131
DLE+ A H+IGH+LGL HS KE VMYP++ P +++ L+I
Sbjct: 61 DLESVATHEIGHVLGLGHSSVKEAVMYPSLSPRRKKVDLRI 183
>AW596048 weakly similar to GP|16901508|gb| matrix metalloproteinase MMP2
{Glycine max}, partial (16%)
Length = 417
Score = 44.7 bits (104), Expect = 1e-05
Identities = 21/46 (45%), Positives = 34/46 (73%)
Frame = +3
Query: 2 TNVFRNAFTRWSQTTRVLNFSETTSYDDANIKIGFYNINYIDGVDD 47
T VFR+AF RW+ + LN +E +Y+ A++K+GFYN++ +GV+D
Sbjct: 288 TKVFRDAFARWAGSVPGLNLTE-MNYNSADLKVGFYNLD--EGVED 416
>TC233494 similar to GB|AAM62930.1|21553837|AY085712 phytochelatin
synthetase-like protein {Arabidopsis thaliana;} ,
partial (29%)
Length = 617
Score = 28.9 bits (63), Expect = 0.73
Identities = 13/38 (34%), Positives = 19/38 (49%)
Frame = +3
Query: 3 NVFRNAFTRWSQTTRVLNFSETTSYDDANIKIGFYNIN 40
+V RN+ W V NFS +Y D N+ + +N N
Sbjct: 39 HVMRNSKKYWRVKVTVTNFSYRMNYSDWNLLVQHHNFN 152
>BU765258 weakly similar to PIR|T14319|T14 protein AX110P - carrot, partial
(32%)
Length = 423
Score = 27.3 bits (59), Expect = 2.1
Identities = 12/18 (66%), Positives = 15/18 (82%)
Frame = +1
Query: 7 NAFTRWSQTTRVLNFSET 24
N +TR S+TT VLNFS+T
Sbjct: 28 NGYTRASRTTLVLNFSKT 81
>TC217067 similar to UP|PE21_ARATH (Q42580) Peroxidase 21 precursor (Atperox
P21) (PRXR5) (ATP2a/ATP2b) , partial (44%)
Length = 892
Score = 26.9 bits (58), Expect = 2.8
Identities = 8/19 (42%), Positives = 15/19 (78%)
Frame = +1
Query: 110 FDKEYVMYPAILPLQQQRK 128
FD+ ++ YP LPL++++K
Sbjct: 736 FDQHFICYPTFLPLERKKK 792
>TC206656
Length = 486
Score = 26.6 bits (57), Expect = 3.6
Identities = 9/21 (42%), Positives = 14/21 (65%)
Frame = -2
Query: 72 KYWVLPTDNYMWSWQNGEFDL 92
KY+ P+ Y+W W++ FDL
Sbjct: 323 KYYKRPSCYYLWFWESPGFDL 261
>CA851175
Length = 537
Score = 25.8 bits (55), Expect = 6.2
Identities = 12/47 (25%), Positives = 23/47 (48%)
Frame = -2
Query: 1 MTNVFRNAFTRWSQTTRVLNFSETTSYDDANIKIGFYNINYIDGVDD 47
+ + R F+RWS LN +++ D GF+ +I+G+ +
Sbjct: 278 LCSAIRKVFSRWSSFNSFLNRAQSRRNIDNTRMCGFFE-EFIEGIQN 141
>BE660494 similar to GP|18252419|gb cytochrome b {Ostrinia nubilalis},
partial (47%)
Length = 565
Score = 25.4 bits (54), Expect = 8.0
Identities = 12/27 (44%), Positives = 16/27 (58%)
Frame = +1
Query: 16 TRVLNFSETTSYDDANIKIGFYNINYI 42
T++L T Y A I+I FY +NYI
Sbjct: 64 TQILTGLFLTIYYTAXIEIXFYRVNYI 144
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.322 0.138 0.424
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,660,462
Number of Sequences: 63676
Number of extensions: 69522
Number of successful extensions: 328
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 319
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 320
length of query: 131
length of database: 12,639,632
effective HSP length: 86
effective length of query: 45
effective length of database: 7,163,496
effective search space: 322357320
effective search space used: 322357320
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 53 (25.0 bits)
Medicago: description of AC144345.5


