

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141865.9 - phase: 0 /pseudo
(547 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
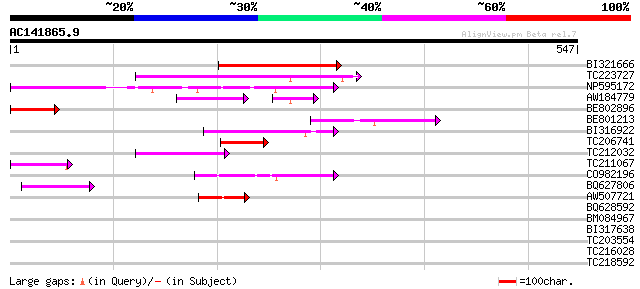
Score E
Sequences producing significant alignments: (bits) Value
BI321666 155 4e-38
TC223727 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 113 2e-25
NP595172 polyprotein [Glycine max] 110 2e-24
AW184779 56 3e-13
BE802896 70 2e-12
BE801213 weakly similar to GP|6691193|gb| F7F22.17 {Arabidopsis ... 63 4e-10
BI316922 62 7e-10
TC206741 58 1e-08
TC212032 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 54 2e-07
TC211067 similar to UP|Q9LQH2 (Q9LQH2) F15O4.13, partial (9%) 47 2e-05
CO982196 45 6e-05
BQ627806 45 1e-04
AW507721 weakly similar to GP|27764548|gb polyprotein {Glycine m... 44 2e-04
BQ628592 34 0.15
BM084967 33 0.43
BI317638 weakly similar to GP|9294238|dbj| contains similarity t... 30 2.1
TC203554 homologue to UP|RL38_ARATH (O22860) 60S ribosomal prote... 30 3.6
TC216028 homologue to UP|Q9ZSQ0 (Q9ZSQ0) Ethylene response senso... 29 4.7
TC218592 weakly similar to UP|Q9LP32 (Q9LP32) T9L6.1 protein, pa... 28 8.1
>BI321666
Length = 430
Score = 155 bits (392), Expect = 4e-38
Identities = 72/119 (60%), Positives = 91/119 (75%)
Frame = +2
Query: 202 SFKILHSGFWWPSLFKDVHLFISKCDKCQRTGNITKRNEMPLNNILEVEIFDVWVIDFIG 261
S +L S F+ PS+FKD ++ C+KCQRT +++KRNE+PL+ ILEVEIFD W IDF+G
Sbjct: 5 STNVLQSRFYLPSIFKDAYVHAQSCNKCQRTRSVSKRNELPLHTILEVEIFDYWGIDFVG 184
Query: 262 PFPSSFVNQYILVAVDYVSKWVEVIASLTNDAQVVIKLFKKNHFSRFGVPRVVISDGGS 320
PFP SF N+YILV VDYVSKWVE +A +DA++VIK KK FSR GVP V+I +GGS
Sbjct: 185 PFPPSFSNEYILVVVDYVSKWVEAVACQKSDAKIVIKFLKKQIFSRLGVPWVLIDNGGS 361
>TC223727 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (9%)
Length = 843
Score = 113 bits (283), Expect = 2e-25
Identities = 65/223 (29%), Positives = 107/223 (47%), Gaps = 5/223 (2%)
Frame = +1
Query: 122 PWYADFVNFLAAGVLPPELSYQQKKKFFNDLKHYYSDEPYLFKRG*DGIFRRCIPENEVS 181
PWY D ++ + PE++ K+ ++ L+KR D RC+ E +
Sbjct: 136 PWYFDIKRYVISKEYLPEIADNDKRTLRRLAAGFFMSGSILYKRNHDMKPLRCVDAREAN 315
Query: 182 SILTHCHSSSYGGHASTQKASFKILHSGFWWPSLFKDVHLFISKCDKCQRTGNITKRNEM 241
++ H S+G HA+ + KIL +G++W ++ D + + KC KCQ +
Sbjct: 316 HMIEEVHEGSFGTHANGHAMARKILRAGYYWLTMESDCCVHVRKCHKCQAFADNVNAPPH 495
Query: 242 PLNNILEVEIFDVWVIDFIGPFPSSFVN--QYILVAVDYVSKWVEVIASLTNDAQVVIKL 299
PLN + F +W ID IG N ++ILVA+DY +KWVE + VV++
Sbjct: 496 PLNVMSSPWPFSMWGIDVIGAIEPKASNGHRFILVAIDYFTKWVEAASYTDVMRGVVVRF 675
Query: 300 FKKNHFSRFGVPRVVISDGG---SQNILKNFCKNLE*GTKLQH 339
KK R+G+PR +I+D G + ++ C+ K+QH
Sbjct: 676 IKKEIICRYGLPRKIITDNGTNLNNKMMGEICEEF----KIQH 792
>NP595172 polyprotein [Glycine max]
Length = 4659
Score = 110 bits (275), Expect = 2e-24
Identities = 94/326 (28%), Positives = 151/326 (45%), Gaps = 9/326 (2%)
Frame = +1
Query: 1 IYYASKTLDGALVNYATTEKELLAVVYAIDKFRQYLVGSKITVYTDHSTIKYLLNKKDAK 60
I Y SK L + + +ELLA+ A+ KFR YL+G+K + TD ++K L+++
Sbjct: 2740 IAYFSKKLAPRMQKQSAYTRELLAITEALSKFRHYLLGNKFIIRTDQRSLKSLMDQSLQT 2919
Query: 61 PRLIQLIFLLQEFDLEIKDKKGVENVVADHLSRLRETNEDELPLDDSFPDDQLFLLAQTN 120
P + +D +I+ K G +N AD LSR +F+LA +
Sbjct: 2920 PEQQAWLHKFLGYDFKIEYKPGKDNQAADALSR-------------------MFMLAWSE 3042
Query: 121 APWYADFVNFLAAGVL--PPELSYQQKKKFFNDLKHYYSDEPYLFKRG*DGIFRRCIPEN 178
++ F+ L A ++ P + K D HY E L+ + R IP
Sbjct: 3043 P--HSIFLEELRARLISDPHLKQLMETYKQGADASHYTVREGLLYWKD-----RVVIPAE 3201
Query: 179 E--VSSILTHCHSSSYGGHASTQKASFKILHSGFWWPSLFKDVHLFISKCDKCQRTGNIT 236
E V+ IL HSS GGHA + + L + F+WP + +DV +I KC CQ+
Sbjct: 3202 EEIVNKILQEYHSSPIGGHAGITRTLAR-LKAQFYWPKMQEDVKAYIQKCLICQQA---K 3369
Query: 237 KRNEMPLNNILEVEI-FDVW---VIDFIGPFPSSFVNQYILVAVDYVSKWVEVIASLTN- 291
N +P + + I VW +DFI P+SF I+V +D ++K+ I +
Sbjct: 3370 SNNTLPAGLLQPLPIPQQVWEDVAMDFITGLPNSFGLSVIMVVIDRLTKYAHFIPLKADY 3549
Query: 292 DAQVVIKLFKKNHFSRFGVPRVVISD 317
+++VV + F + G+PR ++SD
Sbjct: 3550 NSKVVAEAFMSHIVKLHGIPRSIVSD 3627
>AW184779
Length = 432
Score = 55.8 bits (133), Expect(2) = 3e-13
Identities = 24/69 (34%), Positives = 37/69 (52%)
Frame = +1
Query: 162 LFKRG*DGIFRRCIPENEVSSILTHCHSSSYGGHASTQKASFKILHSGFWWPSLFKDVHL 221
L+KR D + RC+ E +L H S+G HA+ + KIL G++W ++ D +
Sbjct: 16 LYKRNHDMVLLRCVDAREAEQMLVEVHEGSFGTHANIHAMAQKILRVGYYWLTMESDCCI 195
Query: 222 FISKCDKCQ 230
+ KC KCQ
Sbjct: 196 HVWKCHKCQ 222
Score = 37.4 bits (85), Expect(2) = 3e-13
Identities = 19/47 (40%), Positives = 27/47 (57%), Gaps = 2/47 (4%)
Frame = +3
Query: 254 VWVIDFIGPFPSSFVN--QYILVAVDYVSKWVEVIASLTNDAQVVIK 298
+W ID IG N +ILVA+DY +KWVE ++ + VVI+
Sbjct: 291 MWGIDVIGAIEPKASNGHHFILVAIDYFTKWVEAVSYASVTRSVVIR 431
>BE802896
Length = 416
Score = 70.1 bits (170), Expect = 2e-12
Identities = 30/48 (62%), Positives = 40/48 (82%)
Frame = -2
Query: 1 IYYASKTLDGALVNYATTEKELLAVVYAIDKFRQYLVGSKITVYTDHS 48
IYY+S+TLD A NY TTEKELLA+V+A++KF YL+G++I VY DH+
Sbjct: 145 IYYSSRTLDAAQANYTTTEKELLAIVFALEKFHSYLLGTRIIVYIDHA 2
>BE801213 weakly similar to GP|6691193|gb| F7F22.17 {Arabidopsis thaliana},
partial (3%)
Length = 416
Score = 62.8 bits (151), Expect = 4e-10
Identities = 50/134 (37%), Positives = 65/134 (48%), Gaps = 9/134 (6%)
Frame = +3
Query: 291 NDAQVVIKLFKKNHFSRFGVPRVVISDGGSQNILKNFCKNLE*GTKLQHPITHKLVDKWR 350
NDA++VIK KKN FSRFG+PR++ISDGGS K L+ + HK+ +
Sbjct: 3 NDAKIVIKFLKKNIFSRFGMPRILISDGGSHFYYSQLNKVLK-----HDSVRHKVETSYH 167
Query: 351 ---------SQIDKSKLSLKKLFQPLGKIGQVSLTMPFGPIERLIKLPLE*LLLN*FMGN 401
S I K KK++ KIG + M FG L KL L *L +G
Sbjct: 168 PQTNGQAKVSNIHKENS*EKKMWHLQEKIGHPN*RMLFGHARLLKKLQLV*LHSKWSIGR 347
Query: 402 LVAFRLSSSIRLIG 415
LV + S S + IG
Sbjct: 348 LVIYP*S*STKHIG 389
>BI316922
Length = 405
Score = 62.0 bits (149), Expect = 7e-10
Identities = 41/135 (30%), Positives = 70/135 (51%), Gaps = 5/135 (3%)
Frame = +3
Query: 188 HSSSYGGHASTQKASFKILHSGFWWPSLFKDVHLFISKCDKCQRTGNITKRNEMPLNNIL 247
H G H+ +Q + ++L G++ ++ K ++ KC++C + GNI+ L+NI+
Sbjct: 9 HRGICGMHSKSQLMTTRVLRVGYY**TMRKYCTEYVKKCEEC*KFGNISHLLVEELHNIV 188
Query: 248 EVEIFDVWVIDFIGPFP-SSFVNQYILVAVDYVSKWVE----VIASLTNDAQVVIKLFKK 302
F + +D + PFP S +Y+LV +D +KW+E I S+ N V K +
Sbjct: 189 APWPFAI*GVDILRPFPLSKRQVKYLLVGIDQFTKWIETEHIAIISIAN----VRKFV*R 356
Query: 303 NHFSRFGVPRVVISD 317
N FG+P +ISD
Sbjct: 357 NIVC*FGIPNTLISD 401
>TC206741
Length = 1191
Score = 57.8 bits (138), Expect = 1e-08
Identities = 21/46 (45%), Positives = 34/46 (73%)
Frame = -1
Query: 204 KILHSGFWWPSLFKDVHLFISKCDKCQRTGNITKRNEMPLNNILEV 249
++ S F+WP+L+K + ++ CD CQRTG I++RNEMP N+I+ +
Sbjct: 312 ELKQSSFYWPTLWKGAYEYVQTCDNCQRTGGISRRNEMPSNSIIPI 175
>TC212032 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (3%)
Length = 803
Score = 53.5 bits (127), Expect = 2e-07
Identities = 25/91 (27%), Positives = 44/91 (47%)
Frame = +2
Query: 122 PWYADFVNFLAAGVLPPELSYQQKKKFFNDLKHYYSDEPYLFKRG*DGIFRRCIPENEVS 181
PWY D ++ + P E S K++ ++ L+KR D + C+ EV
Sbjct: 530 PWYFDIKRYVESKEYPLEASDNDKRRKRRLAAGFFMSGSILYKRNHDMVLLHCVNGKEVE 709
Query: 182 SILTHCHSSSYGGHASTQKASFKILHSGFWW 212
++L H S+G H++ + KIL +G++W
Sbjct: 710 NMLGEVHEGSFGTHSNGHAMARKILRAGYYW 802
>TC211067 similar to UP|Q9LQH2 (Q9LQH2) F15O4.13, partial (9%)
Length = 589
Score = 47.0 bits (110), Expect = 2e-05
Identities = 24/65 (36%), Positives = 39/65 (59%), Gaps = 5/65 (7%)
Frame = +1
Query: 1 IYYASKTLDGALVNYATTEKELLAVVYAIDKFRQYLVGSKITVYTDHSTIKYL-----LN 55
I Y S+ L A +NY T +KEL A++ A + +LV + +++DH ++KY+ LN
Sbjct: 385 IAYFSEKLHSATLNYPTYDKELYALIRAPQTWEHFLVCKEFVIHSDHQSLKYIRGKSKLN 564
Query: 56 KKDAK 60
K+ AK
Sbjct: 565 KRHAK 579
>CO982196
Length = 812
Score = 45.4 bits (106), Expect = 6e-05
Identities = 39/143 (27%), Positives = 66/143 (45%), Gaps = 4/143 (2%)
Frame = +1
Query: 179 EVSSILTHCHSSSYGGHASTQKASFKILHSGFWWPSLFKDVHLFISKCDKCQRTGNITKR 238
++ +L S GGH+ + +FK + + +W + K +++ C+ C+R T
Sbjct: 364 KIPLLLKELQDSPLGGHSGFFR-TFKRVANVVFWQGMKKTTRDYVAACEICRRNKTSTL- 537
Query: 239 NEMPLNNILEVEIFDVWV---IDFIGPFPSSFVNQYILVAVDYVSKWVEVIA-SLTNDAQ 294
+ L +L + VW +DFIG P + ILV VD ++K+ A S A+
Sbjct: 538 SPAGLL*LLPIPT-KVWTDISMDFIGGLPKAQGKDNILVVVDRLTKYAHFFALSHPYTAK 714
Query: 295 VVIKLFKKNHFSRFGVPRVVISD 317
V +LF K G P ++SD
Sbjct: 715 EVAELFIKELVRLHGFPASIVSD 783
>BQ627806
Length = 435
Score = 44.7 bits (104), Expect = 1e-04
Identities = 23/71 (32%), Positives = 39/71 (54%)
Frame = +1
Query: 12 LVNYATTEKELLAVVYAIDKFRQYLVGSKITVYTDHSTIKYLLNKKDAKPRLIQLIFLLQ 71
L+ +T +EL A+ A+ K+RQYL+G + TDH ++K L+++ P + L
Sbjct: 217 LLRASTYVRELAAITVAVKKWRQYLLGHHFVILTDHRSLKELMSQAVQTPEQQIYLARLM 396
Query: 72 EFDLEIKDKKG 82
FD I+ + G
Sbjct: 397 GFDYTIQYRAG 429
>AW507721 weakly similar to GP|27764548|gb polyprotein {Glycine max}, partial
(3%)
Length = 464
Score = 43.9 bits (102), Expect = 2e-04
Identities = 18/49 (36%), Positives = 31/49 (62%)
Frame = +1
Query: 183 ILTHCHSSSYGGHASTQKASFKILHSGFWWPSLFKDVHLFISKCDKCQR 231
I+T H+S GGHA T + +I ++ F+WP + +D+ F+ +C CQ+
Sbjct: 280 IMTESHASKVGGHAGTTRTIVRI-NAQFYWPKMREDIMKFVQECVICQQ 423
>BQ628592
Length = 423
Score = 34.3 bits (77), Expect = 0.15
Identities = 16/63 (25%), Positives = 32/63 (50%)
Frame = -3
Query: 13 VNYATTEKELLAVVYAIDKFRQYLVGSKITVYTDHSTIKYLLNKKDAKPRLIQLIFLLQE 72
+NY+ E+ +V+A + RQY++ + + +KY+ K R+ + LL E
Sbjct: 190 MNYSMLERTCCTLVWASHRLRQYMLSHTTWLISKMDPVKYIFEKPALTGRIARWQVLLSE 11
Query: 73 FDL 75
F++
Sbjct: 10 FNI 2
>BM084967
Length = 426
Score = 32.7 bits (73), Expect = 0.43
Identities = 17/53 (32%), Positives = 24/53 (45%)
Frame = -2
Query: 180 VSSILTHCHSSSYGGHASTQKASFKILHSGFWWPSLFKDVHLFISKCDKCQRT 232
+ ++L HSS H K + L F W + KDV F++ C CQ T
Sbjct: 302 IPTLLLEYHSSPTDAHIGVTKTMAR-LSENFTWIGIRKDVEQFVAACLDCQYT 147
>BI317638 weakly similar to GP|9294238|dbj| contains similarity to reverse
transcriptase~gene_id:K11J14.5 {Arabidopsis thaliana},
partial (5%)
Length = 420
Score = 30.4 bits (67), Expect = 2.1
Identities = 27/97 (27%), Positives = 46/97 (46%), Gaps = 2/97 (2%)
Frame = -2
Query: 223 ISKCDKCQRTGNITKRN-EMPLNNILEVEIFDVWVIDFIGPFPSSFVNQYILVAVDYVSK 281
+ C CQ T TKR ++ ++ ++ +DFI V+ ILV VD+ SK
Sbjct: 335 LPNCLDCQHTKYETKRIVDLLCPLLVPHRPWEDLSLDFITGLLPYHVHTAILVVVDHFSK 156
Query: 282 WVEV-IASLTNDAQVVIKLFKKNHFSRFGVPRVVISD 317
+ + + ++ A V LF + G+PR ++SD
Sbjct: 155 GIHLGMLPSSHTAHTVACLFIDSVAKLHGLPRSLVSD 45
>TC203554 homologue to UP|RL38_ARATH (O22860) 60S ribosomal protein L38,
complete
Length = 1822
Score = 29.6 bits (65), Expect = 3.6
Identities = 21/59 (35%), Positives = 32/59 (53%)
Frame = -1
Query: 47 HSTIKYLLNKKDAKPRLIQLIFLLQEFDLEIKDKKGVENVVADHLSRLRETNEDELPLD 105
HST +LN + + RL+QLI L+ +KD K V+ + A +L + N+ PLD
Sbjct: 1150 HSTALQVLNTQTGRKRLLQLIGFLR-----VKDTKRVQVLGASNL----KLNDIPAPLD 1001
>TC216028 homologue to UP|Q9ZSQ0 (Q9ZSQ0) Ethylene response sensor, partial
(78%)
Length = 1745
Score = 29.3 bits (64), Expect = 4.7
Identities = 14/44 (31%), Positives = 23/44 (51%)
Frame = -2
Query: 186 HCHSSSYGGHASTQKASFKILHSGFWWPSLFKDVHLFISKCDKC 229
H +S+++ GH S+ K S +G+W+P + V FI C
Sbjct: 331 HPNSNNFRGHISSHKRSDPGQWTGWWYPHSLRAVENFIDNRIVC 200
>TC218592 weakly similar to UP|Q9LP32 (Q9LP32) T9L6.1 protein, partial (37%)
Length = 715
Score = 28.5 bits (62), Expect = 8.1
Identities = 12/49 (24%), Positives = 23/49 (46%)
Frame = +3
Query: 243 LNNILEVEIFDVWVIDFIGPFPSSFVNQYILVAVDYVSKWVEVIASLTN 291
L N+L +++ +W IG +F+ +L + Y + W E + N
Sbjct: 357 LGNVLNLQVKGIW----IGMLFGTFIQTVVLTVITYKTDWDEQVTKARN 491
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.336 0.148 0.464
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 23,024,485
Number of Sequences: 63676
Number of extensions: 319598
Number of successful extensions: 2220
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 2203
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2214
length of query: 547
length of database: 12,639,632
effective HSP length: 102
effective length of query: 445
effective length of database: 6,144,680
effective search space: 2734382600
effective search space used: 2734382600
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC141865.9


