

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC143339.11 + phase: 0
(678 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
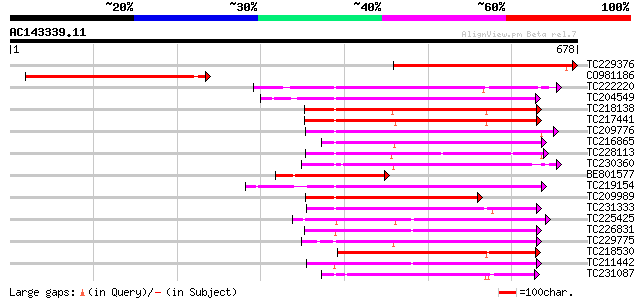
Score E
Sequences producing significant alignments: (bits) Value
TC229376 similar to UP|Q6UY57 (Q6UY57) Lectin-like receptor kina... 331 7e-91
CO981186 234 9e-62
TC222220 weakly similar to UP|Q9ZP16 (Q9ZP16) Receptor-like prot... 226 3e-59
TC204549 similar to UP|Q9M0X5 (Q9M0X5) Receptor protein kinase-l... 226 3e-59
TC218138 homologue to UP|Q8GRK2 (Q8GRK2) Somatic embryogenesis r... 215 6e-56
TC217441 homologue to GB|AAK68074.1|14573459|AF384970 somatic em... 214 1e-55
TC209776 similar to UP|O65470 (O65470) Serine/threonine kinase-l... 210 2e-54
TC216865 similar to UP|Q9LPG0 (Q9LPG0) T3F20.24 protein, partial... 202 5e-52
TC228113 similar to UP|Q9XED4 (Q9XED4) Receptor-like protein kin... 200 1e-51
TC230360 similar to GB|AAP68229.1|31711746|BT008790 At5g63940 {A... 197 1e-50
BE801577 similar to PIR|T49986|T499 lectin-like protein kinase-l... 197 2e-50
TC219154 similar to PIR|T05754|T05754 S-receptor kinase M4I22.1... 193 2e-49
TC209989 similar to UP|Q9XED4 (Q9XED4) Receptor-like protein kin... 191 7e-49
TC231333 similar to UP|Q8L741 (Q8L741) AT3g13690/MMM17_12, parti... 187 1e-47
TC225425 similar to GB|AAN31120.1|23506217|AY149966 At2g17220/T2... 187 2e-47
TC226831 similar to PIR|T52285|T52285 serine/threonine-specific ... 186 4e-47
TC229775 weakly similar to UP|KPEL_DROME (Q05652) Probable serin... 184 8e-47
TC218530 similar to UP|Q9SGU0 (Q9SGU0) T6H22.8.1 protein, partia... 183 2e-46
TC211442 similar to UP|Q84QD9 (Q84QD9) Avr9/Cf-9 rapidly elicite... 183 2e-46
TC231087 similar to UP|Q39203 (Q39203) Receptor-like protein kin... 182 6e-46
>TC229376 similar to UP|Q6UY57 (Q6UY57) Lectin-like receptor kinase 1;1,
partial (29%)
Length = 997
Score = 331 bits (848), Expect = 7e-91
Identities = 159/225 (70%), Positives = 190/225 (83%), Gaps = 6/225 (2%)
Frame = +2
Query: 460 LAWEVRYKVALGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPM 519
L W +RYK+ LGV NAL YLH+DAEQCVLHRDIKSANVLLDT+F+TK+GDFGMAKLVDP
Sbjct: 2 LEWHLRYKIVLGVVNALHYLHEDAEQCVLHRDIKSANVLLDTEFNTKVGDFGMAKLVDPR 181
Query: 520 LRTQRTDVVGTYGYLAPEYINGGRASKESDMYSFGIVALELATGRRFFQDGDFHVPLMNW 579
LRTQRT VVGTYGYLAPEY+N GRAS+ESD+YSFG+V+LE+A+GRR +QDG+FHV LMNW
Sbjct: 182 LRTQRTGVVGTYGYLAPEYVNVGRASRESDIYSFGVVSLEMASGRRTYQDGEFHVSLMNW 361
Query: 580 VWGLYVEGNLMCAADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQLEMA 639
VW LYVEG +M AADEKLN EF+V +M+SLL+VGLWCT+ NDKERPKA +VIKVL LE
Sbjct: 362 VWQLYVEGEIMRAADEKLNNEFEVDQMRSLLVVGLWCTNPNDKERPKAAQVIKVLNLEAP 541
Query: 640 LPELPLDMHDRAPPIVPFRQSNGPS------MAPPMTNSLITSGR 678
LP LPLDM++RAPP+ R + PS M+ P+T+SL++ GR
Sbjct: 542 LPVLPLDMYERAPPMEIIRMPHHPSGKNHSGMSTPITSSLVSVGR 676
>CO981186
Length = 801
Score = 234 bits (597), Expect = 9e-62
Identities = 123/225 (54%), Positives = 149/225 (65%), Gaps = 3/225 (1%)
Frame = +1
Query: 19 ILVLFSLLLVLSYLPKTNSLLFNITNFDDPTVASNMSYQGDGK-STNGSIDLNKVSYLFR 77
I V +L + S L SL FNITNF + A NM Y GDG + NGSI+LN V Y FR
Sbjct: 118 ITVFLLVLAIPSPLKTAESLNFNITNFANSESAKNMLYVGDGAVNKNGSIELNIVDYDFR 297
Query: 78 VGRAFYSQPLHLWDKKTNTLTSFTTRFSFTIDKLND--TTYGDGFVFYLAPLGYQIPPNS 135
VGRA Y QPL LWD + +T F+TRF+FTID+ N+ +Y DGF FY+AP GYQIPPN+
Sbjct: 298 VGRALYGQPLRLWDSSSGVVTDFSTRFTFTIDRGNNKSASYADGFAFYIAPHGYQIPPNA 477
Query: 136 AGGVYGLFNATTNSNFVMNYVVGVEFDTFVGPTDPPMKHVGIDDNALTSVAFGKFDIDKN 195
AGG + LFN T+N N+V+ VEFDTF G DPP +HVGIDDN+L SVA KFDIDKN
Sbjct: 478 AGGTFALFNVTSNPFIPRNHVLAVEFDTFNGTIDPPFQHVGIDDNSLKSVATAKFDIDKN 657
Query: 196 LGRVCYVLIDYNSDEKMLEVFWSFKGRFVKGDGSYGNSSISYQID 240
L C L++YN+ + L V WSF G + NSS+SYQID
Sbjct: 658 LXXKCNALVNYNASNRTLFVSWSFNG---AATPNXXNSSVSYQID 783
>TC222220 weakly similar to UP|Q9ZP16 (Q9ZP16) Receptor-like protein kinase,
RLK3 precursor, partial (35%)
Length = 1304
Score = 226 bits (576), Expect = 3e-59
Identities = 136/378 (35%), Positives = 218/378 (56%), Gaps = 9/378 (2%)
Frame = +3
Query: 292 GSLKTVIVVVAVIVPVILVFLIASIVGWVIVKRKRKNCDEGLDEYGIPFPKKFDLDKATI 351
G+ K+V VV ++VPV++ ++ V + I+KR RK +Y + F + AT+
Sbjct: 18 GNKKSVSRVVLIVVPVVVSIILLLCVCYFILKRSRK-------KYNTLLRENFGEESATL 176
Query: 352 PR-RFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFI 410
+F + AATN F ++ +G GG+G VYKG L G+ +AVK++ F
Sbjct: 177 ESLQFGLVTIEAATNKFSYEKRIGEGGFGVVYKGVLPD-GREIAVKKLSKSSGQGANEFK 353
Query: 411 NEVRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKKS--LAWEVRYKV 468
NE+ +I++L HRNLV +G+C E+ E +L++E++ N SLD LF +S L W RYK+
Sbjct: 354 NEILLIAKLQHRNLVTLLGFCLEEHEKMLIYEFVSNKSLDYFLFDSHRSKQLNWSERYKI 533
Query: 469 ALGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVD-PMLRTQRTDV 527
G+A + YLH+ + V+HRD+K +NVLLD++ + K+ DFGMA++V L+ + +
Sbjct: 534 IEGIAQGISYLHEHSRLKVIHRDLKPSNVLLDSNMNPKISDFGMARIVAIDQLQGKTNRI 713
Query: 528 VGTYGYLAPEYINGGRASKESDMYSFGIVALELATGRR----FFQDGDFHVPLMNWVWGL 583
VGTYGY++PEY G+ S++SD++SFG++ LE+ + +R F D H L+++ W
Sbjct: 714 VGTYGYMSPEYAMHGQFSEKSDVFSFGVIVLEIISAKRNTRSVFSD---HDDLLSYTWEQ 884
Query: 584 YVEGNLMCAADEKLNMEF-DVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQLEMALPE 642
+++ + D+ + EF D SE+ + +GL C +RPK +VI L ++ E
Sbjct: 885 WMDEAPLNIFDQSIEAEFCDHSEVVKCIQIGLLCVQEKPDDRPKITQVIS--YLNSSITE 1058
Query: 643 LPLDMHDRAPPIVPFRQS 660
LPL P P RQS
Sbjct: 1059LPL-------PKKPIRQS 1091
>TC204549 similar to UP|Q9M0X5 (Q9M0X5) Receptor protein kinase-like protein,
partial (54%)
Length = 2032
Score = 226 bits (575), Expect = 3e-59
Identities = 126/341 (36%), Positives = 205/341 (59%), Gaps = 6/341 (1%)
Frame = +3
Query: 300 VVAVIVPVILVFLIASIVGWVIVKRKRKNCDEGLDEYGIPFPKKFDLDKATIPR-RFEYS 358
+VA++VP+ + LI IVG + R+ + +G + G K D T+ +F++S
Sbjct: 642 IVAIVVPITVAVLIF-IVGICFLSRRARKKQQGSVKEG-----KTAYDIPTVDSLQFDFS 803
Query: 359 ELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINEVRIISR 418
+ AATN F D LG GG+G+VYKG LS G++VAVKR+ F NEV ++++
Sbjct: 804 TIEAATNKFSADNKLGEGGFGEVYKGTLSS-GQVVAVKRLSKSSGQGGEEFKNEVVVVAK 980
Query: 419 LIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKKS--LAWEVRYKVALGVANAL 476
L HRNLV+ +G+C + +E +LV+EY+PN SLD LF +K L W RYK+ G+A +
Sbjct: 981 LQHRNLVRLLGFCLQGEEKILVYEYVPNKSLDYILFDPEKQRELDWGRRYKIIGGIARGI 1160
Query: 477 RYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVD-PMLRTQRTDVVGTYGYLA 535
+YLH+D+ ++HRD+K++N+LLD D + K+ DFGMA++ + + +VGTYGY+A
Sbjct: 1161QYLHEDSRLRIIHRDLKASNILLDGDMNPKISDFGMARIFGVDQTQGNTSRIVGTYGYMA 1340
Query: 536 PEYINGGRASKESDMYSFGIVALELATGRR--FFQDGDFHVPLMNWVWGLYVEGNLMCAA 593
PEY G S +SD+YSFG++ +E+ +G++ F D L+++ W L+ +G +
Sbjct: 1341PEYAMHGEFSVKSDVYSFGVLLMEILSGKKNSSFYQTDGAEDLLSYAWQLWKDGTPLELM 1520
Query: 594 DEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVL 634
D L ++ +E+ + +GL C + +RP ++ +L
Sbjct: 1521DPILRESYNQNEVIRSIHIGLLCVQEDPADRPTMATIVLML 1643
>TC218138 homologue to UP|Q8GRK2 (Q8GRK2) Somatic embryogenesis receptor
kinase 1, partial (60%)
Length = 1444
Score = 215 bits (547), Expect = 6e-56
Identities = 121/291 (41%), Positives = 179/291 (60%), Gaps = 8/291 (2%)
Frame = +1
Query: 353 RRFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFAD-FENSERVFIN 411
+RF EL AT+ F + +LGRGG+G+VYKG L+ G +VAVKR+ + E F
Sbjct: 115 KRFSLRELQVATDSFSNKNILGRGGFGKVYKGRLAD-GSLVAVKRLKEERTPGGELQFQT 291
Query: 412 EVRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGD---KKSLAWEVRYKV 468
EV +IS +HRNL++ G+C E LLV+ YM NGS+ + L ++ L W R +V
Sbjct: 292 EVEMISMAVHRNLLRLRGFCMTPTERLLVYPYMANGSVASCLRERPPYQEPLDWPTRKRV 471
Query: 469 ALGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRTDVV 528
ALG A L YLHD + ++HRD+K+AN+LLD +F +GDFG+AKL+D T V
Sbjct: 472 ALGSARGLSYLHDHCDPKIIHRDVKAANILLDEEFEAVVGDFGLAKLMDYKDTHVTTAVR 651
Query: 529 GTYGYLAPEYINGGRASKESDMYSFGIVALELATGRRFFQ----DGDFHVPLMNWVWGLY 584
GT G++APEY++ G++S+++D++ +GI+ LEL TG+R F D V L++WV GL
Sbjct: 652 GTIGHIAPEYLSTGKSSEKTDVFGYGIMLLELITGQRAFDLARLANDDDVMLLDWVKGLL 831
Query: 585 VEGNLMCAADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQ 635
E L D L + +E++ L+ V L CT + +RPK EV+++L+
Sbjct: 832 KEKKLEMLVDPDLQTNYIETEVEQLIQVALLCTQGSPMDRPKMSEVVRMLE 984
>TC217441 homologue to GB|AAK68074.1|14573459|AF384970 somatic embryogenesis
receptor-like kinase 3 {Arabidopsis thaliana;} , partial
(54%)
Length = 1008
Score = 214 bits (544), Expect = 1e-55
Identities = 119/291 (40%), Positives = 180/291 (60%), Gaps = 8/291 (2%)
Frame = +2
Query: 353 RRFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFAD-FENSERVFIN 411
+RF EL AT+ F + +LGRGG+G+VYKG L+ G +VAVKR+ + + F
Sbjct: 113 KRFSLRELQVATDNFSNKHILGRGGFGKVYKGRLAD-GSLVAVKRLKEERTQGG*LQFQT 289
Query: 412 EVRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKKS---LAWEVRYKV 468
EV +IS +HRNL++ G+C E LLV+ YM NGS+ + L ++S L W R ++
Sbjct: 290 EVEMISMAVHRNLLRLRGFCMTPTERLLVYPYMANGSVASCLRERQESQPPLGWPERKRI 469
Query: 469 ALGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRTDVV 528
ALG A L YLHD + ++HRD+K+AN+LLD +F +GDFG+AKL+D T V
Sbjct: 470 ALGSARGLAYLHDHCDPKIIHRDVKAANILLDEEFEAVVGDFGLAKLMDYKDTHVTTAVR 649
Query: 529 GTYGYLAPEYINGGRASKESDMYSFGIVALELATGRRFFQ----DGDFHVPLMNWVWGLY 584
GT G++APEY++ G++S+++D++ +G++ LEL TG+R F D V L++WV GL
Sbjct: 650 GTIGHIAPEYLSTGKSSEKTDVFGYGVMLLELITGQRAFDLARLANDDDVMLLDWVKGLL 829
Query: 585 VEGNLMCAADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQ 635
+ L D L ++ E++ L+ V L CT + ERPK EV+++L+
Sbjct: 830 KDRKLETLVDADLQGSYNDEEVEQLIQVALLCTQGSPMERPKMSEVVRMLE 982
>TC209776 similar to UP|O65470 (O65470) Serine/threonine kinase-like protein
, partial (47%)
Length = 1305
Score = 210 bits (534), Expect = 2e-54
Identities = 117/312 (37%), Positives = 186/312 (59%), Gaps = 9/312 (2%)
Frame = +2
Query: 354 RFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINEV 413
RF++S + AAT F + LG GG+G+VYKG L G+ VAVKR+ F NEV
Sbjct: 41 RFDFSTIEAATQKFSEANKLGEGGFGEVYKGLLPS-GQEVAVKRLSKISGQGGEEFKNEV 217
Query: 414 RIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKK--SLAWEVRYKVALG 471
I+++L HRNLV+ +G+C E +E +LV+E++ N SLD LF +K SL W RYK+ G
Sbjct: 218 EIVAKLQHRNLVRLLGFCLEGEEKILVYEFVANKSLDYILFDPEKQKSLDWTRRYKIVEG 397
Query: 472 VANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVD-PMLRTQRTDVVGT 530
+A ++YLH+D+ ++HRD+K++NVLLD D + K+ DFGMA++ + +VGT
Sbjct: 398 IARGIQYLHEDSRLKIIHRDLKASNVLLDGDMNPKISDFGMARIFGVDQTQANTNRIVGT 577
Query: 531 YGYLAPEYINGGRASKESDMYSFGIVALELATGRR--FFQDGDFHVPLMNWVWGLYVEGN 588
YGY++PEY G S +SD+YSFG++ LE+ +G+R F + D L+++ W L+ +
Sbjct: 578 YGYMSPEYAMHGEYSAKSDVYSFGVLILEIISGKRNSSFYETDVAEDLLSYAWKLWKDEA 757
Query: 589 LMCAADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVL---QLEMALPELP- 644
+ D+ L + +E+ + +GL C + +RP V+ +L + + +P P
Sbjct: 758 PLELMDQSLRESYTRNEVIRCIHIGLLCVQEDPIDRPTMASVVLMLDSYSVTLQVPNQPA 937
Query: 645 LDMHDRAPPIVP 656
++ R P +P
Sbjct: 938 FYINSRTEPNMP 973
>TC216865 similar to UP|Q9LPG0 (Q9LPG0) T3F20.24 protein, partial (28%)
Length = 1358
Score = 202 bits (513), Expect = 5e-52
Identities = 109/275 (39%), Positives = 165/275 (59%), Gaps = 5/275 (1%)
Frame = +1
Query: 373 LGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINEVRIISRLIHRNLVQFIGWCH 432
+G GG+G VYKG S G ++AVK++ + R F+NE+ +IS L H +LV+ G C
Sbjct: 13 IGEGGFGPVYKGCFSD-GTLIAVKQLSSKSRQGNREFLNEIGMISALQHPHLVKLYGCCV 189
Query: 433 EQDELLLVFEYMPNGSLDTHLFGDKK---SLAWEVRYKVALGVANALRYLHDDAEQCVLH 489
E D+LLLV+EYM N SL LFG ++ L W RYK+ +G+A L YLH+++ ++H
Sbjct: 190 EGDQLLLVYEYMENNSLARALFGAEEHQIKLDWTTRYKICVGIARGLAYLHEESRLKIVH 369
Query: 490 RDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRTDVVGTYGYLAPEYINGGRASKESD 549
RDIK+ NVLLD D + K+ DFG+AKL + T + GT+GY+APEY G + ++D
Sbjct: 370 RDIKATNVLLDQDLNPKISDFGLAKLDEEDNTHISTRIAGTFGYMAPEYAMHGYLTDKAD 549
Query: 550 MYSFGIVALELATGR--RFFQDGDFHVPLMNWVWGLYVEGNLMCAADEKLNMEFDVSEMK 607
+YSFGIVALE+ GR + + ++ W L +G++M D +L +EF+ E
Sbjct: 550 VYSFGIVALEIINGRSNTIHRQKEESFSVLEWAHLLREKGDIMDLVDRRLGLEFNKEEAL 729
Query: 608 SLLIVGLWCTHSNDKERPKAYEVIKVLQLEMALPE 642
++ V L CT+ RP V+ +L+ ++ + E
Sbjct: 730 VMIKVALLCTNVTAALRPTMSSVVSMLEGKIVVDE 834
>TC228113 similar to UP|Q9XED4 (Q9XED4) Receptor-like protein kinase homolog
RK20-1, partial (53%)
Length = 1492
Score = 200 bits (509), Expect = 1e-51
Identities = 118/336 (35%), Positives = 184/336 (54%), Gaps = 45/336 (13%)
Frame = +1
Query: 354 RFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINEV 413
+F + + AT F D LG+GG+G VY+G LS G+++AVKR+ + + F NEV
Sbjct: 115 QFNFDTIRVATEDFSDSNKLGQGGFGAVYRGRLSD-GQMIAVKRLSRESSQGDTEFKNEV 291
Query: 414 RIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFG------------------ 455
++++L HRNLV+ +G+C E E LL++EY+PN SLD +FG
Sbjct: 292 LLVAKLQHRNLVRLLGFCLEGKERLLIYEYVPNKSLDYFIFGRS**VHN*TVALYFTSGS 471
Query: 456 --------------------DKKSLAWEVRYKVALGVANALRYLHDDAEQCVLHRDIKSA 495
K L WE+RYK+ GVA L YLH+D+ ++HRD+K++
Sbjct: 472 GQRLNIHIPLKLMAVYADPTKKAQLNWEMRYKIITGVARGLLYLHEDSHLRIIHRDLKAS 651
Query: 496 NVLLDTDFSTKLGDFGMAKLVDPMLRTQRTD--VVGTYGYLAPEYINGGRASKESDMYSF 553
N+LL+ + + K+ DFGMA+LV M +TQ +VGTYGY+APEY G+ S +SD++SF
Sbjct: 652 NILLNEEMNPKIADFGMARLV-LMDQTQANTNRIVGTYGYMAPEYAMHGQFSMKSDVFSF 828
Query: 554 GIVALELATGRR--FFQDGDFHVPLMNWVWGLYVEGNLMCAADEKLNMEFDVSEMKSLLI 611
G++ LE+ +G + + G+ L+++ W + EG + D LN +EM +
Sbjct: 829 GVLVLEIISGHKNSGIRHGENVEDLLSFAWRNWREGTAVKIVDPSLNNN-SRNEMLRCIH 1005
Query: 612 VGLWCTHSNDKERPKAYEVIKVL---QLEMALPELP 644
+GL C N +RP ++ +L L + +P P
Sbjct: 1006IGLLCVQENLADRPTMTTIMLMLNSYSLSLPIPSKP 1113
>TC230360 similar to GB|AAP68229.1|31711746|BT008790 At5g63940 {Arabidopsis
thaliana;} , partial (42%)
Length = 1533
Score = 197 bits (501), Expect = 1e-50
Identities = 116/317 (36%), Positives = 175/317 (54%), Gaps = 5/317 (1%)
Frame = +2
Query: 349 ATIPRRFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERV 408
+++ R + EL++AT+ F D ++GRGG VY+G L GK +AVK I EN +
Sbjct: 404 SSLCRLYRLQELLSATSNFASDNLIGRGGCSYVYRGCLPD-GKELAVK-ILKPSENVIKE 577
Query: 409 FINEVRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDK---KSLAWEVR 465
F+ E+ II+ L H+N++ G+C E + LLLV++++ GSL+ +L G+K + W+ R
Sbjct: 578 FVQEIEIITTLRHKNIISISGFCLEGNHLLLVYDFLSRGSLEENLHGNKVDCSAFGWQER 757
Query: 466 YKVALGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRT 525
YKVA+GVA AL YLH+ Q V+HRD+KS+N+LL DF +L DFG+A T
Sbjct: 758 YKVAVGVAEALDYLHNGCAQAVIHRDVKSSNILLADDFEPQLSDFGLASWGSSSSHITCT 937
Query: 526 DVVGTYGYLAPEYINGGRASKESDMYSFGIVALELATGRRFF--QDGDFHVPLMNWVWGL 583
DV GT+GYLAPEY GR + + D+Y+FG+V LEL + R+ + L+ W +
Sbjct: 938 DVAGTFGYLAPEYFMHGRVTDKIDVYAFGVVLLELLSNRKPINNESPKGQESLVMWATPI 1117
Query: 584 YVEGNLMCAADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQLEMALPEL 643
G D L E++ ++K +++ C + RP Q+ +AL
Sbjct: 1118LEGGKFSQLLDPSLGSEYNTCQIKRMILAATLCIRRIPRLRP---------QINLAL--- 1261
Query: 644 PLDMHDRAPPIVPFRQS 660
LD+ D I QS
Sbjct: 1262-LDLEDDTVSISSTEQS 1309
>BE801577 similar to PIR|T49986|T499 lectin-like protein kinase-like -
Arabidopsis thaliana, partial (7%)
Length = 421
Score = 197 bits (500), Expect = 2e-50
Identities = 99/138 (71%), Positives = 112/138 (80%), Gaps = 2/138 (1%)
Frame = -2
Query: 319 WVIVKRKRKNCDEGLD--EYGIPFPKKFDLDKATIPRRFEYSELVAATNGFDDDRMLGRG 376
WVI+ +KR+ + D E G KFDLD+ TIPRRF+Y ELV ATNGF DD LGRG
Sbjct: 411 WVIIMKKRRGKGDYYDNDESGHT-SAKFDLDRETIPRRFDYKELVVATNGFADDTRLGRG 235
Query: 377 GYGQVYKGALSYLGKIVAVKRIFADFENSERVFINEVRIISRLIHRNLVQFIGWCHEQDE 436
G GQVYKG LS+LG++VAVKRIF + ENSERVFINEVRIISRLIHRNLVQF+GWCHEQ E
Sbjct: 234 GSGQVYKGVLSHLGRVVAVKRIFTNSENSERVFINEVRIISRLIHRNLVQFVGWCHEQGE 55
Query: 437 LLLVFEYMPNGSLDTHLF 454
LLVFE+MPNGSLD+HLF
Sbjct: 54 FLLVFEFMPNGSLDSHLF 1
>TC219154 similar to PIR|T05754|T05754 S-receptor kinase M4I22.110 precursor
- Arabidopsis thaliana {Arabidopsis thaliana;} ,
partial (31%)
Length = 1363
Score = 193 bits (490), Expect = 2e-49
Identities = 128/367 (34%), Positives = 196/367 (52%), Gaps = 7/367 (1%)
Frame = +1
Query: 283 SLVEGNDGKGSLKTVIVVVAVIVPVILVFLIASIVGWVIVKRKR--KNCDEGLDEYGIPF 340
S +E K S K +I+ +V P+ +V I I I + + K+ D L + +P
Sbjct: 136 SELESIKSKKSSK-IIIGTSVAAPLGVVLAICFIYRRNIADKSKTKKSIDRQLQDVDVPL 312
Query: 341 PKKFDLDKATIPRRFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFA 400
F+ + AAT+ F + +G GG+G VYKG L G+ +AVKR+ +
Sbjct: 313 --------------FDMLTITAATDNFLLNNKIGEGGFGPVYKGKLVG-GQEIAVKRLSS 447
Query: 401 DFENSERVFINEVRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKKS- 459
FI EV++I++L HRNLV+ +G C + E LLV+EY+ NGSL++ +F KS
Sbjct: 448 LSGQGITEFITEVKLIAKLQHRNLVKLLGCCIKGQEKLLVYEYVVNGSLNSFIFDQIKSK 627
Query: 460 -LAWEVRYKVALGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDP 518
L W R+ + LG+A L YLH D+ ++HRD+K++NVLLD + K+ DFGMA+
Sbjct: 628 LLDWPRRFNIILGIARGLLYLHQDSRLRIIHRDLKASNVLLDEKLNPKISDFGMARAFGG 807
Query: 519 MLRTQRTD-VVGTYGYLAPEYINGGRASKESDMYSFGIVALELATG--RRFFQDGDFHVP 575
T+ VVGTYGY+APEY G S +SD++SFGI+ LE+ G + + +
Sbjct: 808 DQTEGNTNRVVGTYGYMAPEYAFDGNFSIKSDVFSFGILLLEIVCGIKNKSLCHENQTLN 987
Query: 576 LMNWVWGLYVEGNLMCAADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQ 635
L+ + W L+ E N + D + + E+ + V L C ++RP VI++L
Sbjct: 988 LVGYAWALWKEQNALQLIDSGIKDSCVIPEVLRCIHVSLLCVQQYPEDRPTMTSVIQMLG 1167
Query: 636 LEMALPE 642
EM + E
Sbjct: 1168SEMDMVE 1188
>TC209989 similar to UP|Q9XED4 (Q9XED4) Receptor-like protein kinase homolog
RK20-1, partial (32%)
Length = 738
Score = 191 bits (486), Expect = 7e-49
Identities = 96/215 (44%), Positives = 145/215 (66%), Gaps = 3/215 (1%)
Frame = +3
Query: 354 RFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINEV 413
+FE++ + ATN F D LG+GG+G VYKG LS G+ +A+KR+ + E F NE+
Sbjct: 96 QFEFATIKFATNNFSDANKLGQGGFGIVYKGTLSD-GQEIAIKRLSINSNQGETEFKNEI 272
Query: 414 RIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKK--SLAWEVRYKVALG 471
+ RL HRNLV+ +G+C + E LL++E++PN SLD +F K +L E+RYK+ G
Sbjct: 273 LLTGRLQHRNLVRLLGFCFARRERLLIYEFVPNKSLDYFIFDPNKRVNLN*EIRYKIIRG 452
Query: 472 VANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVD-PMLRTQRTDVVGT 530
+A L YLH+D+ V+HRD+K++N+LLD + + K+ DFGMA+L + T +VGT
Sbjct: 453 IARGLLYLHEDSRLNVVHRDLKTSNILLDGELNPKISDFGMARLFEINQTEASTTTIVGT 632
Query: 531 YGYLAPEYINGGRASKESDMYSFGIVALELATGRR 565
+GY+APEYI G+ S +SD++SFG++ LE+ G+R
Sbjct: 633 FGYMAPEYIKHGQFSIKSDVFSFGVMILEIVCGQR 737
>TC231333 similar to UP|Q8L741 (Q8L741) AT3g13690/MMM17_12, partial (38%)
Length = 1009
Score = 187 bits (475), Expect = 1e-47
Identities = 107/288 (37%), Positives = 163/288 (56%), Gaps = 7/288 (2%)
Frame = +3
Query: 355 FEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINEVR 414
F +SEL AT GF L GG+G V++G L G+++AVK+ ++ F +EV
Sbjct: 6 FTFSELQLATGGFSQANFLAEGGFGSVHRGVLPD-GQVIAVKQYKLASTQGDKEFCSEVE 182
Query: 415 IISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDKKS-LAWEVRYKVALGVA 473
++S HRN+V IG+C + LLV+EY+ NGSLD+H++ K++ L W R K+A+G A
Sbjct: 183 VLSCAQHRNVVMLIGFCVDDGRRLLVYEYICNGSLDSHIYRRKQNVLEWSARQKIAVGAA 362
Query: 474 NALRYLHDDAE-QCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRTDVVGTYG 532
LRYLH++ C++HRD++ N+LL DF +GDFG+A+ T V+GT+G
Sbjct: 363 RGLRYLHEECRVGCIVHRDMRPNNILLTHDFEALVGDFGLARWQPDGDMGVETRVIGTFG 542
Query: 533 YLAPEYINGGRASKESDMYSFGIVALELATGRRFFQDGDFHVP-----LMNWVWGLYVEG 587
YLAPEY G+ ++++D+YSFGIV LEL TGR+ D + P L W L +
Sbjct: 543 YLAPEYAQSGQITEKADVYSFGIVLLELVTGRKAV---DINRPKGQQCLSEWARPLLEKQ 713
Query: 588 NLMCAADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQ 635
D L + E+ +L C + RP+ +V+++L+
Sbjct: 714 ATYKLIDPSLRNCYVDQEVYRMLKCSSLCIGRDPHLRPRMSQVLRMLE 857
>TC225425 similar to GB|AAN31120.1|23506217|AY149966 At2g17220/T23A1.8
{Arabidopsis thaliana;} , partial (80%)
Length = 1386
Score = 187 bits (474), Expect = 2e-47
Identities = 114/322 (35%), Positives = 177/322 (54%), Gaps = 14/322 (4%)
Frame = +1
Query: 339 PFPKKFDLDKATIPRRFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYL-------GK 391
P+P L + + R F ++EL AAT F D +LG GG+G+V+KG L G
Sbjct: 67 PYPNGQILPTSNL-RIFTFAELKAATRNFKADTVLGEGGFGKVFKGWLEEKATSKGGSGT 243
Query: 392 IVAVKRIFADFENSERVFINEVRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDT 451
++AVK++ ++ + +EV + RL H NLV+ +G+C E+ ELLLV+E+M GSL+
Sbjct: 244 VIAVKKLNSESLQGLEEWQSEVNFLGRLSHTNLVKLLGYCLEESELLLVYEFMQKGSLEN 423
Query: 452 HLFGDKKS---LAWEVRYKVALGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLG 508
HLFG + L W++R K+A+G A L +LH + V++RD K++N+LLD ++ K+
Sbjct: 424 HLFGRGSAVQPLPWDIRLKIAIGAARGLAFLH--TSEKVIYRDFKASNILLDGSYNAKIS 597
Query: 509 DFGMAKLVDPMLRTQ-RTDVVGTYGYLAPEYINGGRASKESDMYSFGIVALELATGRRFF 567
DFG+AKL ++ T V+GT+GY APEY+ G +SD+Y FG+V +E+ TG R
Sbjct: 598 DFGLAKLGPSASQSHVTTRVMGTHGYAAPEYVATGHLYVKSDVYGFGVVLVEILTGLRAL 777
Query: 568 QDG--DFHVPLMNWVWG-LYVEGNLMCAADEKLNMEFDVSEMKSLLIVGLWCTHSNDKER 624
L WV L+ L D +L +F + + + C S K R
Sbjct: 778 DSNRPSGQHKLTEWVKPYLHDRRKLKGIMDSRLEGKFPSKAAFRIAQLSMKCLASEPKHR 957
Query: 625 PKAYEVIKVLQLEMALPELPLD 646
P +V++ L+ A E P++
Sbjct: 958 PSMKDVLENLERIQAANEKPVE 1023
>TC226831 similar to PIR|T52285|T52285 serine/threonine-specific protein
kinase APK2a [imported] - Arabidopsis thaliana
{Arabidopsis thaliana;} , partial (82%)
Length = 1693
Score = 186 bits (471), Expect = 4e-47
Identities = 114/297 (38%), Positives = 167/297 (55%), Gaps = 14/297 (4%)
Frame = +1
Query: 353 RRFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSY---------LGKIVAVKRIFADFE 403
+ F ++EL AT F D +LG GG+G VYKG + G +VAVK++ +
Sbjct: 433 KAFTFNELKNATRNFRPDSLLGEGGFGYVYKGWIDEHTFTASKPGSGMVVAVKKLKPEGL 612
Query: 404 NSERVFINEVRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLF-GDKKSLAW 462
+ ++ EV + +L H+NLV+ IG+C + + LLV+E+M GSL+ HLF + L+W
Sbjct: 613 QGHKEWLTEVDYLGQLHHQNLVKLIGYCADGENRLLVYEFMSKGSLENHLFRRGPQPLSW 792
Query: 463 EVRYKVALGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRT 522
VR KVA+G A L +LH +A+ V++RD K++N+LLD +F+ KL DFG+AK RT
Sbjct: 793 SVRMKVAIGAARGLSFLH-NAKSQVIYRDFKASNILLDAEFNAKLSDFGLAKAGPTGDRT 969
Query: 523 Q-RTDVVGTYGYLAPEYINGGRASKESDMYSFGIVALELATGRRFFQDGDFHVP--LMNW 579
T V+GT GY APEY+ GR + +SD+YSFG+V LEL +GRR V L+ W
Sbjct: 970 HVSTQVMGTQGYAAPEYVATGRLTAKSDVYSFGVVLLELLSGRRAVDRSKAGVEQNLVEW 1149
Query: 580 VWG-LYVEGNLMCAADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQ 635
L + L D KL ++ + L C + K RP EV++ L+
Sbjct: 1150AKPYLGDKRRLFRIMDTKLGGQYPQKGAYMAATLALKCLNREAKGRPPITEVLQTLE 1320
>TC229775 weakly similar to UP|KPEL_DROME (Q05652) Probable
serine/threonine-protein kinase pelle , partial (8%)
Length = 1250
Score = 184 bits (468), Expect = 8e-47
Identities = 103/291 (35%), Positives = 165/291 (56%), Gaps = 4/291 (1%)
Frame = +1
Query: 349 ATIPRRFEYSELVAATNGFDDDRMLGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERV 408
A+ R++ ++ELV TN F R+LGRGG+G+VY G + VAVK +
Sbjct: 31 ASKQRQYTFNELVKITNNFT--RILGRGGFGKVYHGFID--DTQVAVKMLSPSAVRGYEQ 198
Query: 409 FINEVRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGDK---KSLAWEVR 465
F+ EV+++ R+ HRNL +G+C+E++ + L++EYM NG+LD + G K L WE R
Sbjct: 199 FLAEVKLLMRVHHRNLTSLVGYCNEENNIGLIYEYMANGNLDEIVSGKSSRAKFLTWEDR 378
Query: 466 YKVALGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRT 525
++AL A L YLH+ + ++HRD+K AN+LL+ +F KL DFG++K + +
Sbjct: 379 LQIALDAAQGLEYLHNGCKPPIIHRDVKCANILLNENFQAKLADFGLSKSFPTDGGSYMS 558
Query: 526 DVV-GTYGYLAPEYINGGRASKESDMYSFGIVALELATGRRFFQDGDFHVPLMNWVWGLY 584
VV GT GYL PEY R +++SD+YSFG+V LE+ TG+ + WV +
Sbjct: 559 TVVAGTPGYLDPEYSISSRLTEKSDVYSFGVVLLEMVTGQPAIAKTPDKTHISQWVKSML 738
Query: 585 VEGNLMCAADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQ 635
G++ AD + +FD S + ++ +G+ + +RP ++ L+
Sbjct: 739 SNGDIKNIADSRFKEDFDTSSVWRIVEIGMASVSISPFKRPSMSYIVNELK 891
>TC218530 similar to UP|Q9SGU0 (Q9SGU0) T6H22.8.1 protein, partial (25%)
Length = 1190
Score = 183 bits (465), Expect = 2e-46
Identities = 98/245 (40%), Positives = 149/245 (60%), Gaps = 3/245 (1%)
Frame = +3
Query: 393 VAVKRIFADFENSERVFINEVRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTH 452
+AVK++ + FI E+ IS + HRNLV+ G C E + LLV+EY+ N SLD
Sbjct: 6 IAVKQLSVGSHQGKSQFITEIATISAVQHRNLVKLYGCCIEGSKRLLVYEYLENKSLDQA 185
Query: 453 LFGDKKSLAWEVRYKVALGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGM 512
LFG +L W RY + LGVA L YLH+++ ++HRD+K++N+LLD + K+ DFG+
Sbjct: 186 LFGKCLTLNWSTRYDICLGVARGLTYLHEESRLRIVHRDVKASNILLDYELIPKISDFGL 365
Query: 513 AKLVDPMLRTQRTDVVGTYGYLAPEYINGGRASKESDMYSFGIVALELATGRRFFQ---D 569
AKL D T V GT GYLAPEY G ++++D++SFG+VALEL +GR +
Sbjct: 366 AKLYDDKKTHISTGVAGTIGYLAPEYAMRGHLTEKADVFSFGVVALELVSGRPNSDSSLE 545
Query: 570 GDFHVPLMNWVWGLYVEGNLMCAADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYE 629
G+ V L+ W W L+ + ++ D++L+ EF+ E+K ++ + L CT ++ RP
Sbjct: 546 GE-KVYLLEWAWQLHEKNCIIDLVDDRLS-EFNEEEVKRVVGIALLCTQTSPTLRPSMSR 719
Query: 630 VIKVL 634
V+ +L
Sbjct: 720 VVAML 734
>TC211442 similar to UP|Q84QD9 (Q84QD9) Avr9/Cf-9 rapidly elicited protein
264, partial (73%)
Length = 1047
Score = 183 bits (465), Expect = 2e-46
Identities = 111/292 (38%), Positives = 163/292 (55%), Gaps = 11/292 (3%)
Frame = +2
Query: 355 FEYSELVAATNGFDDDRMLGRGGYGQVYKGALS------YLGKIVAVKRIFADFENSERV 408
F EL ATN F MLG GG+G VYKG + + VAVKR+ D R
Sbjct: 35 FTLEELREATNSFSWSNMLGEGGFGPVYKGFVDDKLRSGLKAQTVAVKRLDLDGLQGHRE 214
Query: 409 FINEVRIISRLIHRNLVQFIGWCHEQDELLLVFEYMPNGSLDTHLFGD-KKSLAWEVRYK 467
++ E+ + +L H +LV+ IG+C+E + LL++EYMP GSL+ LF ++ W R K
Sbjct: 215 WLAEIIFLGQLRHPHLVKLIGYCYEDEHRLLMYEYMPRGSLENQLFRRYSAAMPWSTRMK 394
Query: 468 VALGVANALRYLHDDAEQCVLHRDIKSANVLLDTDFSTKLGDFGMAKLVDPMLRTQ-RTD 526
+ALG A L +LH +A++ V++RD K++N+LLD+DF+ KL DFG+AK T T
Sbjct: 395 IALGAAKGLAFLH-EADKPVIYRDFKASNILLDSDFTAKLSDFGLAKDGPEGEDTHVTTR 571
Query: 527 VVGTYGYLAPEYINGGRASKESDMYSFGIVALELATGRRFFQDGDFH--VPLMNWVWGLY 584
++GT GY APEYI G + +SD+YS+G+V LEL TGRR + L+ W L
Sbjct: 572 IMGTQGYAAPEYIMTGHLTTKSDVYSYGVVLLELLTGRRVVDKSRSNEGKSLVEWARPLL 751
Query: 585 -VEGNLMCAADEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKVLQ 635
+ + D +L +F + + ++ C + RP +VIKVL+
Sbjct: 752 RDQKKVYNIIDRRLEGQFPMKGAMKVAMLAFKCLSHHPNARPTMSDVIKVLE 907
>TC231087 similar to UP|Q39203 (Q39203) Receptor-like protein kinase
precursor, partial (30%)
Length = 826
Score = 182 bits (461), Expect = 6e-46
Identities = 101/280 (36%), Positives = 158/280 (56%), Gaps = 19/280 (6%)
Frame = +1
Query: 373 LGRGGYGQVYKGALSYLGKIVAVKRIFADFENSERVFINEVRIISRLIHRNLVQFIGWCH 432
+G GG+G V++G LS +VAVKR+ E+ F EV I + H NLV+ G+C
Sbjct: 1 VGHGGFGTVFQGELSD-ASVVAVKRLERP-GGGEKEFRAEVSTIGNIQHVNLVRLRGFCS 174
Query: 433 EQDELLLVFEYMPNGSLDTHLFGDKKSLAWEVRYKVALGVANALRYLHDDAEQCVLHRDI 492
E LLV+EYM NG+L+ +L + L+W+VR++VA+G A + YLH++ C++H DI
Sbjct: 175 ENSHRLLVYEYMQNGALNVYLRKEGPCLSWDVRFRVAVGTAKGIAYLHEECRCCIIHCDI 354
Query: 493 KSANVLLDTDFSTKLGDFGMAKLVDPMLRTQRTDVVGTYGYLAPEYINGGRASKESDMYS 552
K N+LLD DF+ K+ DFG+AKL+ + GT+GY+APE+I+G + ++D+YS
Sbjct: 355 KPENILLDGDFTAKVSDFGLAKLIGRDFSRVLVTMRGTWGYVAPEWISGVAITTKADVYS 534
Query: 553 FGIVALELATGRRFF-------------QDGD------FHVPLMNWVWGLYVEGNLMCAA 593
+G+ LEL GRR + GD F P W +EGN+
Sbjct: 535 YGMTLLELIGGRRNVEAPLSAGGGGGGGESGDEMGGKWFFPP---WAAQRIIEGNVSDVM 705
Query: 594 DEKLNMEFDVSEMKSLLIVGLWCTHSNDKERPKAYEVIKV 633
D++L +++ E + + +V +WC ++ RP V+K+
Sbjct: 706 DKRLGNAYNIEEARRVALVAVWCIQDDEAMRPTMGMVVKM 825
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.320 0.139 0.413
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 27,639,171
Number of Sequences: 63676
Number of extensions: 371495
Number of successful extensions: 3832
Number of sequences better than 10.0: 987
Number of HSP's better than 10.0 without gapping: 2998
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3043
length of query: 678
length of database: 12,639,632
effective HSP length: 104
effective length of query: 574
effective length of database: 6,017,328
effective search space: 3453946272
effective search space used: 3453946272
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 62 (28.5 bits)
Medicago: description of AC143339.11


