

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC142143.13 - phase: 0
(259 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
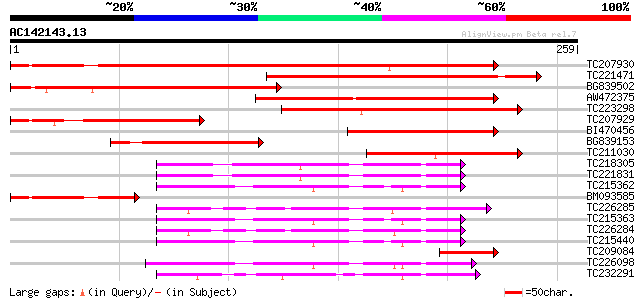
Score E
Sequences producing significant alignments: (bits) Value
TC207930 similar to UP|O49459 (O49459) Predicted protein, partia... 361 e-100
TC221471 similar to UP|O49459 (O49459) Predicted protein, partia... 204 3e-53
BG839502 similar to GP|2842493|emb| predicted protein {Arabidops... 204 4e-53
AW472375 156 1e-38
TC223298 similar to UP|Q9SS95 (Q9SS95) F4P13.14 protein, partial... 128 3e-30
TC207929 similar to UP|O49459 (O49459) Predicted protein, partia... 114 4e-26
BI470456 100 1e-21
BG839153 91 7e-19
TC211030 similar to UP|Q6NQK2 (Q6NQK2) At1g25580, partial (8%) 82 2e-16
TC218305 similar to UP|Q8LJS1 (Q8LJS1) No apical meristem-like p... 69 2e-12
TC221831 UP|Q8LJS1 (Q8LJS1) No apical meristem-like protein, com... 66 1e-11
TC215362 homologue to UP|Q8LRL5 (Q8LRL5) Nam-like protein 10, pa... 63 1e-10
BM093585 63 2e-10
TC226285 similar to UP|Q9FY93 (Q9FY93) NAM-like protein (AT5g131... 63 2e-10
TC215363 homologue to UP|Q8LRL5 (Q8LRL5) Nam-like protein 10, pa... 62 3e-10
TC226284 similar to UP|Q9FY93 (Q9FY93) NAM-like protein (AT5g131... 60 8e-10
TC215440 homologue to UP|Q7Y1A6 (Q7Y1A6) NAC-domain protein 18, ... 59 2e-09
TC209084 homologue to UP|O49459 (O49459) Predicted protein, part... 59 2e-09
TC226098 similar to UP|Q84TD6 (Q84TD6) At3g04070, partial (45%) 59 2e-09
TC232291 similar to UP|O81790 (O81790) NAM / CUC2-like protein, ... 59 2e-09
>TC207930 similar to UP|O49459 (O49459) Predicted protein, partial (62%)
Length = 1474
Score = 361 bits (926), Expect = e-100
Identities = 176/225 (78%), Positives = 191/225 (84%), Gaps = 2/225 (0%)
Frame = +3
Query: 1 MTWCNDSDDQDRGIELITSTVIPGTSNESPLVTDDETKPTEVRTLTCPSCGHNIEFQDQG 60
MTWCNDSD+ DR IE T T+ + + +T+ E+RT+TCPSCGHNIEF+DQG
Sbjct: 87 MTWCNDSDE-DRVIESTTPTITITPNTDGIKLTE------EIRTITCPSCGHNIEFKDQG 245
Query: 61 GINELPGLPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHPEK 120
GI++LPGLPAGVKFDPNDQEIL+HLEAKV D KLHPLIDEFIPTLEGENGICYTHPEK
Sbjct: 246 GIHDLPGLPAGVKFDPNDQEILEHLEAKVASDACKLHPLIDEFIPTLEGENGICYTHPEK 425
Query: 121 LPGVKKDGQIRHFFHRPSKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVI--IDGVMK 178
LPGV KDGQIRHFFHRPSKAYTTGTRKRRKV TD+EGSETRWHKTGKTR V G +K
Sbjct: 426 LPGVSKDGQIRHFFHRPSKAYTTGTRKRRKVHTDDEGSETRWHKTGKTRAVFAAAGGAVK 605
Query: 179 GFKKILVLYTNYGRQKKPEKTNWVMHQYHLGSNEEEKDGELVVSK 223
GFKKILVLYTNYGRQKKPEKT WVMHQYHLG+ EEEKDGELVVSK
Sbjct: 606 GFKKILVLYTNYGRQKKPEKTYWVMHQYHLGNTEEEKDGELVVSK 740
>TC221471 similar to UP|O49459 (O49459) Predicted protein, partial (36%)
Length = 757
Score = 204 bits (519), Expect = 3e-53
Identities = 96/126 (76%), Positives = 109/126 (86%)
Frame = +2
Query: 118 PEKLPGVKKDGQIRHFFHRPSKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVIIDGVM 177
P ++ GV+KDGQIRHFFHRPSKAYTTGTRKRRKV TD+EGSETRWHKTGKTRPV + G +
Sbjct: 80 PGRITGVRKDGQIRHFFHRPSKAYTTGTRKRRKVHTDKEGSETRWHKTGKTRPVFVGGAV 259
Query: 178 KGFKKILVLYTNYGRQKKPEKTNWVMHQYHLGSNEEEKDGELVVSKKILEYLSSSERRLA 237
KGFKKILVLYTNYGRQKKPEKTNWVMHQYHLG++EEEKDGELVVSK + + R+
Sbjct: 260 KGFKKILVLYTNYGRQKKPEKTNWVMHQYHLGTSEEEKDGELVVSK---IFYQTQPRQCG 430
Query: 238 CNSVIR 243
NS+I+
Sbjct: 431 NNSIIK 448
>BG839502 similar to GP|2842493|emb| predicted protein {Arabidopsis
thaliana}, partial (25%)
Length = 593
Score = 204 bits (518), Expect = 4e-53
Identities = 96/126 (76%), Positives = 110/126 (87%), Gaps = 2/126 (1%)
Frame = -1
Query: 1 MTWCNDSDDQDRGIE-LITSTVIPGTSNESPLVTDDE-TKPTEVRTLTCPSCGHNIEFQD 58
MTWCNDSD++ RGI+ L+ + P ++N + ++T D KPTE+RT+TCPSCGHNIEFQD
Sbjct: 512 MTWCNDSDEE-RGIDQLVAPAITPTSTNNTTVLTHDHHRKPTEIRTITCPSCGHNIEFQD 336
Query: 59 QGGINELPGLPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHP 118
Q GIN+LPGLPAGVKFDPNDQEIL+HLEAKV DVPKLHPLIDEFIPTLEGENGICYTHP
Sbjct: 335 QTGINDLPGLPAGVKFDPNDQEILEHLEAKVFSDVPKLHPLIDEFIPTLEGENGICYTHP 156
Query: 119 EKLPGV 124
EKLPGV
Sbjct: 155 EKLPGV 138
>AW472375
Length = 443
Score = 156 bits (394), Expect = 1e-38
Identities = 69/111 (62%), Positives = 86/111 (77%)
Frame = +3
Query: 113 ICYTHPEKLPGVKKDGQIRHFFHRPSKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVI 172
ICYTHP+ LPGVK+DG HFFHR AY TGTRKRRK+ + G + RWHKTG+T+PV+
Sbjct: 3 ICYTHPQNLPGVKQDGSASHFFHRAINAYNTGTRKRRKILGQDFG-DVRWHKTGRTKPVV 179
Query: 173 IDGVMKGFKKILVLYTNYGRQKKPEKTNWVMHQYHLGSNEEEKDGELVVSK 223
+G+ KG KKI+VLY + R + EKTNWVMHQYHLG+ E+EKDGE ++SK
Sbjct: 180 FNGIQKGCKKIMVLYVSNVRGGRAEKTNWVMHQYHLGTEEDEKDGEYIISK 332
>TC223298 similar to UP|Q9SS95 (Q9SS95) F4P13.14 protein, partial (21%)
Length = 438
Score = 128 bits (321), Expect = 3e-30
Identities = 64/112 (57%), Positives = 75/112 (66%), Gaps = 2/112 (1%)
Frame = +2
Query: 125 KKDGQIRHFFHRPSKAYTTGTRKRRKVQTDEEGSE--TRWHKTGKTRPVIIDGVMKGFKK 182
K DG HFFH+ AY TG RKRRK+ + +E RWHKTG+T+ VI DGV KGFKK
Sbjct: 20 KTDGSSVHFFHKTVNAYATGPRKRRKIHHQDGMTEEHVRWHKTGRTKAVIEDGVHKGFKK 199
Query: 183 ILVLYTNYGRQKKPEKTNWVMHQYHLGSNEEEKDGELVVSKKILEYLSSSER 234
I+VLY + KP KTNWVMHQYHLGS E+EKD E VVSK + SE+
Sbjct: 200 IMVLYIRSKKGSKPYKTNWVMHQYHLGSEEDEKDDEYVVSKIFYQRQKQSEK 355
>TC207929 similar to UP|O49459 (O49459) Predicted protein, partial (16%)
Length = 432
Score = 114 bits (285), Expect = 4e-26
Identities = 58/91 (63%), Positives = 68/91 (73%), Gaps = 2/91 (2%)
Frame = +2
Query: 1 MTWCNDSDDQDRGIELITS--TVIPGTSNESPLVTDDETKPTEVRTLTCPSCGHNIEFQD 58
MTWCNDSD+ +R IE IT T+ P T DD+ E+RT CPSCGHNI F+D
Sbjct: 179 MTWCNDSDE-NRVIESITPRITITPNT--------DDKKLTQEIRTTACPSCGHNIAFKD 331
Query: 59 QGGINELPGLPAGVKFDPNDQEILQHLEAKV 89
+GGI++LPGLPAGVKFDPNDQEIL+HLEAKV
Sbjct: 332 KGGIHDLPGLPAGVKFDPNDQEILEHLEAKV 424
>BI470456
Length = 421
Score = 99.8 bits (247), Expect = 1e-21
Identities = 42/69 (60%), Positives = 55/69 (78%)
Frame = +1
Query: 155 EEGSETRWHKTGKTRPVIIDGVMKGFKKILVLYTNYGRQKKPEKTNWVMHQYHLGSNEEE 214
++ + RWHKTG+T+PV++ GV KG KKI+VLY + R K EKTNWVMHQYHLG+ E+E
Sbjct: 1 QDFGDVRWHKTGRTKPVVLSGVQKGCKKIMVLYVSNVRGGKAEKTNWVMHQYHLGTEEDE 180
Query: 215 KDGELVVSK 223
KDGE ++SK
Sbjct: 181 KDGEYIISK 207
>BG839153
Length = 715
Score = 90.5 bits (223), Expect = 7e-19
Identities = 44/70 (62%), Positives = 50/70 (70%)
Frame = +2
Query: 47 CPSCGHNIEFQDQGGINELPGLPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLIDEFIPT 106
CP CGH E G + GLPA VKFDP DQE+++HLEAKV K HPLIDEFIPT
Sbjct: 521 CPGCGHKFE-----GKPDWLGLPAEVKFDPTDQELIEHLEAKVEAKNMKSHPLIDEFIPT 685
Query: 107 LEGENGICYT 116
+EGE+GICYT
Sbjct: 686 IEGEDGICYT 715
>TC211030 similar to UP|Q6NQK2 (Q6NQK2) At1g25580, partial (8%)
Length = 856
Score = 82.0 bits (201), Expect = 2e-16
Identities = 39/72 (54%), Positives = 50/72 (69%), Gaps = 1/72 (1%)
Frame = +1
Query: 164 KTGKTRPVIIDGVMKGFKKILVLYTNYGRQ-KKPEKTNWVMHQYHLGSNEEEKDGELVVS 222
KTGKT+ +I DGV KGFKKI+V+Y KP K+NWVMHQYHLG+ E+EK+ E VVS
Sbjct: 4 KTGKTKAIIEDGVHKGFKKIMVIYIRSSENGSKPYKSNWVMHQYHLGTEEDEKEAEYVVS 183
Query: 223 KKILEYLSSSER 234
K + +E+
Sbjct: 184 KVFYQQQKQTEK 219
>TC218305 similar to UP|Q8LJS1 (Q8LJS1) No apical meristem-like protein,
partial (58%)
Length = 935
Score = 68.9 bits (167), Expect = 2e-12
Identities = 47/142 (33%), Positives = 64/142 (44%), Gaps = 1/142 (0%)
Frame = +3
Query: 68 LPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHPEKLPGVKKD 127
LP G +F P D+E++ H K + D +F GE + + P LP K
Sbjct: 192 LPPGFRFHPTDEELISHYLYKKVIDT--------KFCARAIGEVDLNKSEPWDLPWKAKM 347
Query: 128 GQIR-HFFHRPSKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVIIDGVMKGFKKILVL 186
G+ +FF + Y TG R R + W TGK + + + G KK LV
Sbjct: 348 GEKEWYFFCVRDRKYPTGLRTNRATEAGY------WKATGKDKEIFRGKSLVGMKKTLVF 509
Query: 187 YTNYGRQKKPEKTNWVMHQYHL 208
Y GR K EK+NWVMH+Y L
Sbjct: 510 YR--GRAPKGEKSNWVMHEYRL 569
>TC221831 UP|Q8LJS1 (Q8LJS1) No apical meristem-like protein, complete
Length = 1216
Score = 66.2 bits (160), Expect = 1e-11
Identities = 46/142 (32%), Positives = 64/142 (44%), Gaps = 1/142 (0%)
Frame = +2
Query: 68 LPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHPEKLPGVKKD 127
LP G +F P D+E++ H + + D F GE + + P LP K
Sbjct: 170 LPPGFRFHPTDEELISHYLYRKVTDT--------NFSARAIGEVDLNRSEPWDLPWKAKM 325
Query: 128 GQIR-HFFHRPSKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVIIDGVMKGFKKILVL 186
G+ +FF + Y TG R R ++ W TGK + + + G KK LV
Sbjct: 326 GEKEWYFFCVRDRKYPTGLRTNRATESGY------WKATGKDKEIFRGKSLVGMKKTLVF 487
Query: 187 YTNYGRQKKPEKTNWVMHQYHL 208
Y GR K EKT+WVMH+Y L
Sbjct: 488 YK--GRAPKGEKTDWVMHEYRL 547
>TC215362 homologue to UP|Q8LRL5 (Q8LRL5) Nam-like protein 10, partial (63%)
Length = 1565
Score = 63.2 bits (152), Expect = 1e-10
Identities = 46/144 (31%), Positives = 64/144 (43%), Gaps = 3/144 (2%)
Frame = +3
Query: 68 LPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHPEKLPGVKKD 127
LP G +F P DQE++ H + P P+I E + P LPG+
Sbjct: 150 LPPGFRFHPTDQELVLHYLCRKCASQPIAVPIIAEI--------DLYKYDPWDLPGLASY 305
Query: 128 GQIRHFFHRP-SKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVIIDGVMK--GFKKIL 184
G+ +F P + Y G+R R T W TG +P+ G K G KK L
Sbjct: 306 GEKEWYFFSPRDRKYPNGSRPNRAAGTGY------WKATGADKPI---GHPKPVGIKKAL 458
Query: 185 VLYTNYGRQKKPEKTNWVMHQYHL 208
V Y G+ K +K+NW+MH+Y L
Sbjct: 459 VFYA--GKAPKGDKSNWIMHEYRL 524
>BM093585
Length = 382
Score = 62.8 bits (151), Expect = 2e-10
Identities = 31/59 (52%), Positives = 40/59 (67%)
Frame = +1
Query: 1 MTWCNDSDDQDRGIELITSTVIPGTSNESPLVTDDETKPTEVRTLTCPSCGHNIEFQDQ 59
MTWCNDSD+ DR IE T T+ + + +T+ E+RT+TCPSCGHNIEF+DQ
Sbjct: 187 MTWCNDSDE-DRVIESTTPTITITPNTDGIKLTE------EIRTITCPSCGHNIEFKDQ 342
>TC226285 similar to UP|Q9FY93 (Q9FY93) NAM-like protein
(AT5g13180/T19L5_140), partial (59%)
Length = 1332
Score = 62.8 bits (151), Expect = 2e-10
Identities = 48/157 (30%), Positives = 70/157 (44%), Gaps = 4/157 (2%)
Frame = +2
Query: 68 LPAGVKFDPNDQE-ILQHLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHPEKLPGVKK 126
LP G +F P D+E +LQ+L+ KV PL IP L +C + P LPG +
Sbjct: 476 LPPGFRFHPTDEELVLQYLKRKVFSC-----PLPASIIPELH----VCKSDPWDLPGDLE 628
Query: 127 DGQIRHFFHRPSKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVII---DGVMKGFKKI 183
Q R+FF Y G R R + W TG + ++ + + G KK
Sbjct: 629 --QERYFFSTKVAKYPNGNRSNRATNSGY------WKATGLDKQIVTSKGNNQVVGMKKT 784
Query: 184 LVLYTNYGRQKKPEKTNWVMHQYHLGSNEEEKDGELV 220
LV Y G+ +T+W+MH+Y L N + +V
Sbjct: 785 LVFYR--GKPPNGSRTDWIMHEYRLILNASQSQSHVV 889
>TC215363 homologue to UP|Q8LRL5 (Q8LRL5) Nam-like protein 10, partial (64%)
Length = 1464
Score = 62.0 bits (149), Expect = 3e-10
Identities = 45/144 (31%), Positives = 64/144 (44%), Gaps = 3/144 (2%)
Frame = +1
Query: 68 LPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHPEKLPGVKKD 127
LP G +F P D+E++ H + P P+I E + P LPG+
Sbjct: 79 LPPGFRFHPTDEELVLHYLCRKCASQPIAVPIIAEI--------DLYKYDPWDLPGLATY 234
Query: 128 GQIRHFFHRP-SKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVIIDGVMK--GFKKIL 184
G+ +F P + Y G+R R T W TG +P+ G K G KK L
Sbjct: 235 GEKEWYFFSPRDRKYPNGSRPNRAAGTGY------WKATGADKPI---GQPKPVGIKKAL 387
Query: 185 VLYTNYGRQKKPEKTNWVMHQYHL 208
V Y G+ K +K+NW+MH+Y L
Sbjct: 388 VFYA--GKAPKGDKSNWIMHEYRL 453
>TC226284 similar to UP|Q9FY93 (Q9FY93) NAM-like protein
(AT5g13180/T19L5_140), partial (59%)
Length = 1601
Score = 60.5 bits (145), Expect = 8e-10
Identities = 45/144 (31%), Positives = 66/144 (45%), Gaps = 3/144 (2%)
Frame = +2
Query: 68 LPAGVKFDPNDQE-ILQHLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHPEKLPGVKK 126
LP G +F P D+E +LQ+L+ KV PL IP ++ +C + P LPG +
Sbjct: 215 LPPGFRFHPTDEELVLQYLKRKVFSC-----PLPASIIPEVD----VCKSDPWDLPGDLE 367
Query: 127 DGQIRHFFHRPSKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVIID--GVMKGFKKIL 184
Q R+FF Y G R R + W TG + ++ + G KK L
Sbjct: 368 --QERYFFSTKEAKYPNGNRSNRATNSGY------WKATGLDKQIVTSKGNQVVGMKKTL 523
Query: 185 VLYTNYGRQKKPEKTNWVMHQYHL 208
V Y G+ +T+W+MH+Y L
Sbjct: 524 VFYR--GKPPHGSRTDWIMHEYRL 589
>TC215440 homologue to UP|Q7Y1A6 (Q7Y1A6) NAC-domain protein 18, partial
(70%)
Length = 983
Score = 59.3 bits (142), Expect = 2e-09
Identities = 46/144 (31%), Positives = 62/144 (42%), Gaps = 3/144 (2%)
Frame = +1
Query: 68 LPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHPEKLPGVKKD 127
LP G +F P D+E++ H + P+I E + P LPG+
Sbjct: 91 LPPGFRFHPTDEELVVHYLCRKCASQEIAVPIIAEI--------DLYKYDPWDLPGMALY 246
Query: 128 GQIRHFFHRP-SKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVIIDGVMK--GFKKIL 184
G+ +F P + Y G+R R T W TG +PV G K G KK L
Sbjct: 247 GEKEWYFFTPRDRKYPNGSRPNRSAGTGY------WKATGADKPV---GKPKPVGIKKAL 399
Query: 185 VLYTNYGRQKKPEKTNWVMHQYHL 208
V Y G+ K KTNW+MH+Y L
Sbjct: 400 VFYA--GKAPKGVKTNWIMHEYRL 465
>TC209084 homologue to UP|O49459 (O49459) Predicted protein, partial (12%)
Length = 972
Score = 58.9 bits (141), Expect = 2e-09
Identities = 26/27 (96%), Positives = 26/27 (96%)
Frame = +3
Query: 197 EKTNWVMHQYHLGSNEEEKDGELVVSK 223
EKTNWVMHQYHLGS EEEKDGELVVSK
Sbjct: 6 EKTNWVMHQYHLGSTEEEKDGELVVSK 86
>TC226098 similar to UP|Q84TD6 (Q84TD6) At3g04070, partial (45%)
Length = 1562
Score = 58.9 bits (141), Expect = 2e-09
Identities = 46/157 (29%), Positives = 65/157 (41%), Gaps = 6/157 (3%)
Frame = +1
Query: 63 NELPGLPAGVKFDPNDQEILQHLEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHPEKLP 122
N LP G +F P D+E++ H +K + +P +I E I P LP
Sbjct: 154 NPESNLPPGFRFHPTDEELILHYLSKKVASIPLPVSIIAEV--------DIYKLDPWDLP 309
Query: 123 GVKKDGQIRHFFHRP-SKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVIID---GVMK 178
G+ +F P + Y G R R + W TG + ++ G +
Sbjct: 310 AKATFGEKEWYFFSPRDRKYPNGARPNRAAASGY------WKATGTDKTIVTSLQGGAQE 471
Query: 179 --GFKKILVLYTNYGRQKKPEKTNWVMHQYHLGSNEE 213
G KK LV Y GR K KTNW+MH+Y L N +
Sbjct: 472 SVGVKKALVFYK--GRPPKGVKTNWIMHEYRLVDNNK 576
>TC232291 similar to UP|O81790 (O81790) NAM / CUC2-like protein, partial
(20%)
Length = 716
Score = 58.9 bits (141), Expect = 2e-09
Identities = 49/154 (31%), Positives = 71/154 (45%), Gaps = 6/154 (3%)
Frame = +1
Query: 68 LPAGVKFDPNDQEILQH-LEAKVLFDVPKLHPLIDEFIPTLEGENGICYTHPEKLPG--- 123
+P G +F P D+E++ + L+ K+L D +H IP ++ +C P +PG
Sbjct: 181 MPVGFRFRPTDEELVDYYLKHKLLADDFPVH-----IIPEID----LCKVEPWDVPGRSV 333
Query: 124 VKKDGQIRHFFHRPSKAYTTGTRKRRKVQTDEEGSETRWHKTGKTRPVIIDGVMK--GFK 181
+K D FF Y R R T + G W TG R V I G G K
Sbjct: 334 IKSDDPEWFFFSPVDYKYLKSKRFNR---TTKRGY---WKTTGNDRNVKIPGTSNVIGTK 495
Query: 182 KILVLYTNYGRQKKPEKTNWVMHQYHLGSNEEEK 215
K LV + GR + KTNWV+H+YH ++ E +
Sbjct: 496 KTLVFHE--GRGPRGVKTNWVIHEYHAVTSHESQ 591
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.316 0.136 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,683,519
Number of Sequences: 63676
Number of extensions: 154704
Number of successful extensions: 917
Number of sequences better than 10.0: 82
Number of HSP's better than 10.0 without gapping: 896
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 900
length of query: 259
length of database: 12,639,632
effective HSP length: 95
effective length of query: 164
effective length of database: 6,590,412
effective search space: 1080827568
effective search space used: 1080827568
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 58 (26.9 bits)
Medicago: description of AC142143.13


