

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
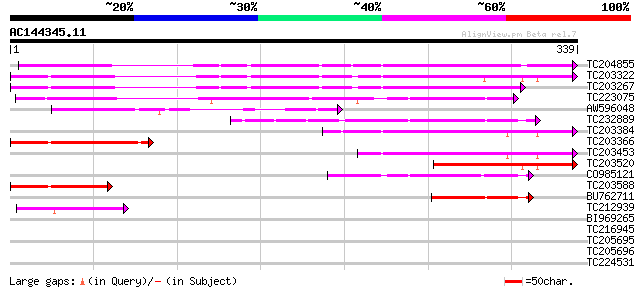
Query= AC144345.11 - phase: 0
(339 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC204855 UP|Q93Z89 (Q93Z89) Matrix metalloproteinase MMP2, complete 189 1e-48
TC203322 similar to UP|Q93Z89 (Q93Z89) Matrix metalloproteinase ... 183 1e-46
TC203267 similar to UP|Q93Z89 (Q93Z89) Matrix metalloproteinase ... 176 1e-44
TC223075 UP|MEP1_SOYBN (P29136) Metalloendoproteinase 1 precurso... 122 3e-28
AW596048 weakly similar to GP|16901508|gb| matrix metalloprotein... 106 1e-23
TC232889 weakly similar to UP|O65340 (O65340) Metalloproteinase ... 99 4e-21
TC203384 similar to UP|Q93Z89 (Q93Z89) Matrix metalloproteinase ... 88 6e-18
TC203366 similar to UP|Q93Z89 (Q93Z89) Matrix metalloproteinase ... 79 2e-15
TC203453 similar to UP|Q93Z89 (Q93Z89) Matrix metalloproteinase ... 75 4e-14
TC203520 similar to UP|Q93Z89 (Q93Z89) Matrix metalloproteinase ... 66 2e-11
CO985121 64 1e-10
TC203588 similar to UP|Q93Z89 (Q93Z89) Matrix metalloproteinase ... 58 5e-09
BU762711 57 9e-09
TC212939 weakly similar to UP|Q9ZUJ5 (Q9ZUJ5) T2K10.2 protein, p... 56 3e-08
BI969265 29 3.5
TC216945 similar to UP|Q43854 (Q43854) Peroxidase precursor , p... 28 4.5
TC205695 28 4.5
TC205696 similar to UP|Q8VZG3 (Q8VZG3) AT3g48690/T8P19_200, part... 28 5.9
TC224531 similar to UP|VATD_ARATH (Q9XGM1) Vacuolar ATP synthase... 28 7.7
>TC204855 UP|Q93Z89 (Q93Z89) Matrix metalloproteinase MMP2, complete
Length = 1307
Score = 189 bits (481), Expect = 1e-48
Identities = 125/336 (37%), Positives = 175/336 (51%), Gaps = 2/336 (0%)
Frame = +1
Query: 6 LSQIKKHLSTFGYFRQFLLKFDDVLDEETISAIKTYQQFFNLQVTGNLNTETLQKISLPR 65
+S IK +LS +GY + + E ISAIKTYQQ+F+LQ TG LN ETLQ++
Sbjct: 277 VSLIKDYLSNYGYIESSGPLSNSMDQETIISAIKTYQQYFSLQPTGKLNNETLQQM---- 444
Query: 66 CGIPDMRYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLLRCGIPDMRYEYGF 125
S LRCG+PD+ +Y F
Sbjct: 445 --------------------------------------------SFLRCGVPDINIDYNF 492
Query: 126 DVGSDVSFPK-GNKWFPKGTKKLTYGFLPGKKFSLDMIEGFRNAFTRWSQTTRVLNFSET 184
++S+PK G++WFP LTYGFLP + +M + FR++F RW+Q + VLN +ET
Sbjct: 493 -TDDNMSYPKAGHRWFPH--TNLTYGFLPENQIPANMTKVFRDSFARWAQASGVLNLTET 663
Query: 185 TSYDDADIKIGFYHIYNNDIVDDVVVGDSFISRNLDSNVKSGMIRLE-RSKFWVSPTTTY 243
T YD+ADI++GFY+ I D V G S I DS K G+I L+ +K W P+
Sbjct: 664 T-YDNADIQVGFYNFTYLGI-DIEVYGGSLIFLQPDST-KKGVILLDGTNKLWALPSENG 834
Query: 244 FKKWELQEFDLETVVMHQIGHLLGLDHSSDKESIMYPLIDPLQEKKVQITDSDNQAIQQL 303
WE DLE+ MH+IGHLLGLDHS+ ++S+MYP I P ++KVQ++ SD +Q
Sbjct: 835 RLSWEEGVLDLESAAMHEIGHLLGLDHSNKEDSVMYPCILPSHQRKVQLSKSDKTNVQHQ 1014
Query: 304 YTNTAKANPNSDHSGCFKLFESSMFLLAFCPLVLLL 339
+ N ++ H G + + L F L+LLL
Sbjct: 1015FAN---VEDSAGHVGRLGVSLITTLSLGFAYLLLLL 1113
>TC203322 similar to UP|Q93Z89 (Q93Z89) Matrix metalloproteinase MMP2,
partial (74%)
Length = 1664
Score = 183 bits (464), Expect = 1e-46
Identities = 124/350 (35%), Positives = 181/350 (51%), Gaps = 11/350 (3%)
Frame = +2
Query: 1 MKIQGLSQIKKHLSTFGYFRQFLLKFDDVLDEETISAIKTYQQFFNLQVTGNLNTETLQK 60
+ I+GLS +K +LS +GY F D D+E ISAIKTYQ F NLQVTG LN + +Q+
Sbjct: 410 INIKGLSVVKDYLSDYGYIESSG-PFTDSFDQEIISAIKTYQNFSNLQVTGGLNKQLIQQ 586
Query: 61 ISLPRCGIPDMRYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLLRCGIPDMR 120
+ +RCG+PD+
Sbjct: 587 MLS------------------------------------------------IRCGVPDVN 622
Query: 121 YEYGFDVGSDVSFPK-GNKWFPKGTKKLTYGFLPGKKFSLDMIEGFRNAFTRWSQTTRVL 179
++Y F ++S+PK G++WFP + LTYGFLP + +M + FR++F RW+Q + L
Sbjct: 623 FDYNF-TDDNISYPKAGHRWFPN--RNLTYGFLPENQIPDNMTKVFRDSFARWAQASGTL 793
Query: 180 NFSETTSYDDADIKIGFYHIYNNDIVDDVVVGDSFISRNLDSNVKSGMIRLERSKFWVSP 239
+ ETT YD+ADI++GFY+ N+ I V G S I DS+ K G++ L+ + W+ P
Sbjct: 794 SLRETT-YDNADIQVGFYNFTNSRIE---VYGGSTIFLQPDSS-KKGVVLLDGNMGWLLP 958
Query: 240 TTTYFKKWELQEFDLETVVMHQIGHLLGLDHSSDKESIMYPLI--DPLQEKKVQITDSDN 297
+ + DLETV MHQIGHLLGLDHS ++S+MYP I +++KVQ+++SD
Sbjct: 959 SENATLSKDDGVLDLETVAMHQIGHLLGLDHSHKEDSVMYPYILSSQSEQRKVQLSNSDK 1138
Query: 298 QAIQQLYT----NTAKANPNS----DHSGCFKLFESSMFLLAFCPLVLLL 339
I + + PNS H G + + F L F ++LLL
Sbjct: 1139ANIHLQFAKHD*DVTSLLPNSHDSASHGGGLDVLLVTTFSLGFAYVLLLL 1288
>TC203267 similar to UP|Q93Z89 (Q93Z89) Matrix metalloproteinase MMP2,
partial (61%)
Length = 1338
Score = 176 bits (447), Expect = 1e-44
Identities = 112/309 (36%), Positives = 165/309 (53%), Gaps = 1/309 (0%)
Frame = +3
Query: 1 MKIQGLSQIKKHLSTFGYFRQFLLKFDDVLDEETISAIKTYQQFFNLQVTGNLNTETLQK 60
+ I+GLS +K +LS +GY F++ D+ET+SAIKTYQ+F NL VTG N + +Q+
Sbjct: 291 INIKGLSVVKDYLSEYGYIESSR-PFNNSFDQETMSAIKTYQKFSNLPVTGVPNKQLIQQ 467
Query: 61 ISLPRCGIPDMRYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLLRCGIPDMR 120
+ LRCG+PD+
Sbjct: 468 MLS------------------------------------------------LRCGVPDVN 503
Query: 121 YEYGFDVGSDVSFPK-GNKWFPKGTKKLTYGFLPGKKFSLDMIEGFRNAFTRWSQTTRVL 179
++Y F + S+PK G++WFP + LTYGFLP + +M + FR++F RW+Q + L
Sbjct: 504 FDYNF-TDDNTSYPKAGHRWFPN--RNLTYGFLPENQIPDNMTKVFRDSFARWAQASGTL 674
Query: 180 NFSETTSYDDADIKIGFYHIYNNDIVDDVVVGDSFISRNLDSNVKSGMIRLERSKFWVSP 239
+ +ETT YD+ADI++GFY+ N+ I V G S I DS+ K G++ L + W+ P
Sbjct: 675 SLTETT-YDNADIQVGFYNFTNSRIE---VYGGSLIFLQPDSS-KKGVVLLNGNMGWLLP 839
Query: 240 TTTYFKKWELQEFDLETVVMHQIGHLLGLDHSSDKESIMYPLIDPLQEKKVQITDSDNQA 299
+ + DLET MHQIGHLLGLDHS ++S+MYP I Q++KVQ+++SD
Sbjct: 840 SENATLSKDDGVLDLETAAMHQIGHLLGLDHSHKEDSVMYPYILSSQQRKVQLSNSDKAN 1019
Query: 300 IQQLYTNTA 308
I + A
Sbjct: 1020IHLQFAKHA 1046
>TC223075 UP|MEP1_SOYBN (P29136) Metalloendoproteinase 1 precursor (SMEP1) ,
complete
Length = 1198
Score = 122 bits (305), Expect = 3e-28
Identities = 93/309 (30%), Positives = 135/309 (43%), Gaps = 8/309 (2%)
Frame = +3
Query: 4 QGLSQIKKHLSTFGYFRQFLLKFDDVLDEETISAIKTYQQFFNLQVTGNLNTETLQKISL 63
+GLS +K + GY FDD D+ +SAIKTYQ+ +NL VTG + TL++I
Sbjct: 198 KGLSNVKNYFHHLGYIPN-APHFDDNFDDTLVSAIKTYQKNYNLNVTGKFDINTLKQIMT 374
Query: 64 PRCGIPDMRYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLLRCGIPDM---- 119
P RCG+PD+
Sbjct: 375 P------------------------------------------------RCGVPDIIINT 410
Query: 120 RYEYGFDVGSDVSFPKGNKWFPKGTKKLTYGFLPGKKFSLDMIEGFRNAFTRWSQTTRVL 179
F + SD +F K + GT +LTY F P + AF++W+ +
Sbjct: 411 NKTTSFGMISDYTFFKDMPRWQAGTTQLTYAFSPEPRLDDTFKSAIARAFSKWTPVVNIA 590
Query: 180 NFSETTSYDDADIKIGFYHIYNNDIVD----DVVVGDSFISRNLDSNVKSGMIRLERSKF 235
F ETTSY+ A+IKI F + D ++G +F + G + ++
Sbjct: 591 -FQETTSYETANIKILFASKNHGDPYPFDGPGGILGHAFAPTD-------GRCHFDADEY 746
Query: 236 WVSPTTTYFKKWELQEFDLETVVMHQIGHLLGLDHSSDKESIMYPLIDPLQEKKVQITDS 295
WV+ + K FDLE+V +H+IGHLLGL HSSD +IMYP I P + +KV +
Sbjct: 747 WVA-SGDVTKSPVTSAFDLESVAVHEIGHLLGLGHSSDLRAIMYPSIPP-RTRKVNLAQD 920
Query: 296 DNQAIQQLY 304
D I++LY
Sbjct: 921 DIDGIRKLY 947
>AW596048 weakly similar to GP|16901508|gb| matrix metalloproteinase MMP2
{Glycine max}, partial (16%)
Length = 417
Score = 106 bits (265), Expect = 1e-23
Identities = 67/178 (37%), Positives = 92/178 (51%), Gaps = 4/178 (2%)
Frame = +3
Query: 26 FDDVLDEETISAIKTYQQFFNLQVTGNLNTETLQKISLPRCGIPDMRYEYGFDVGSD-VS 84
F+D LD+ T+SAI TYQ+FFNL++TG+L ETLQ+ISLPRCG+PDM ++Y DV D VS
Sbjct: 24 FNDDLDQATVSAITTYQRFFNLKITGDLTNETLQQISLPRCGVPDMNFDY--DVSKDNVS 197
Query: 85 FPKG---NKWFPKGTKKLTYGFLPGKKFSLLRCGIPDMRYEYGFDVGSDVSFPKGNKWFP 141
+P +WFP + LTYGF P K P
Sbjct: 198 WPMSRYHRRWFP--DRNLTYGFSPASK-------------------------------IP 278
Query: 142 KGTKKLTYGFLPGKKFSLDMIEGFRNAFTRWSQTTRVLNFSETTSYDDADIKIGFYHI 199
K+ FR+AF RW+ + LN +E +Y+ AD+K+GFY++
Sbjct: 279 SNATKV-----------------FRDAFARWAGSVPGLNLTE-MNYNSADLKVGFYNL 398
>TC232889 weakly similar to UP|O65340 (O65340) Metalloproteinase , partial
(38%)
Length = 734
Score = 98.6 bits (244), Expect = 4e-21
Identities = 71/186 (38%), Positives = 103/186 (55%), Gaps = 1/186 (0%)
Frame = +1
Query: 133 FPKGNKWFPKGTKKLTYGFLPGKKFSLDMIEG-FRNAFTRWSQTTRVLNFSETTSYDDAD 191
FP +W P+GT++LTY F PG S D ++G F AF RW++ T L F ET SY AD
Sbjct: 19 FPGMPRW-PEGTQELTYAFFPGNGLS-DAVKGVFAAAFARWAEVTS-LKFRETASYFGAD 189
Query: 192 IKIGFYHIYNNDIVDDVVVGDSFISRNLDSNVKSGMIRLERSKFWVSPTTTYFKKWELQE 251
I+IGF+ + D D S + + +G L+ ++ WV + +
Sbjct: 190 IRIGFF---SGDHGDGEPFDGSLGTLAHAFSPTNGRFHLDAAEDWVV-SGDVTRSALPTA 357
Query: 252 FDLETVVMHQIGHLLGLDHSSDKESIMYPLIDPLQEKKVQITDSDNQAIQQLYTNTAKAN 311
DLE+V +H+IGHLLGL HSS +E++M+P I ++KKV + D + IQ LY +N
Sbjct: 358 VDLESVAVHEIGHLLGLGHSSVEEAVMFPTISS-RKKKVVLARDDIEGIQFLY----GSN 522
Query: 312 PNSDHS 317
PN + S
Sbjct: 523 PNFNGS 540
>TC203384 similar to UP|Q93Z89 (Q93Z89) Matrix metalloproteinase MMP2,
partial (32%)
Length = 643
Score = 87.8 bits (216), Expect = 6e-18
Identities = 59/168 (35%), Positives = 91/168 (54%), Gaps = 16/168 (9%)
Frame = +1
Query: 188 DDADIKIGFYHIYNNDIVDDVVVGDSFISRNLDSNVKSGMIRLERSKFWVSPTTTYFKKW 247
D+ADI++GFY+ + ++ V G S I DS+ K GM+ ++ + W+ P+
Sbjct: 1 DNADIQVGFYN-FTALSIEVEVYGGSLIFLQPDSS-KKGMVLMDGNMGWLLPSENATLSK 174
Query: 248 ELQEFDLETVVMHQIGHLLGLDHSSDKESIMYPLIDPLQEKKVQITDSD--NQAIQQLYT 305
+ DLETV MHQIGHLLGLDHS ++S+MYP I Q++KV++++SD N +Q
Sbjct: 175 DDGVLDLETVAMHQIGHLLGLDHSHKEDSVMYPYILSSQQRKVKLSNSDKANIHLQFASH 354
Query: 306 NTAKANPNS--------------DHSGCFKLFESSMFLLAFCPLVLLL 339
++ +PNS H G + + F L F ++LLL
Sbjct: 355 DSDMTSPNSHDSDLTSPNSHDSASHGGRLDVLLVTTFSLGFAYVLLLL 498
>TC203366 similar to UP|Q93Z89 (Q93Z89) Matrix metalloproteinase MMP2,
partial (18%)
Length = 576
Score = 79.3 bits (194), Expect = 2e-15
Identities = 40/86 (46%), Positives = 57/86 (65%)
Frame = +1
Query: 1 MKIQGLSQIKKHLSTFGYFRQFLLKFDDVLDEETISAIKTYQQFFNLQVTGNLNTETLQK 60
+ I+GLS +K +LS +GY F+D D+E ISAIKTYQ F NL VTG+LN + +Q+
Sbjct: 325 INIKGLSVVKDYLSDYGYIESSG-PFNDSFDQEIISAIKTYQNFSNLNVTGDLNKQLIQQ 501
Query: 61 ISLPRCGIPDMRYEYGFDVGSDVSFP 86
I RCG+PD+ ++Y F + S+P
Sbjct: 502 ILSIRCGVPDVNFDYNF-TDDNTSYP 576
>TC203453 similar to UP|Q93Z89 (Q93Z89) Matrix metalloproteinase MMP2,
partial (29%)
Length = 622
Score = 75.1 bits (183), Expect = 4e-14
Identities = 51/147 (34%), Positives = 77/147 (51%), Gaps = 16/147 (10%)
Frame = +1
Query: 209 VVGDSFISRNLDSNVKSGMIRLERSKFWVSPTTTYFKKWELQEFDLETVVMHQIGHLLGL 268
V G S I DS+ K G++ ++ + W+ P+ + DLETV MHQIGHLLGL
Sbjct: 34 VYGGSLIFLQPDSS-KKGVVLMDGNMGWLLPSENATLSKDDGVLDLETVAMHQIGHLLGL 210
Query: 269 DHSSDKESIMYPLIDPLQEKKVQITDSD--NQAIQQLYTNTAKANPNS------------ 314
DHS ++S+MYP I Q++KV++++SD N +Q ++ +PNS
Sbjct: 211 DHSHKEDSVMYPYILSSQQRKVKLSNSDKANIHLQFASHDSDLTSPNSHDSDLTSPNSHD 390
Query: 315 --DHSGCFKLFESSMFLLAFCPLVLLL 339
H G + + F L F ++LLL
Sbjct: 391 SASHGGRLDVLLVTTFSLGFAYVLLLL 471
>TC203520 similar to UP|Q93Z89 (Q93Z89) Matrix metalloproteinase MMP2,
partial (13%)
Length = 434
Score = 66.2 bits (160), Expect = 2e-11
Identities = 38/92 (41%), Positives = 56/92 (60%), Gaps = 6/92 (6%)
Frame = +2
Query: 254 LETVVMHQIGHLLGLDHSSDKESIMYPLIDPLQEKKVQITDSDNQAIQQLYT--NTAKAN 311
LETV MHQIGHLLGL+HS ++S+MYP I Q++KV++++SD I + ++ +
Sbjct: 2 LETVAMHQIGHLLGLEHSPKEDSVMYPYILSSQQRKVKLSNSDKANIHLEFAKHDSDLTS 181
Query: 312 PNS----DHSGCFKLFESSMFLLAFCPLVLLL 339
PNS H G + + F L F ++LLL
Sbjct: 182 PNSHDSASHGGRLDVLLVTTFSLGFAYVLLLL 277
>CO985121
Length = 535
Score = 63.5 bits (153), Expect = 1e-10
Identities = 46/123 (37%), Positives = 67/123 (54%)
Frame = -2
Query: 191 DIKIGFYHIYNNDIVDDVVVGDSFISRNLDSNVKSGMIRLERSKFWVSPTTTYFKKWELQ 250
DI+IGFY + D V + +N GM L+ ++ WV+ + K
Sbjct: 534 DIRIGFYGGDHGDGEPFDGVLGTLAHAXXPTN---GMFHLDSAEDWVA-SGDVTKASLSN 367
Query: 251 EFDLETVVMHQIGHLLGLDHSSDKESIMYPLIDPLQEKKVQITDSDNQAIQQLYTNTAKA 310
DLE+V +H+IGHLLGL HSS +++IMYP I + +KV++ + D Q IQ LY +
Sbjct: 366 AVDLESVAVHEIGHLLGLGHSSVEDAIMYPTI-TARTRKVELNEDDIQGIQVLY----GS 202
Query: 311 NPN 313
NPN
Sbjct: 201 NPN 193
>TC203588 similar to UP|Q93Z89 (Q93Z89) Matrix metalloproteinase MMP2,
partial (14%)
Length = 514
Score = 58.2 bits (139), Expect = 5e-09
Identities = 31/61 (50%), Positives = 40/61 (64%)
Frame = +3
Query: 1 MKIQGLSQIKKHLSTFGYFRQFLLKFDDVLDEETISAIKTYQQFFNLQVTGNLNTETLQK 60
+ I GL +K +LS +GY F+D D+E ISAIKTYQ F NLQVTG LN E +Q+
Sbjct: 333 INITGLYIVKDYLSDYGYIESSG-PFNDSFDQEIISAIKTYQNFSNLQVTGGLNKELIQQ 509
Query: 61 I 61
+
Sbjct: 510 M 512
>BU762711
Length = 428
Score = 57.4 bits (137), Expect = 9e-09
Identities = 30/61 (49%), Positives = 40/61 (65%)
Frame = +1
Query: 253 DLETVVMHQIGHLLGLDHSSDKESIMYPLIDPLQEKKVQITDSDNQAIQQLYTNTAKANP 312
DLE+V H+IGH+LGL HSS KE++MYP + P + KKV + D +Q LY +NP
Sbjct: 61 DLESVATHEIGHVLGLGHSSVKEAVMYPSLSP-RRKKVDLRIDDVVGVQALY----GSNP 225
Query: 313 N 313
N
Sbjct: 226 N 228
>TC212939 weakly similar to UP|Q9ZUJ5 (Q9ZUJ5) T2K10.2 protein, partial (13%)
Length = 821
Score = 55.8 bits (133), Expect = 3e-08
Identities = 27/70 (38%), Positives = 40/70 (56%), Gaps = 3/70 (4%)
Frame = +1
Query: 5 GLSQIKKHLSTFGYFRQFLLK---FDDVLDEETISAIKTYQQFFNLQVTGNLNTETLQKI 61
GLS +K + FGY K F D D+ +A++ YQ+ FNL +TG L+ T+ +I
Sbjct: 592 GLSNLKNYFHYFGYISNVSSKSSNFSDDFDDALEAAVRAYQKNFNLNITGELDDPTMNQI 771
Query: 62 SLPRCGIPDM 71
PRCG+ D+
Sbjct: 772 VKPRCGVADI 801
>BI969265
Length = 770
Score = 28.9 bits (63), Expect = 3.5
Identities = 13/41 (31%), Positives = 22/41 (52%)
Frame = +3
Query: 212 DSFISRNLDSNVKSGMIRLERSKFWVSPTTTYFKKWELQEF 252
+S + NL + K+ + E+ K WVS T+F W + +F
Sbjct: 156 ESHTTSNLS*SRKAFIASFEKGKNWVSRGKTFFDNWMVHKF 278
>TC216945 similar to UP|Q43854 (Q43854) Peroxidase precursor , partial (96%)
Length = 1274
Score = 28.5 bits (62), Expect = 4.5
Identities = 14/51 (27%), Positives = 22/51 (42%)
Frame = +2
Query: 238 SPTTTYFKKWELQEFDLETVVMHQIGHLLGLDHSSDKESIMYPLIDPLQEK 288
S +T + D VV GH +G+ H + +YP DP+ +K
Sbjct: 539 SNASTILSSLATKNLDPTDVVALSGGHTIGISHCGSFTNRLYPTQDPVMDK 691
>TC205695
Length = 1332
Score = 28.5 bits (62), Expect = 4.5
Identities = 14/36 (38%), Positives = 19/36 (51%)
Frame = -2
Query: 254 LETVVMHQIGHLLGLDHSSDKESIMYPLIDPLQEKK 289
L TV +H H HS+ +YPL DP Q++K
Sbjct: 995 LRTVPLHLASHSSSQPHSTQP---LYPLSDPFQQRK 897
>TC205696 similar to UP|Q8VZG3 (Q8VZG3) AT3g48690/T8P19_200, partial (18%)
Length = 545
Score = 28.1 bits (61), Expect = 5.9
Identities = 14/37 (37%), Positives = 19/37 (50%)
Frame = -2
Query: 254 LETVVMHQIGHLLGLDHSSDKESIMYPLIDPLQEKKV 290
L TV +H H HS+ +YPL DP Q++ V
Sbjct: 343 LRTVPLHLASHSSSQPHSTQP---LYPLSDPFQQRNV 242
>TC224531 similar to UP|VATD_ARATH (Q9XGM1) Vacuolar ATP synthase subunit D
(V-ATPase D subunit) (Vacuolar proton pump D subunit) ,
partial (42%)
Length = 374
Score = 27.7 bits (60), Expect = 7.7
Identities = 11/24 (45%), Positives = 18/24 (74%)
Frame = -2
Query: 191 DIKIGFYHIYNNDIVDDVVVGDSF 214
D + GF+H++ +D++ DVV GD F
Sbjct: 286 DAEGGFFHVFEDDVL-DVVAGDVF 218
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.321 0.140 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,698,802
Number of Sequences: 63676
Number of extensions: 189038
Number of successful extensions: 860
Number of sequences better than 10.0: 39
Number of HSP's better than 10.0 without gapping: 743
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 838
length of query: 339
length of database: 12,639,632
effective HSP length: 98
effective length of query: 241
effective length of database: 6,399,384
effective search space: 1542251544
effective search space used: 1542251544
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 59 (27.3 bits)
Medicago: description of AC144345.11


