

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC142143.3 + phase: 0 /pseudo
(212 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
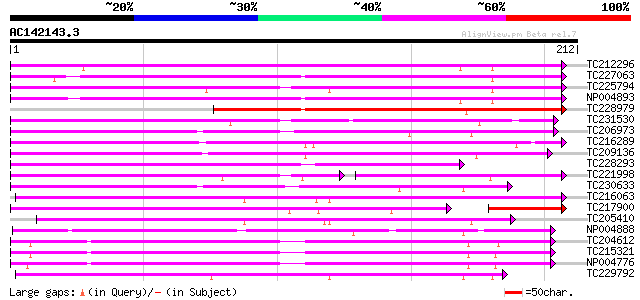
Score E
Sequences producing significant alignments: (bits) Value
TC212296 UP|C933_SOYBN (O81973) Cytochrome P450 93A3 (P450 CP5)... 137 4e-33
TC227063 UP|C931_SOYBN (Q42798) Cytochrome P450 93A1 , complete 129 1e-30
TC225794 UP|Q9XHC6 (Q9XHC6) Cytochrome P450 monooxygenaseCYP93D1... 127 5e-30
NP004893 cytochrome P-450 (CYP93A2) 124 4e-29
TC228979 weakly similar to UP|O23389 (O23389) Cytochrome P450 li... 108 2e-24
TC231530 homologue to UP|C7DA_SOYBN (O48923) Cytochrome P450 71D... 99 2e-21
TC206973 UP|C7D8_SOYBN (O81974) Cytochrome P450 71D8 (P450 CP7)... 97 5e-21
TC216289 UP|Q8W3Y5 (Q8W3Y5) Flavonoid 3'-hydroxylase, complete 96 1e-20
TC209136 similar to UP|Q9FQL9 (Q9FQL9) Cytochrome P450, partial ... 95 2e-20
TC228293 weakly similar to UP|Q8W228 (Q8W228) Cytochrome P450, p... 94 5e-20
TC221998 weakly similar to UP|C727_ARATH (Q96514) Cytochrome P45... 73 6e-20
TC230633 similar to UP|C7D8_SOYBN (O81974) Cytochrome P450 71D8 ... 93 8e-20
TC216063 UP|C822_SOYBN (O81972) Cytochrome P450 82A2 (P450 CP4)... 93 8e-20
TC217900 UP|O48924 (O48924) CYP83D1p (Fragment), complete 87 1e-18
TC205410 similar to UP|C824_SOYBN (O49859) Cytochrome P450 82A4 ... 88 3e-18
NP004888 putative cytochrome P450 87 6e-18
TC204612 UP|Q9SWR5 (Q9SWR5) Cytochrome P450 monooxygenase CYP93C... 84 4e-17
TC215321 UP|Q9M6D6 (Q9M6D6) Isoflavone synthase 1, complete 83 8e-17
NP004776 cytochrome P450 H2O2-dependent urate-degrading peroxidase 82 1e-16
TC229792 UP|C824_SOYBN (O49859) Cytochrome P450 82A4 (P450 CP9)... 82 2e-16
>TC212296 UP|C933_SOYBN (O81973) Cytochrome P450 93A3 (P450 CP5) , complete
Length = 1646
Score = 137 bits (345), Expect = 4e-33
Identities = 76/215 (35%), Positives = 127/215 (58%), Gaps = 7/215 (3%)
Frame = +1
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSY-FITAPYGPYWRFMKKICVTKLLSSSQLGRFMHI 59
+KTH+ FS RP + + L G F+ APYGPYW+FMKK+C+++LL L +F+ +
Sbjct: 274 LKTHEPAFSNRPANTVAVETLTYGFQDFLFAPYGPYWKFMKKLCMSELLGGHMLDQFLPV 453
Query: 60 REQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLV 119
R+ E +K +K +L + D G +F TL+NNI+ RM + T + + E ++ LV
Sbjct: 454 RQXETKKFIKRVLQKGISGEAVDFGGEFITLSNNIVSRMIVSQTSTTEDENEVEEMRKLV 633
Query: 120 REFMHVGAKLSMGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKE-HEERINTEDCQ 178
++ + K ++ + + L +FDL G+ K+L KI FD +L+ +K+ EER N +
Sbjct: 634 KDAAELSGKFNISDFVSFLKRFDLQGFNKRLEKIRDCFDTVLDRIIKQREEERRNKNETV 813
Query: 179 G-----DMMDILLQVYRNENAEVRLTRNDIKAFFL 208
G DM+D+L + +E++E++L + +IKAF L
Sbjct: 814 GKREFKDMLDVLFDISEDESSEIKLNKENIKAFIL 918
>TC227063 UP|C931_SOYBN (Q42798) Cytochrome P450 93A1 , complete
Length = 1769
Score = 129 bits (324), Expect = 1e-30
Identities = 70/222 (31%), Positives = 124/222 (55%), Gaps = 14/222 (6%)
Frame = +1
Query: 1 MKTHDLNFSYRPQFG--------SSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQ 52
+KTH++NFS RP S D+L F AP+GPYW+FMKK+C+++LLS
Sbjct: 307 LKTHEINFSNRPGQNVAVKGLAYDSQDFL-----FAFAPFGPYWKFMKKLCMSELLSGRM 471
Query: 53 LGRFMHIREQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDP 112
+ +F+ +R+QE ++ + + + D G + TL+NNI+ RM++ + N
Sbjct: 472 MDQFLPVRQQETKRFISRVFRKGVAGEAVDFGDELMTLSNNIVSRMTLSQKTSENDN-QA 648
Query: 113 EVIQCLVREFMHVGAKLSMGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERI 172
E ++ LV + K ++ + + L FDL G+ +K+++ FD +++G +K+ +E
Sbjct: 649 EEMKKLVSNIAELMGKFNVSDFIWYLKPFDLQGFNRKIKETRDRFDVVVDGIIKQRQEER 828
Query: 173 NTEDCQG------DMMDILLQVYRNENAEVRLTRNDIKAFFL 208
G DM+D+LL ++ +ENAE++L + +IKAF +
Sbjct: 829 RKNKETGTAKQFKDMLDVLLDMHEDENAEIKLDKKNIKAFIM 954
>TC225794 UP|Q9XHC6 (Q9XHC6) Cytochrome P450 monooxygenaseCYP93D1, complete
Length = 1905
Score = 127 bits (318), Expect = 5e-30
Identities = 80/216 (37%), Positives = 116/216 (53%), Gaps = 8/216 (3%)
Frame = +3
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
+KT + F RP +S Y + + PYG YWRF+KK+C+T+LLS L F+ IR
Sbjct: 333 LKTSEEAFCNRPLMIASESLTYGAADYFFIPYGTYWRFLKKLCMTELLSGKTLEHFVRIR 512
Query: 61 EQELEKLLKSLL-ICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCL- 118
E E+E LK ++ I N N + + T TNNI+ RM MG KK N + + + L
Sbjct: 513 ESEVEAFLKRMMEISGNGNYEVVMRKELITHTNNIITRMIMG----KKSNAENDEVARLR 680
Query: 119 --VREFMHVGAKLSMGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINTED 176
VRE + ++G+V+G + DL G+GKK + + D ++E L+EHEE ED
Sbjct: 681 KVVREVGELLGAFNLGDVIGFMRPLDLQGFGKKNMETHHKVDAMMEKVLREHEEARAKED 860
Query: 177 CQG----DMMDILLQVYRNENAEVRLTRNDIKAFFL 208
D+ DILL + + A+ +LTR KAF L
Sbjct: 861 ADSDRKKDLFDILLNLIEADGADNKLTRESAKAFAL 968
>NP004893 cytochrome P-450 (CYP93A2)
Length = 1509
Score = 124 bits (310), Expect = 4e-29
Identities = 69/214 (32%), Positives = 125/214 (58%), Gaps = 6/214 (2%)
Frame = +1
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
+KTH++NFS RP + +L ++ PYGP +F+KK+C+++LL L +F+ +R
Sbjct: 256 LKTHEINFSNRPGQNVAVQFLT----YVFGPYGPSVKFIKKLCMSELLGGRMLDQFLPVR 423
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVR 120
+QE +K +K +L + D G +F L+NNI+ RM+M T + E ++ LV
Sbjct: 424 QQETKKFIKRVLQKGIAGEAVDFGGEFMRLSNNIISRMTMNQTSSEDEK-QAEEMRMLVA 600
Query: 121 EFMHVGAKLSMGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKE-HEERINTEDCQG 179
+ + ++ + + L FDL G+ K++RK FD +L+ +K+ EER N ++ G
Sbjct: 601 DVAELMGTFNVSDFIWFLKPFDLQGFNKRIRKTRIRFDAVLDRIIKQREEERRNNKEIGG 780
Query: 180 -----DMMDILLQVYRNENAEVRLTRNDIKAFFL 208
D++D+LL + ++++E++LT+ +IKAF +
Sbjct: 781 TRQFKDILDVLLDIGEDDSSEIKLTKENIKAFIM 882
>TC228979 weakly similar to UP|O23389 (O23389) Cytochrome P450 like protein,
partial (26%)
Length = 1207
Score = 108 bits (269), Expect = 2e-24
Identities = 59/136 (43%), Positives = 83/136 (60%), Gaps = 4/136 (2%)
Frame = +2
Query: 77 ENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVREFMHVGAKLSMGEVLG 136
E + DLG +FT TNN+ CR +M T+C +K D E I+ LV+E + AKL G+VLG
Sbjct: 8 ETVALDLGSEFTKFTNNVTCRTAMSTSCAEKCE-DAERIRKLVKESFELAAKLCFGDVLG 184
Query: 137 PLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHE----ERINTEDCQGDMMDILLQVYRNE 192
P + + YGKK + +D++LE LKEHE R N + + D+MDILL VY +
Sbjct: 185 PFKELSFWVYGKKAIDMSTRYDELLEEVLKEHEHKRLSRANGDQSERDLMDILLDVYHDA 364
Query: 193 NAEVRLTRNDIKAFFL 208
+AE ++T IKAFF+
Sbjct: 365 HAEFKITMAHIKAFFM 412
>TC231530 homologue to UP|C7DA_SOYBN (O48923) Cytochrome P450 71D10 ,
complete
Length = 1661
Score = 98.6 bits (244), Expect = 2e-21
Identities = 65/210 (30%), Positives = 111/210 (51%), Gaps = 5/210 (2%)
Frame = +1
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
MKTHDLNFS RP F S Y GS + + +G YWR ++KIC +LL++ ++ F IR
Sbjct: 289 MKTHDLNFSDRPDFVLSRIVSYNGSGIVFSQHGDYWRQLRKICTVELLTAKRVQSFRSIR 468
Query: 61 EQELEKLLKSLLICSNENRST--DLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCL 118
E+E+ +L+K + ++E + +L ++T I R + G KK I +
Sbjct: 469 EEEVAELVKKIAATASEEGGSIFNLTQSIYSMTFGIAARAAFG----KKSRYQQVFISNM 636
Query: 119 VREFMHVGAKLSMGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINT---E 175
++ M +G S+ ++ F + G KL K+ D++L+ + EH+ R +
Sbjct: 637 HKQLMLLGG-FSVADLYPSSRVFQMMGATGKLEKVHRVTDRVLQDIIDEHKNRNRSSEER 813
Query: 176 DCQGDMMDILLQVYRNENAEVRLTRNDIKA 205
+ D++D+LL+ + +E RLT ++IKA
Sbjct: 814 EAVEDLVDVLLKF--QKESEFRLTDDNIKA 897
>TC206973 UP|C7D8_SOYBN (O81974) Cytochrome P450 71D8 (P450 CP7) , complete
Length = 1800
Score = 97.1 bits (240), Expect = 5e-21
Identities = 68/213 (31%), Positives = 109/213 (50%), Gaps = 8/213 (3%)
Frame = +2
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
MKTHD++F RPQ + +Y + APYG YWR ++KIC +LLS+ ++ F HIR
Sbjct: 350 MKTHDVHFVQRPQLLAPQFMVYGATDIAFAPYGDYWRQIRKICTLELLSAKRVQSFSHIR 529
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVR 120
+ E +KL++S I S+ DL +L + R + G K N D + LVR
Sbjct: 530 QDENKKLIQS--IHSSAGSPIDLSGKLFSLLGTTVSRAAFG-----KENDDQDEFMSLVR 688
Query: 121 EFMHVGAKLSMGEVLGPLGKFDLFGYGK-KLRKIVGEFDKILEGFLKEHEER-------I 172
+ + + + ++ L L K K+ + DKILE L++H E+
Sbjct: 689 KAITMTGGFEVDDMFPSLKPLHLLTRQKAKVEHVHQRADKILEDILRKHMEKRTRVKEGN 868
Query: 173 NTEDCQGDMMDILLQVYRNENAEVRLTRNDIKA 205
+E Q D++D+LL++ + + EV +T +IKA
Sbjct: 869 GSEAEQEDLVDVLLRLKESGSLEVPMTMENIKA 967
>TC216289 UP|Q8W3Y5 (Q8W3Y5) Flavonoid 3'-hydroxylase, complete
Length = 1883
Score = 95.9 bits (237), Expect = 1e-20
Identities = 62/214 (28%), Positives = 103/214 (47%), Gaps = 6/214 (2%)
Frame = +2
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
+K HD NFS RP + Y + APYGP WR ++K+ L S + F H+R
Sbjct: 326 LKIHDSNFSSRPPNAGAKYIAYNYQDLVFAPYGPRWRLLRKLTSVHLFSGKAMNEFRHLR 505
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYN--IDP--EVIQ 116
++E+ +L +L S++ ++ +LG T N L R +G F N DP + +
Sbjct: 506 QEEVARLTCNL--ASSDTKAVNLGQLLNVCTTNALARAMIGRRVFNDGNGGCDPRADEFK 679
Query: 117 CLVREFMHVGAKLSMGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINTED 176
+V E M + ++G+ + L DL G K++K+ FD L ++EH + +
Sbjct: 680 AMVMEVMVLAGVFNIGDFIPSLEWLDLQGVQAKMKKLHKRFDAFLTSIIEEHNNSSSKNE 859
Query: 177 CQGDMMDILLQV--YRNENAEVRLTRNDIKAFFL 208
+ + ILL + R+++ LT +IKA L
Sbjct: 860 NHKNFLSILLSLKDVRDDHGN-HLTDTEIKALLL 958
>TC209136 similar to UP|Q9FQL9 (Q9FQL9) Cytochrome P450, partial (63%)
Length = 1028
Score = 95.1 bits (235), Expect = 2e-20
Identities = 58/208 (27%), Positives = 112/208 (52%), Gaps = 5/208 (2%)
Frame = +3
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
+KTHD + RP+F + Y S + YGPYWR +++C+ +L + +L + +I+
Sbjct: 315 LKTHDATLAGRPKFAAGKYTTYNYSDITWSKYGPYWRQARRMCLMELFIAKRLQEY*YIK 494
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYN---IDPEVIQC 117
+QEL LL L ++ N++ L ++L+ N++ RM +G ++ + P+ +
Sbjct: 495 KQELHCLLNELF--NSANKTILLKDHLSSLSLNVISRMVLGKKYLEESQNAVVSPDEFKK 668
Query: 118 LVREFMHVGAKLSMGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERIN--TE 175
++ E + + ++G+ + + DL GY K+++ + +FD +E L EH ER +
Sbjct: 669 MLDELILLNGVYNIGDFIPWIDFLDLQGYIKRMKTLSKKFDMCMEHVLNEHIERKKGIKD 848
Query: 176 DCQGDMMDILLQVYRNENAEVRLTRNDI 203
DM+D+LLQ+ + EV+L R+ +
Sbjct: 849 YVA*DMVDVLLQLDEDPTLEVKLERHGV 932
>TC228293 weakly similar to UP|Q8W228 (Q8W228) Cytochrome P450, partial (40%)
Length = 905
Score = 94.0 bits (232), Expect = 5e-20
Identities = 53/170 (31%), Positives = 92/170 (53%)
Frame = +3
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
+KTHD F+ RP+ +S Y + YGPYWR +KK+C T+LLS+S++ F +R
Sbjct: 393 LKTHDTIFASRPKTLASEYMSYGSKGLAFSEYGPYWRNVKKLCTTQLLSASKVEMFAPLR 572
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVR 120
+EL +KSL + +L L +NI+CRM +G + +++ ++ L R
Sbjct: 573 REELGVFVKSLEKAAASRDVVNLSEQVGELISNIVCRMILGRSKDDRFD-----LKGLAR 737
Query: 121 EFMHVGAKLSMGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEE 170
E + + ++ + + G DL G K++K+ FD++ E +K+HE+
Sbjct: 738 EVLRLAGVFNIADYVPWTGFLDLQGLKGKIKKMSKAFDEVFEQIIKDHED 887
>TC221998 weakly similar to UP|C727_ARATH (Q96514) Cytochrome P450 71B7 ,
partial (20%)
Length = 1123
Score = 72.8 bits (177), Expect(2) = 6e-20
Identities = 42/129 (32%), Positives = 72/129 (55%), Gaps = 4/129 (3%)
Frame = +1
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
+KTHDL F+ RP + ++ A YG YWR M+K+C +LLS +++ F +R
Sbjct: 325 LKTHDLVFASRPPHEAIKYIAWEQRNLGFAEYGSYWRNMRKMCTLELLSQTKINSFRIVR 504
Query: 61 EQELEKLLKSLLICSNEN-RSTDLGLDFTTLTNNILCRMSMGTTCFKKY---NIDPEVIQ 116
E+EL+ +K L SN+ + D+ ++ ++ CRM +G KKY ++D + +
Sbjct: 505 EEELDLSIKLLREASNDGAAAVDISAKVARISADVACRMVLG----KKYMDQDLDEKGFK 672
Query: 117 CLVREFMHV 125
+V+E MH+
Sbjct: 673 AVVQEVMHL 699
Score = 41.2 bits (95), Expect(2) = 6e-20
Identities = 23/80 (28%), Positives = 44/80 (54%), Gaps = 1/80 (1%)
Frame = +2
Query: 130 SMGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERINTEDCQ-GDMMDILLQV 188
++G+ + +GK DL G K+++ + FD+ + + EH + +D + D +D++L
Sbjct: 713 NIGDYIPYIGKLDLQGLIKRMKAVRKIFDEFFDKIIDEHIQSDKGQDNKVKDFVDVMLGF 892
Query: 189 YRNENAEVRLTRNDIKAFFL 208
+E E R+ R +IKA L
Sbjct: 893 VGSEEYEYRIERPNIKAILL 952
>TC230633 similar to UP|C7D8_SOYBN (O81974) Cytochrome P450 71D8 (P450 CP7)
, partial (56%)
Length = 876
Score = 93.2 bits (230), Expect = 8e-20
Identities = 65/197 (32%), Positives = 97/197 (48%), Gaps = 9/197 (4%)
Frame = +3
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
MKTHDL F RPQ + Y + APYG YWR M+KIC +LLS+ ++ F HIR
Sbjct: 300 MKTHDLAFVQRPQLLAPQYMAYGATDIAFAPYGEYWRQMRKICTLELLSAKRVQSFSHIR 479
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVR 120
+ E KL++S I S+ DL +L + R + G N D + LVR
Sbjct: 480 QDENRKLIQS--IQSSAGSPIDLSSKLFSLLGTTVSRAAFGNK-----NDDQDEFMSLVR 638
Query: 121 EFMHVGAKLSMGEVLGPLGKFDLF-GYGKKLRKIVGEFDKILEGFLKEH--------EER 171
+ + + + ++ L L G K+ +I D+ILE L++H EE
Sbjct: 639 KAVAMTGGFELDDMFPSLKPLHLLTGQKAKVEEIHKRADRILEDILRKHVEKRTRAKEEG 818
Query: 172 INTEDCQGDMMDILLQV 188
N+E Q D++D+LL++
Sbjct: 819 NNSEAQQEDLVDVLLRI 869
>TC216063 UP|C822_SOYBN (O81972) Cytochrome P450 82A2 (P450 CP4) , complete
Length = 1766
Score = 93.2 bits (230), Expect = 8e-20
Identities = 62/216 (28%), Positives = 105/216 (47%), Gaps = 10/216 (4%)
Frame = +2
Query: 3 THDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIREQ 62
T+D+ S P S++ Y S + APYGPYWR ++KI +++ LS S++ + H+R
Sbjct: 311 TNDIAVSSLPDLISANLLCYNRSMIVVAPYGPYWRQLRKILMSEFLSPSRVEQLHHVRVS 490
Query: 63 ELEKLLKSLLICSNENRSTDLGLD-------FTTLTNNILCRMSMGTTCFKKYNIDPE-V 114
E++ + L N++ G F+ L N++ RM G F D E
Sbjct: 491 EVQSSITELFRDWRSNKNVQSGFATVELKQWFSLLVFNMILRMVCGKRYFSASTSDDEKA 670
Query: 115 IQCL--VREFMHVGAKLSMGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEERI 172
+C+ V EF+ + A ++G+ + L FD GY +R+ E D+I+ +L EH ++
Sbjct: 671 NRCVKAVDEFVRLAATFTVGDAIPYLRWFDFGGYENDMRETGKELDEIIGEWLDEHRQKR 850
Query: 173 NTEDCQGDMMDILLQVYRNENAEVRLTRNDIKAFFL 208
+ D+M +LL + + E IK+F L
Sbjct: 851 KMGENVQDLMSVLLSLLEGKTIEGMNVDIVIKSFVL 958
>TC217900 UP|O48924 (O48924) CYP83D1p (Fragment), complete
Length = 1672
Score = 87.0 bits (214), Expect(2) = 1e-18
Identities = 55/176 (31%), Positives = 88/176 (49%), Gaps = 11/176 (6%)
Frame = +1
Query: 1 MKTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIR 60
+KTHDLNF+ RP F Y G APYGPYWR MKK+C+ L S+ ++ F IR
Sbjct: 259 LKTHDLNFASRPLFVGPRKLSYDGLDMGFAPYGPYWREMKKLCIVHLFSAQRVRSFRPIR 438
Query: 61 EQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTT--CFKKYNIDPEV---- 114
E E+ K+++ L +L + TN+++CR+++G + C + + EV
Sbjct: 439 ENEVAKMVRKLSEHEASGTVVNLTETLMSFTNSLICRIALGKSYGCEYEEVVVDEVLGNR 618
Query: 115 ---IQCLVREFMHVGAKLSMGEVLGPLGKF--DLFGYGKKLRKIVGEFDKILEGFL 165
+Q L+ E + ++ + P+GK+ + G +L K E D E F+
Sbjct: 619 RSRLQVLLNEAQALLSEFFFSDYFPPIGKWVDRVTGILSRLDKTFKELDACYERFI 786
Score = 22.3 bits (46), Expect(2) = 1e-18
Identities = 11/29 (37%), Positives = 18/29 (61%)
Frame = +2
Query: 180 DMMDILLQVYRNENAEVRLTRNDIKAFFL 208
D++DILLQ+ + + LT + IKA +
Sbjct: 848 DIIDILLQLLDDRSFTFDLTLDHIKAVLM 934
>TC205410 similar to UP|C824_SOYBN (O49859) Cytochrome P450 82A4 (P450 CP9)
, partial (73%)
Length = 1401
Score = 88.2 bits (217), Expect = 3e-18
Identities = 56/193 (29%), Positives = 99/193 (51%), Gaps = 14/193 (7%)
Frame = +3
Query: 11 RPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIREQELEKLLKS 70
RP+ + Y + F APYGPYWR +KI ++L+S ++ + H+R E++ +K
Sbjct: 6 RPKLLAIELMCYNQAMFGFAPYGPYWREQRKITTLEILTSRRVEQLQHVRVSEVQSSIKE 185
Query: 71 LLICSNENRSTDLGLD-------FTTLTNNILCRMSMGTTCFKKYNIDPEVIQ-CL--VR 120
L + N++ + G F+ LT N++ RM +G F +D E Q C+ V+
Sbjct: 186 LFNVWSSNKNNESGYALLELKQWFSQLTYNMVLRMVVGKRLFGARTMDDEKAQRCVKAVK 365
Query: 121 EFMHVGAKLSMGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEER----INTED 176
EFM + ++ + + L FD GY K +++ + D+I +L+EH++ N D
Sbjct: 366 EFMRLMGVFTVADAIPFLRWFDFGGYEKAMKETAKDLDEIFGEWLEEHKQNRAFGENNVD 545
Query: 177 CQGDMMDILLQVY 189
D MD++L ++
Sbjct: 546 GIQDFMDVMLSLF 584
>NP004888 putative cytochrome P450
Length = 1500
Score = 87.0 bits (214), Expect = 6e-18
Identities = 67/210 (31%), Positives = 107/210 (50%), Gaps = 7/210 (3%)
Frame = +1
Query: 2 KTHDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIRE 61
K HD FS RP +++ Y GS APYG YWR M+KI + +LLS ++ F +R
Sbjct: 271 KNHDSVFSGRPSLYAANRLGY-GSTVSFAPYGEYWREMRKIMILELLSPKRVQSFEAVRF 447
Query: 62 QELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLVRE 121
+E++ LL+++ + ++L L +LTNNI+CR+++G + +V + L
Sbjct: 448 EEVKLLLQTIALSHGPVNLSELTL---SLTNNIVCRIALGKRNRSGADDANKVSEMLKET 618
Query: 122 FMHVGA--KLSMGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEH-----EERINT 174
+G + LG L KF G +L KI E D + +KEH ER
Sbjct: 619 QAMLGGFFPVDFFPRLGWLNKFS--GLENRLEKIFREMDNFYDQVIKEHIADNSSERSGA 792
Query: 175 EDCQGDMMDILLQVYRNENAEVRLTRNDIK 204
E D++D+LL+V ++ N + +T + IK
Sbjct: 793 E--HEDVVDVLLRVQKDPNQAIAITDDQIK 876
>TC204612 UP|Q9SWR5 (Q9SWR5) Cytochrome P450 monooxygenase CYP93C1v2p,
complete
Length = 1912
Score = 84.3 bits (207), Expect = 4e-17
Identities = 53/213 (24%), Positives = 111/213 (51%), Gaps = 9/213 (4%)
Frame = +3
Query: 1 MKTHDL-NFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHI 59
++TH+ +F+ R Q + Y S + P+GPYW+F++K+ + LL+++ + + +
Sbjct: 351 LQTHEATSFNTRFQTSAIRRLTYDSSVAMV-PFGPYWKFVRKLIMNDLLNATTVNKLRPL 527
Query: 60 REQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLV 119
R Q++ K L+ + + + DL + TN+ + M +G + E I+ +
Sbjct: 528 RTQQIRKFLRVMAQGAEAQKPLDLTEELLKWTNSTISMMMLG---------EAEEIRDIA 680
Query: 120 REFMHVGAKLSMGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEE----RINTE 175
RE + + + S+ + + PL + Y K++ I+ +FD ++E +K+ E R N E
Sbjct: 681 REVLKIFGEYSLTDFIWPLKHLKVGKYEKRIDDILNKFDPVVERVIKKRREIVRRRKNGE 860
Query: 176 DCQGDM----MDILLQVYRNENAEVRLTRNDIK 204
+G++ +D LL+ +E E+++T++ IK
Sbjct: 861 VVEGEVSGVFLDTLLEFAEDETMEIKITKDHIK 959
>TC215321 UP|Q9M6D6 (Q9M6D6) Isoflavone synthase 1, complete
Length = 1888
Score = 83.2 bits (204), Expect = 8e-17
Identities = 52/213 (24%), Positives = 109/213 (50%), Gaps = 9/213 (4%)
Frame = +1
Query: 1 MKTHDL-NFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHI 59
++TH+ +F+ R Q + Y S + P+GPYW+F++K+ + LL+++ + + +
Sbjct: 358 LQTHEATSFNTRFQTSAIRRLTYDNSVAMV-PFGPYWKFVRKLIMNDLLNATTVNKLRPL 534
Query: 60 REQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLV 119
R Q++ K L+ + + + D+ + TN+ + M +G + E I+ +
Sbjct: 535 RTQQIRKFLRVMAQSAEAQKPLDVTEELLKWTNSTISMMMLG---------EAEEIRDIA 687
Query: 120 REFMHVGAKLSMGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEE----RINTE 175
RE + + + S+ + + PL + Y K++ I+ +FD ++E +K+ E R N E
Sbjct: 688 REVLKIFGEYSLTDFIWPLKYLKVGKYEKRIDDILNKFDPVVERVIKKRREIVRRRKNGE 867
Query: 176 DCQGD----MMDILLQVYRNENAEVRLTRNDIK 204
+G+ +D LL+ +E E+++T+ IK
Sbjct: 868 VVEGEASGVFLDTLLEFAEDETMEIKITKEQIK 966
>NP004776 cytochrome P450 H2O2-dependent urate-degrading peroxidase
Length = 1536
Score = 82.4 bits (202), Expect = 1e-16
Identities = 52/213 (24%), Positives = 109/213 (50%), Gaps = 9/213 (4%)
Frame = +1
Query: 1 MKTHD-LNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHI 59
++TH+ +F+ R Q + Y S + P+GPYW+F++K+ + LL+++ + +
Sbjct: 277 LQTHEGTSFNTRFQTSAIRRLTYDNSVAMV-PFGPYWKFVRKLIMNDLLNATTDNKLRPL 453
Query: 60 REQELEKLLKSLLICSNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQCLV 119
R Q++ K L+ + + + D+ + TN+ + M +G + E+I+ +
Sbjct: 454 RTQQIRKFLRVMAQSAEAQKPLDVTEELLKWTNSTISMMMLG---------EAEMIRDIA 606
Query: 120 REFMHVGAKLSMGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEHEE----RINTE 175
RE + + + S+ + + PL + Y K++ I+ +FD ++E +K+ E R N E
Sbjct: 607 REVLKIFGEYSLTDFIWPLKYLKVGKYEKRIDDILNKFDPVVERVIKKRREIVRRRKNGE 786
Query: 176 DCQGD----MMDILLQVYRNENAEVRLTRNDIK 204
+G+ +D LL+ +E E+++T+ IK
Sbjct: 787 VVEGEASGVFLDTLLEFAEDETMEIKITKEQIK 885
>TC229792 UP|C824_SOYBN (O49859) Cytochrome P450 82A4 (P450 CP9) , complete
Length = 1769
Score = 81.6 bits (200), Expect = 2e-16
Identities = 55/195 (28%), Positives = 98/195 (50%), Gaps = 11/195 (5%)
Frame = +3
Query: 3 THDLNFSYRPQFGSSHDYLYKGSYFITAPYGPYWRFMKKICVTKLLSSSQLGRFMHIREQ 62
T+D+ S RP+ + Y + + APYGPYWR ++KI VT++LSSS++ + +R
Sbjct: 291 TNDVAVSARPKLLVAELMCYNNAMLLVAPYGPYWRELRKIIVTEILSSSRVEQLQDVRVS 470
Query: 63 ELEKLLKSLLIC------SNENRSTDLGLDFTTLTNNILCRMSMGTTCFKKYNIDPEVIQ 116
E++ + L ++ S +L F N++ RM +G D + +
Sbjct: 471 EVQNSIVELYDVWRSQKNESDYASVELKQWFAQPIFNMVLRMVVGKRFLSATATDEKAEK 650
Query: 117 CL--VREFMHVGAKLSMGEVLGPLGKFDLFGYGKKLRKIVGEFDKILEGFLKEH-EERIN 173
C+ V EFM + ++G+ + L D GY K +++ E D ++ +L+EH ++R
Sbjct: 651 CVKAVDEFMRLAGVFTVGDAIPYLRWLDFGGYEKAMKETAKELDVMISEWLEEHRQKRAL 830
Query: 174 TEDCQG--DMMDILL 186
E G D M+++L
Sbjct: 831 GEGVDGAQDFMNVML 875
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.325 0.142 0.428
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,908,997
Number of Sequences: 63676
Number of extensions: 142329
Number of successful extensions: 864
Number of sequences better than 10.0: 115
Number of HSP's better than 10.0 without gapping: 840
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 841
length of query: 212
length of database: 12,639,632
effective HSP length: 93
effective length of query: 119
effective length of database: 6,717,764
effective search space: 799413916
effective search space used: 799413916
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 57 (26.6 bits)
Medicago: description of AC142143.3


