

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC143341.5 + phase: 0
(485 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
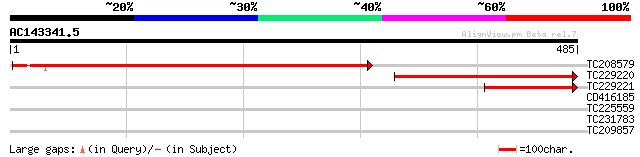
Sequences producing significant alignments: (bits) Value
TC208579 similar to UP|PURA_ARATH (Q96529) Adenylosuccinate synt... 529 e-150
TC229220 similar to PDB|1DJ2_A.0|7546402|1DJ2_A Chain A, Structu... 296 2e-80
TC229221 similar to PDB|1DJ2_A.0|7546402|1DJ2_A Chain A, Structu... 143 2e-34
CD416185 similar to GP|5139695|dbj| expressed in cucumber hypoco... 29 5.4
TC225559 similar to UP|Q9XIV1 (Q9XIV1) Cucumis sativus mRNA expr... 29 5.4
TC231783 homologue to UP|Q9LZQ9 (Q9LZQ9) ATP-dependent RNA helic... 28 7.0
TC209857 28 9.2
>TC208579 similar to UP|PURA_ARATH (Q96529) Adenylosuccinate synthetase,
chloroplast precursor (IMP--aspartate ligase) (AdSS)
(AMPSase) , partial (53%)
Length = 979
Score = 529 bits (1363), Expect = e-150
Identities = 267/310 (86%), Positives = 282/310 (90%), Gaps = 2/310 (0%)
Frame = +3
Query: 3 ISSLRLDSHAICTHQPQRPFAFHNRFP--RNVVVCAAKPVAPPPTKLAAADSSLRRIESL 60
ISSL LDSHAIC PQR + P RNVVVC+AKPVAPPPTKLAA D+ RI L
Sbjct: 54 ISSLTLDSHAICN--PQRSISLRQVRPTTRNVVVCSAKPVAPPPTKLAADDTYGCRIGKL 227
Query: 61 SQVSGVLGCQWGDEGKGKLVDILAQHFQIVARCQGGANAGHTIYNAEGKKFALHLVPSGI 120
SQ SGVLGCQWGD+GKGKLVDILAQHF+IVARCQGGANAGHTIYNAEGKKFALHLVPSGI
Sbjct: 228 SQDSGVLGCQWGDQGKGKLVDILAQHFEIVARCQGGANAGHTIYNAEGKKFALHLVPSGI 407
Query: 121 LNEDTLCVIGNGVVVHLPGLFQEIDNLESNGVSCKGRILISDRAHLLFDFHQTVDGLREA 180
LNEDTLCVIGNGVVVHLPGLF+EID LESNGVSCKGRILISDRAHLLFDFHQ VDGLREA
Sbjct: 408 LNEDTLCVIGNGVVVHLPGLFKEIDGLESNGVSCKGRILISDRAHLLFDFHQVVDGLREA 587
Query: 181 ELSKSFIGTTKRGIGPCYSSKANRNGIRVGDLRYMETLPQKLDLLLSDAALRFKDFKYGP 240
EL+KSFIGTTKRGIGPCYSSK NRNG+RVGDLR+M+T PQKLDL+LSDAALRFKDF YGP
Sbjct: 588 ELAKSFIGTTKRGIGPCYSSKVNRNGLRVGDLRHMDTFPQKLDLILSDAALRFKDFNYGP 767
Query: 241 DVLREEVEKYKRYAERLEPYIADTVHVMNEAITQKKKVLVEGGQATMLDIDFGTYPFVTS 300
+LREEVEKYKRYAERLEP+IADTV VMN+AI QKKK+LVEGGQATMLDIDFGTYPFVTS
Sbjct: 768 YMLREEVEKYKRYAERLEPFIADTVLVMNDAIEQKKKILVEGGQATMLDIDFGTYPFVTS 947
Query: 301 SSPSAGGICT 310
SSPSAGGICT
Sbjct: 948 SSPSAGGICT 977
>TC229220 similar to PDB|1DJ2_A.0|7546402|1DJ2_A Chain A, Structures Of
Adenylosuccinate Synthetase From Triticum Aestivum And
Arabidopsis Thaliana. {Arabidopsis thaliana;} , partial
(35%)
Length = 769
Score = 296 bits (757), Expect = 2e-80
Identities = 143/156 (91%), Positives = 151/156 (96%)
Frame = +1
Query: 330 TTRVGSGPFPTEILGPGGDLLRFAGQEFGTTTGRPRRCGWLDIVALSYSCQINGFSSLNL 389
TTRVGSGPFPTEILG GGDLLRFAGQEFGTTTGRPRRCGWLD+VAL YSCQINGFSSLNL
Sbjct: 10 TTRVGSGPFPTEILGSGGDLLRFAGQEFGTTTGRPRRCGWLDLVALKYSCQINGFSSLNL 189
Query: 390 TKLDVLSDLDEIQLGVSYKHADGTPVQSFPSDLRLLEQLKVEYESLPGWKTDISSIRNYS 449
TKLDVLSDL+EIQLGVSYK ADGTP++SFPSDLRLLEQLKVEYE LPGWK DISSIRNYS
Sbjct: 190 TKLDVLSDLEEIQLGVSYKLADGTPIKSFPSDLRLLEQLKVEYEVLPGWKYDISSIRNYS 369
Query: 450 DLPRAAQLYVERIEELVGVPIHYIGVGPGRDALIFK 485
DLP+AA+LYVERIEELVGVPIHYIG+GPGRDALI+K
Sbjct: 370 DLPKAARLYVERIEELVGVPIHYIGIGPGRDALIYK 477
>TC229221 similar to PDB|1DJ2_A.0|7546402|1DJ2_A Chain A, Structures Of
Adenylosuccinate Synthetase From Triticum Aestivum And
Arabidopsis Thaliana. {Arabidopsis thaliana;} , partial
(18%)
Length = 584
Score = 143 bits (360), Expect = 2e-34
Identities = 68/79 (86%), Positives = 75/79 (94%)
Frame = +3
Query: 407 YKHADGTPVQSFPSDLRLLEQLKVEYESLPGWKTDISSIRNYSDLPRAAQLYVERIEELV 466
YK ADGTP++SF SDL LLEQLKVEYE LPGWK+DISSIRNYSDLP+AA+LYVERIEELV
Sbjct: 3 YKLADGTPIKSFXSDLXLLEQLKVEYEVLPGWKSDISSIRNYSDLPKAARLYVERIEELV 182
Query: 467 GVPIHYIGVGPGRDALIFK 485
GVPIHYIG+GPGRDALI+K
Sbjct: 183 GVPIHYIGIGPGRDALIYK 239
>CD416185 similar to GP|5139695|dbj| expressed in cucumber hypocotyls
{Cucumis sativus}, partial (32%)
Length = 530
Score = 28.9 bits (63), Expect = 5.4
Identities = 13/28 (46%), Positives = 16/28 (56%)
Frame = +2
Query: 298 VTSSSPSAGGICTGLGIAPRVIGDLVGV 325
V +AGG+ TG G A V+GD GV
Sbjct: 434 VAGGGEAAGGLVTGAGAAGVVVGDAAGV 517
>TC225559 similar to UP|Q9XIV1 (Q9XIV1) Cucumis sativus mRNA expressed in
cucumber hypocotyls, complete cds (Arabinogalactan
protein), partial (47%)
Length = 870
Score = 28.9 bits (63), Expect = 5.4
Identities = 13/28 (46%), Positives = 16/28 (56%)
Frame = -3
Query: 298 VTSSSPSAGGICTGLGIAPRVIGDLVGV 325
V +AGG+ TG G A V+GD GV
Sbjct: 358 VAGGGEAAGGLVTGAGAAGVVVGDAAGV 275
>TC231783 homologue to UP|Q9LZQ9 (Q9LZQ9) ATP-dependent RNA helicase-like
protein (At3g62310), partial (24%)
Length = 529
Score = 28.5 bits (62), Expect = 7.0
Identities = 15/48 (31%), Positives = 26/48 (53%)
Frame = -1
Query: 265 VHVMNEAITQKKKVLVEGGQATMLDIDFGTYPFVTSSSPSAGGICTGL 312
+ N I + K+++E T I+ T+P+++SSS S +CT L
Sbjct: 442 IRSQNFGIGLRLKIVIE----TFFSIESKTFPWLSSSSTSRSLVCTSL 311
>TC209857
Length = 857
Score = 28.1 bits (61), Expect = 9.2
Identities = 13/33 (39%), Positives = 19/33 (57%)
Frame = +3
Query: 283 GQATMLDIDFGTYPFVTSSSPSAGGICTGLGIA 315
G L+ FG P +++P +GG+C GLG A
Sbjct: 306 GARARLETLFGPGP*SLAAAPVSGGVCMGLGQA 404
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.320 0.140 0.416
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,926,465
Number of Sequences: 63676
Number of extensions: 297406
Number of successful extensions: 1323
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 1317
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1322
length of query: 485
length of database: 12,639,632
effective HSP length: 101
effective length of query: 384
effective length of database: 6,208,356
effective search space: 2384008704
effective search space used: 2384008704
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC143341.5


