

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
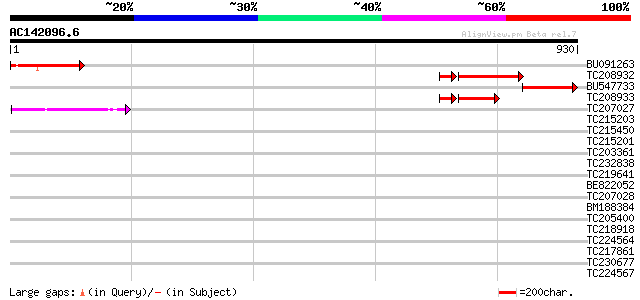
Query= AC142096.6 - phase: 0 /pseudo
(930 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BU091263 166 4e-41
TC208932 weakly similar to UP|Q9LQB3 (Q9LQB3) F4N2.4, partial (10%) 118 1e-28
BU547733 100 2e-21
TC208933 similar to UP|Q9LTR4 (Q9LTR4) Similarity to EH domain c... 75 5e-16
TC207027 weakly similar to UP|Q9LW95 (Q9LW95) KED, partial (17%) 44 4e-04
TC215203 similar to UP|Q70Z19 (Q70Z19) Nucleosome assembly prote... 42 0.001
TC215450 similar to UP|P93488 (P93488) NAP1Ps, partial (30%) 38 0.018
TC215201 similar to UP|Q70Z19 (Q70Z19) Nucleosome assembly prote... 37 0.031
TC203361 similar to UP|Q6VBJ4 (Q6VBJ4) Epa5p, partial (7%) 37 0.040
TC232838 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {A... 37 0.053
TC219641 similar to UP|Q6VBJ3 (Q6VBJ3) Epa4p, partial (6%) 36 0.090
BE822052 similar to GP|4557063|gb|A expressed protein {Arabidops... 35 0.12
TC207028 similar to UP|Q9LW95 (Q9LW95) KED, partial (13%) 35 0.12
BM188384 homologue to GP|22093701|dbj OJ1316_A04.14 {Oryza sativ... 35 0.20
TC205400 weakly similar to GB|AAP37850.1|30725656|BT008491 At4g2... 35 0.20
TC218918 UP|Q39892 (Q39892) Nucleosome assembly protein 1, complete 34 0.26
TC224564 similar to PIR|T51159|T51159 HMG protein [imported] - A... 33 0.44
TC217861 similar to UP|Q761Y3 (Q761Y3) BRI1-KD interacting prote... 33 0.58
TC230677 similar to UP|Q7Q3H3 (Q7Q3H3) EbiP7281 (Fragment), part... 32 0.99
TC224567 similar to PIR|T51159|T51159 HMG protein [imported] - A... 32 0.99
>BU091263
Length = 424
Score = 166 bits (420), Expect = 4e-41
Identities = 85/126 (67%), Positives = 99/126 (78%), Gaps = 4/126 (3%)
Frame = +1
Query: 1 MAKSSKRSIANGTNTSNKSTSKKKKNNKMGPEAVAMKAKAQKTN----TNPFESIWSRRK 56
MAKSSKRS+ ++T +K K K++ K GPE VAMK KA N +NPFESIWSRRK
Sbjct: 49 MAKSSKRSVGT-SSTDSKKKKKNKRSKKFGPEGVAMKVKANNNNNGTASNPFESIWSRRK 225
Query: 57 FEVMGQKRKGDTKRMGLARSLAVEKRKKTLLKEYEQSTKSSEFIDRRIGENDEGLDDFGK 116
FEV+GQKRKG+ +RMGLARSLA++KR TLLKEY QS KSS F+D+RIGE DE LDDFGK
Sbjct: 226 FEVLGQKRKGEARRMGLARSLAIQKRNDTLLKEYHQSAKSSLFVDKRIGEKDEALDDFGK 405
Query: 117 AILRSQ 122
AILRSQ
Sbjct: 406 AILRSQ 423
>TC208932 weakly similar to UP|Q9LQB3 (Q9LQB3) F4N2.4, partial (10%)
Length = 705
Score = 118 bits (296), Expect(2) = 1e-28
Identities = 74/107 (69%), Positives = 80/107 (74%)
Frame = +3
Query: 736 NRHA*RLSILHYGQFQGKCISYRCRNPTRIYQCL**LEFVSRNIFTNFEITT*DSRTEKY 795
NRHA RL LH FQG CIS N TRI L* L+F+S NI TNFEI *+SRTEKY
Sbjct: 159 NRHARRLFFLHLC*FQG*CISCSH*NSTRIR*RL*RLKFIS*NIPTNFEIIK*NSRTEKY 338
Query: 796 AKCITG*N*RCG*AD*VESG*TSYFEATTSNAETKASSNQITES*V* 842
KCI *N*RCG*A *VESG S+F+ATTSNAETKAS+NQITES V*
Sbjct: 339 VKCIMR*N*RCG*AH*VESGQASHFKATTSNAETKASTNQITESQV* 479
Score = 27.7 bits (60), Expect(2) = 1e-28
Identities = 17/28 (60%), Positives = 18/28 (63%)
Frame = +2
Query: 706 ISSYGTERSQATIAH**GSRQD*RFEFL 733
ISSYGTE SQA+ H S D* EFL
Sbjct: 68 ISSYGTESSQASPIHTPYSE*D*SSEFL 151
>BU547733
Length = 669
Score = 100 bits (250), Expect = 2e-21
Identities = 62/90 (68%), Positives = 69/90 (75%)
Frame = -1
Query: 841 V*GKLR*G*RL*SRPCAS*KKKIEERGKA*S*RGCS*NT*RQLLLA*CEGKRKVFNGESK 900
V*G+L G RL*S P S* +KIEE +A*S*RGC+ *RQLLLA EGKR+VF G+
Sbjct: 669 V*GELCQGQRL*S*P*TS*TEKIEETFEA*S*RGCTRIA*RQLLLAGGEGKREVFTGKR* 490
Query: 901 S*EVWKD*SFPSRARTCFQIWTTRKREEEK 930
S EVWK *SFPSRARTC QIWTTRKR EE+
Sbjct: 489 SREVWKS*SFPSRARTCLQIWTTRKR*EEE 400
>TC208933 similar to UP|Q9LTR4 (Q9LTR4) Similarity to EH domain containing
proteins, partial (12%)
Length = 648
Score = 75.1 bits (183), Expect(2) = 5e-16
Identities = 44/68 (64%), Positives = 50/68 (72%)
Frame = +3
Query: 736 NRHA*RLSILHYGQFQGKCISYRCRNPTRIYQCL**LEFVSRNIFTNFEITT*DSRTEKY 795
NRHA RL LH+ FQG CIS N TRI+Q L* L+F+S NI TNFEI *+SRT+KY
Sbjct: 444 NRHARRLFFLHFC*FQG*CISCSR*NSTRIHQRL*RLKFIS*NIPTNFEIIK*NSRTKKY 623
Query: 796 AKCITG*N 803
AKCIT *N
Sbjct: 624 AKCITR*N 647
Score = 28.5 bits (62), Expect(2) = 5e-16
Identities = 17/28 (60%), Positives = 18/28 (63%)
Frame = +2
Query: 706 ISSYGTERSQATIAH**GSRQD*RFEFL 733
ISSYGTE SQA+ H S D* EFL
Sbjct: 353 ISSYGTESSQASPIHTPDSE*D*SSEFL 436
>TC207027 weakly similar to UP|Q9LW95 (Q9LW95) KED, partial (17%)
Length = 1005
Score = 43.5 bits (101), Expect = 4e-04
Identities = 50/198 (25%), Positives = 81/198 (40%), Gaps = 2/198 (1%)
Frame = +3
Query: 3 KSSKRSIANGTNTSNKSTSKKKKNNKMGPEAVAMKAKAQKTNTNPFESIWSRRKFEVMGQ 62
K K T + KK+ + K E K K K + + K E G
Sbjct: 144 KKKKEKKDKDGKTDAEKIKKKEDDGKDDEEKKEKKKKYDKIDAXKVKGKEDDGKDE--GN 317
Query: 63 KRKGDTKRMGLARSLAVEKRKKTLLKEYEQSTKSSEFID-RRIGENDEGLDDFG-KAILR 120
K K D ++ EK+ K K+ E+ K E + G+NDEG DD G K +
Sbjct: 318 KEKKDKEKGDGDGEEKKEKKDKEKEKKKEKKDKDEETDTLKEKGKNDEGEDDEGNKKKKK 497
Query: 121 SQRERQLNVKSSKKSKYHLSEEDDDEFEGIDGLGRDDFEDEMLGEDDDETDET*NQEGSD 180
++E++ + K KK K +ED + + ++ + E GE +D+ ++G
Sbjct: 498 DKKEKEKDHKKEKKDKEEGEKEDSKVEVSVRDIDIEEIKKE--GEKEDK-----GKDGGK 656
Query: 181 ERSYCKKQVLQG*KSKRK 198
E KK+ + K K+K
Sbjct: 657 EVKEKKKKEDKDKKEKKK 710
>TC215203 similar to UP|Q70Z19 (Q70Z19) Nucleosome assembly protein 1-like
protein 1, partial (31%)
Length = 729
Score = 42.4 bits (98), Expect = 0.001
Identities = 37/127 (29%), Positives = 59/127 (46%), Gaps = 7/127 (5%)
Frame = +2
Query: 81 KRKKTLLKEYEQSTKSSEFIDRRIGENDEGLDDFGKAILRSQRERQLNVKSSKKSK---- 136
K K + K + + + F ++ E+D+ +DD L + E ++ S+ + K
Sbjct: 5 KNAKPITKTEKCESFFNFFNPPQVPEDDDDIDDDAVEELENLMEHDYDIGSTIRDKIIPH 184
Query: 137 ---YHLSEEDDDEFEGIDGLGRDDFEDEMLGEDDDETDET*NQEGSDERSYCKKQVLQG* 193
+ E +FE ID D+EDE G+DDD+ DE +++G DE KK
Sbjct: 185 AVSWFTGEAAQSDFEDID---EADYEDEE-GDDDDDEDEDEDEDGEDEEE--KKG----- 331
Query: 194 KSKRKGK 200
KSK K K
Sbjct: 332 KSKSKSK 352
>TC215450 similar to UP|P93488 (P93488) NAP1Ps, partial (30%)
Length = 672
Score = 38.1 bits (87), Expect = 0.018
Identities = 25/84 (29%), Positives = 45/84 (52%), Gaps = 4/84 (4%)
Frame = +3
Query: 102 RRIGENDEGLDDFGKAILRSQRERQLNVKSSKKSK---YHLSEEDDDEFEGID-GLGRDD 157
R+ E+DE +D+ L++Q E+ ++ S+ + K + +S + +G + G DD
Sbjct: 39 RKFPEDDEDIDEDAAXELQNQMEQDYDIGSTIRDKIIPHAVSWFTGEAIQGDEFGDLEDD 218
Query: 158 FEDEMLGEDDDETDET*NQEGSDE 181
+DE + EDDDE +E + DE
Sbjct: 219 EDDEDIEEDDDEEEEEDEDDDDDE 290
>TC215201 similar to UP|Q70Z19 (Q70Z19) Nucleosome assembly protein 1-like
protein 1, partial (55%)
Length = 904
Score = 37.4 bits (85), Expect = 0.031
Identities = 29/123 (23%), Positives = 56/123 (44%), Gaps = 7/123 (5%)
Frame = +3
Query: 81 KRKKTLLKEYEQSTKSSEFIDRRIGENDEGLDDFGKAILRSQRERQLNVKSSKKSK---- 136
K K + K + + + F + E+D+ +D+ L++ ER ++ S+ + K
Sbjct: 276 KNAKPITKTEKCESFFNFFNPPEVPEDDDDIDEDAVEELQNLMERDYDIGSTIRDKIIPH 455
Query: 137 ---YHLSEEDDDEFEGIDGLGRDDFEDEMLGEDDDETDET*NQEGSDERSYCKKQVLQG* 193
+ E +FE ID + D++DE ED+DE ++ +E +E K + G
Sbjct: 456 AVSWFTGEAAQSDFEDIDEV---DYDDEEGDEDEDEDEDEEEEEEEEEEKKGKSRSKSGG 626
Query: 194 KSK 196
+ K
Sbjct: 627 RVK 635
>TC203361 similar to UP|Q6VBJ4 (Q6VBJ4) Epa5p, partial (7%)
Length = 1090
Score = 37.0 bits (84), Expect = 0.040
Identities = 31/124 (25%), Positives = 55/124 (44%), Gaps = 10/124 (8%)
Frame = -1
Query: 67 DTKRMGLARSLAVEKRKKTLLKEYEQSTKSSEFIDRRIGENDEGLDDFGKAILRSQRE-- 124
D R+ L + + +KT ++ ++ + + D ++D+G FG+ E
Sbjct: 682 DLYRLLLTNTPTTKGERKTQSQDKQRDAEGEDGNDDEGDDDDDGDGGFGEGEEELSSEDG 503
Query: 125 --------RQLNVKSSKKSKYHLSEEDDDEFEGIDGLGRDDFEDEMLGEDDDETDET*NQ 176
+ N K + + +EE+ +E E DG +DD ED+ +DDDE DE +
Sbjct: 502 GGYGNNSNNKSNSKKAPEGGAGGAEENGEEDEDEDGDDQDDDEDDD-DDDDDEEDEGGEE 326
Query: 177 EGSD 180
E D
Sbjct: 325 EDED 314
>TC232838 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {Arabidopsis
thaliana;} , partial (46%)
Length = 902
Score = 36.6 bits (83), Expect = 0.053
Identities = 19/41 (46%), Positives = 27/41 (65%), Gaps = 2/41 (4%)
Frame = +2
Query: 139 LSEEDDDEFEGIDGLGRD--DFEDEMLGEDDDETDET*NQE 177
L EEDD+E+E +D + D D EDE +DDDE D+ ++E
Sbjct: 275 LVEEDDEEYESVDEVVDDAEDDEDEEEDDDDDEGDDNDDEE 397
Score = 35.8 bits (81), Expect = 0.090
Identities = 29/119 (24%), Positives = 53/119 (44%), Gaps = 2/119 (1%)
Frame = +2
Query: 66 GDTKRMGLARSLAVEKRKKTLLKEYEQSTKSSEFIDRRIGENDEGLDDFGKAILRSQRER 125
G +R A+ ++ + ++ +L + K+++ + ++ EN+ K ++ E
Sbjct: 137 GGKRREEAAQRISKSRGERNVLWFLDSKMKATKELSEQVEENEV------KVLVEEDDEE 298
Query: 126 QLNVKS--SKKSKYHLSEEDDDEFEGIDGLGRDDFEDEMLGEDDDETDET*NQEGSDER 182
+V EEDDD+ EG D DD ED+ DD+ DE +EG +R
Sbjct: 299 YESVDEVVDDAEDDEDEEEDDDDDEGDD---NDDEEDDAPDGGDDDDDEDDEEEGDVQR 466
Score = 29.3 bits (64), Expect = 8.4
Identities = 24/104 (23%), Positives = 47/104 (45%), Gaps = 1/104 (0%)
Frame = +2
Query: 80 EKRKKTLLKEYEQSTKS-SEFIDRRIGENDEGLDDFGKAILRSQRERQLNVKSSKKSKYH 138
E K L++E ++ +S E +D + DE DD + + + +
Sbjct: 257 ENEVKVLVEEDDEEYESVDEVVDDAEDDEDEEEDDDD-----DEGDDNDDEEDDAPDGGD 421
Query: 139 LSEEDDDEFEGIDGLGRDDFEDEMLGEDDDETDET*NQEGSDER 182
+++DDE EG G + +D+ +DD + DE ++E +E+
Sbjct: 422 DDDDEDDEEEGDVQRGGEPDDDDNDSDDDSDDDEDEDEEDEEEQ 553
>TC219641 similar to UP|Q6VBJ3 (Q6VBJ3) Epa4p, partial (6%)
Length = 942
Score = 35.8 bits (81), Expect = 0.090
Identities = 23/68 (33%), Positives = 34/68 (49%), Gaps = 1/68 (1%)
Frame = +1
Query: 106 ENDEGLDDF-GKAILRSQRERQLNVKSSKKSKYHLSEEDDDEFEGIDGLGRDDFEDEMLG 164
+++EG +DF G+ + E + S ++DD + D G DD EDE G
Sbjct: 436 DDEEGDEDFSGEGEEEADPEDDPEANGAGGSDSDGDDDDDGGDDDDDDNGDDDGEDEEEG 615
Query: 165 EDDDETDE 172
ED+DE DE
Sbjct: 616 EDEDEEDE 639
Score = 34.7 bits (78), Expect = 0.20
Identities = 28/97 (28%), Positives = 42/97 (42%), Gaps = 4/97 (4%)
Frame = +1
Query: 106 ENDEGLDDFGKAILRSQRERQLNVKSSKKSKYHLSEEDDDEFEGIDGLGRDDFEDEMLGE 165
E+D+ DD Q + + + S + + EDD E G G D +D+ G+
Sbjct: 409 EDDDDADD--------QDDEEGDEDFSGEGEEEADPEDDPEANGAGGSDSDGDDDDDGGD 564
Query: 166 DDD----ETDET*NQEGSDERSYCKKQVLQG*KSKRK 198
DDD + D +EG DE ++ V Q KRK
Sbjct: 565 DDDDDNGDDDGEDEEEGEDEDEEDEEDVPQPPAKKRK 675
>BE822052 similar to GP|4557063|gb|A expressed protein {Arabidopsis
thaliana}, partial (28%)
Length = 668
Score = 35.4 bits (80), Expect = 0.12
Identities = 27/102 (26%), Positives = 46/102 (44%), Gaps = 3/102 (2%)
Frame = -1
Query: 100 IDRRIGENDEGLDDFGKAILRSQRERQLNVKSSKKSKYHLSEEDDDEFEGIDGLGRDDFE 159
+++ ++DE D+ Q + + + S + + EDD E G G DD
Sbjct: 539 LNKDASDSDEDEDEEDDHDTNDQNDEEGDEDFSGEGEEEADPEDDPEANGAGGSNDDDDS 360
Query: 160 DEMLGED---DDETDET*NQEGSDERSYCKKQVLQG*KSKRK 198
DE +D DD+ ++ +EG DE +++V Q KRK
Sbjct: 359 DEDDDDDNDGDDDGEDEDEKEGEDEED--EEEVPQPPTKKRK 240
Score = 32.0 bits (71), Expect = 1.3
Identities = 23/84 (27%), Positives = 36/84 (42%)
Frame = -1
Query: 89 EYEQSTKSSEFIDRRIGENDEGLDDFGKAILRSQRERQLNVKSSKKSKYHLSEEDDDEFE 148
E E + D+ E DE G+ + + + N E+DDD+ +
Sbjct: 512 EDEDEEDDHDTNDQNDEEGDEDFSGEGEEEADPEDDPEANGAGGSNDDDDSDEDDDDDND 333
Query: 149 GIDGLGRDDFEDEMLGEDDDETDE 172
G D G D EDE GED+++ +E
Sbjct: 332 GDDD-GED--EDEKEGEDEEDEEE 270
>TC207028 similar to UP|Q9LW95 (Q9LW95) KED, partial (13%)
Length = 490
Score = 35.4 bits (80), Expect = 0.12
Identities = 26/93 (27%), Positives = 48/93 (50%), Gaps = 1/93 (1%)
Frame = +1
Query: 80 EKRKKTLLKEYEQSTKSSEFIDRRIGENDEGLDDFGKAILRSQRERQLNVKSSKKSKYHL 139
EK++K KE + + ++ + + G+NDEG DD G + +++++ K KK K +
Sbjct: 163 EKKEKEKKKEKKDKDEETDTLKEK-GKNDEGEDDEGNK--KKKKDKKEKEKDHKKEKKDI 333
Query: 140 SEEDDDEFEGIDGLGRD-DFEDEMLGEDDDETD 171
EE + E ++ RD D E++ + D D
Sbjct: 334 -EEGEKEDSKVEASVRDIDIEEKKVTGKDKTKD 429
>BM188384 homologue to GP|22093701|dbj OJ1316_A04.14 {Oryza sativa (japonica
cultivar-group)}, partial (2%)
Length = 437
Score = 34.7 bits (78), Expect = 0.20
Identities = 25/90 (27%), Positives = 45/90 (49%)
Frame = +1
Query: 73 LARSLAVEKRKKTLLKEYEQSTKSSEFIDRRIGENDEGLDDFGKAILRSQRERQLNVKSS 132
+ R V+K K+ + K + S+K + I+ D G+D K I+ S+ + + +
Sbjct: 106 VCRKSKVDKNKEVVEKMGDGSSKKT--INEDDERKDSGVDSGKKVIMESKVDDGKDSLVA 279
Query: 133 KKSKYHLSEEDDDEFEGIDGLGRDDFEDEM 162
K + EE+D E++ I+ L R D E E+
Sbjct: 280 NKHSGNKKEEEDLEWDEIENLSRIDEEKEV 369
>TC205400 weakly similar to GB|AAP37850.1|30725656|BT008491 At4g26630
{Arabidopsis thaliana;} , partial (20%)
Length = 794
Score = 34.7 bits (78), Expect = 0.20
Identities = 40/155 (25%), Positives = 70/155 (44%), Gaps = 5/155 (3%)
Frame = +1
Query: 23 KKKNNKMGPEAVAMKAKAQKTNTNPFESIWSRRKFEVMG-QKRKGDTKRMGLARSLAVEK 81
KKKN K G + + ++ +P ++SR+K E G QKR TK ++ V K
Sbjct: 1 KKKNEKGGKQKID-----DDSDESPNPKVFSRKKNEKGGKQKRSTPTKSTSKEKTEKVTK 165
Query: 82 RKKTLLKEYEQSTKSS----EFIDRRIGENDEGLDDFGKAILRSQRERQLNVKSSKKSKY 137
K K+ E+S+ S + I + + E D F + + ++ +++ K S
Sbjct: 166 GKG---KKKEKSSPSDNQLRDAICKILKEVDFNTATFTDILKQLAKQFDMDLTPKKASIK 336
Query: 138 HLSEEDDDEFEGIDGLGRDDFEDEMLGEDDDETDE 172
+ +E+ + D+ +DE GE+D E DE
Sbjct: 337 LMIQEELTKLA-------DEADDEEDGEEDAEKDE 420
>TC218918 UP|Q39892 (Q39892) Nucleosome assembly protein 1, complete
Length = 1507
Score = 34.3 bits (77), Expect = 0.26
Identities = 32/128 (25%), Positives = 53/128 (41%), Gaps = 8/128 (6%)
Frame = +1
Query: 81 KRKKTLLKEYEQSTKSSEFIDRRIGENDEGLDDFGKAILRSQRERQLNVKSSKKSK---- 136
K K + K + + F + E+D +D+ L++Q E+ ++ S+ + K
Sbjct: 769 KNAKPITKTESCESFFNFFKPPEVPEDDADIDEDLAEELQNQMEQDYDIGSTLRDKIIPH 948
Query: 137 ----YHLSEEDDDEFEGIDGLGRDDFEDEMLGEDDDETDET*NQEGSDERSYCKKQVLQG 192
+ DEFE ++ D EDE ED+DE +E E +E K
Sbjct: 949 AVSWFTGEAAQGDEFEDLE-----DDEDEEEDEDEDEDEEDDEDEDDEEEDDTKT----- 1098
Query: 193 *KSKRKGK 200
K K+ GK
Sbjct: 1099-KKKKSGK 1119
>TC224564 similar to PIR|T51159|T51159 HMG protein [imported] - Arabidopsis
thaliana {Arabidopsis thaliana;} , partial (82%)
Length = 833
Score = 33.5 bits (75), Expect = 0.44
Identities = 43/185 (23%), Positives = 73/185 (39%), Gaps = 3/185 (1%)
Frame = +1
Query: 6 KRSIANGTNTSNKSTSKKKKNNKMGPEAVAMKAKAQKTNTNPFESIWSRRKFEVMGQKRK 65
K + G ++K + K + K+G +K + P S S K E +K
Sbjct: 88 KNAKGKGAARASKESLKPVDDRKVGK---------RKASGKPGRS--SAPKKEKKAKKDP 234
Query: 66 GDTKRMGLARSLAVEKRKKTLLKEYEQSTKSSEFIDRRIGENDEGLDDFGKAILRSQ--- 122
KR A + +E+ +KT E + K+ + + GE + L KA ++
Sbjct: 235 NKPKRPPSAFFVFLEEFRKTFKAE-NPNVKAVSVVGKAGGEKWKSLSSAEKAPYEAKAAK 411
Query: 123 RERQLNVKSSKKSKYHLSEEDDDEFEGIDGLGRDDFEDEMLGEDDDETDET*NQEGSDER 182
R+ + K S DD+E + D ED+ GE+++E DE + D+
Sbjct: 412 RKAEYEKLIKAYDKKQASSADDEESDKSKSEVND--EDDASGEEEEEDDEEEEDDEDDD* 585
Query: 183 SYCKK 187
C K
Sbjct: 586 D*CPK 600
>TC217861 similar to UP|Q761Y3 (Q761Y3) BRI1-KD interacting protein 132
(Fragment), partial (27%)
Length = 1293
Score = 33.1 bits (74), Expect = 0.58
Identities = 34/130 (26%), Positives = 64/130 (49%), Gaps = 1/130 (0%)
Frame = +1
Query: 18 KSTSKKKKNNKMGPEAVAMKAKAQKTNTNPFESIWSRRKFEVMGQKRKGDTKRMGLARSL 77
KS KKKK +K+ E++A + N ES+ + + +V + G + +
Sbjct: 394 KSKEKKKKKSKLVSESLATNVE-----DNHLESVATVEENKV----KDGVSTDAKVINGS 546
Query: 78 AVEKRKKTLLKEYEQSTKSSEFIDRRIGENDEGLDDFGKAILRSQRERQLNV-KSSKKSK 136
EK+ K+ +K+ ++ + + I++ IG+ +E + + I S +E V K SKK K
Sbjct: 547 EKEKKSKSKMKKKDKQSGQGDVIEQ-IGDPNETVSK-EENIEASNKEMMHEVEKDSKKRK 720
Query: 137 YHLSEEDDDE 146
+SEE+ +
Sbjct: 721 RPISEENGQQ 750
>TC230677 similar to UP|Q7Q3H3 (Q7Q3H3) EbiP7281 (Fragment), partial (26%)
Length = 1076
Score = 32.3 bits (72), Expect = 0.99
Identities = 15/34 (44%), Positives = 21/34 (61%)
Frame = +3
Query: 136 KYHLSEEDDDEFEGIDGLGRDDFEDEMLGEDDDE 169
K L EE +D+ EG+D DD ED+ E+D+E
Sbjct: 3 KSMLGEESEDDEEGLDSESDDDDEDDDSDEEDEE 104
>TC224567 similar to PIR|T51159|T51159 HMG protein [imported] - Arabidopsis
thaliana {Arabidopsis thaliana;} , partial (71%)
Length = 876
Score = 32.3 bits (72), Expect = 0.99
Identities = 44/176 (25%), Positives = 69/176 (39%), Gaps = 7/176 (3%)
Frame = +2
Query: 13 TNTSNKSTSKKKKNNKMGPEAVA----MKAKAQKTNTNPFESIWSRRKFEVMGQKRKGDT 68
T T T+K K + E++ K +K + P +S K E +K
Sbjct: 113 TGT*TMKTAKGKGAARPSKESLKPVDDRKVGKRKASGKPEKS--RAPKKEKKAKKDPNKP 286
Query: 69 KRMGLARSLAVEKRKKTLLKEYEQSTKSSEFIDRRIGENDEGLDDFGKAILRSQ---RER 125
KR A + +E+ +KT E K+ + + GE + L KA ++ R+
Sbjct: 287 KRPPSAFFVFLEEFRKTFKAE-NPLVKAVSVVGKAGGEKWKSLSSAEKAPYEAKAAKRKA 463
Query: 126 QLNVKSSKKSKYHLSEEDDDEFEGIDGLGRDDFEDEMLGEDDDETDET*NQEGSDE 181
+ K S DDDE + D ED+ GE+D + DE +E DE
Sbjct: 464 EYEKLIKAYEKKQASSADDDESDKSKSEVND--EDDASGEEDHQEDEDDEEEEDDE 625
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.377 0.171 0.654
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 35,998,479
Number of Sequences: 63676
Number of extensions: 476982
Number of successful extensions: 9649
Number of sequences better than 10.0: 61
Number of HSP's better than 10.0 without gapping: 2991
Number of HSP's successfully gapped in prelim test: 402
Number of HSP's that attempted gapping in prelim test: 6154
Number of HSP's gapped (non-prelim): 3868
length of query: 930
length of database: 12,639,632
effective HSP length: 106
effective length of query: 824
effective length of database: 5,889,976
effective search space: 4853340224
effective search space used: 4853340224
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 13 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 35 (21.6 bits)
S2: 63 (28.9 bits)
Medicago: description of AC142096.6


