

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141863.5 - phase: 0
(139 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
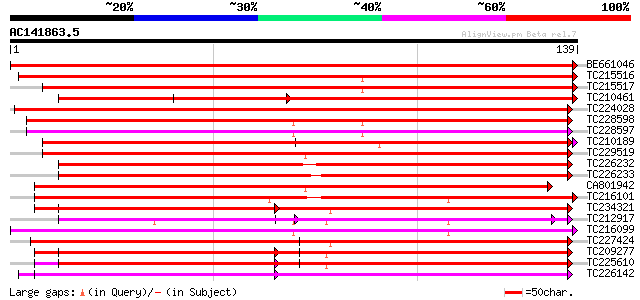
Score E
Sequences producing significant alignments: (bits) Value
BE661046 210 2e-55
TC215516 weakly similar to UP|Q8VWY6 (Q8VWY6) Pollen-specific ca... 189 4e-49
TC215517 186 3e-48
TC210461 164 1e-41
TC224028 similar to UP|Q9AXG2 (Q9AXG2) Regulator of gene silenci... 155 8e-39
TC228598 weakly similar to UP|Q8L3R2 (Q8L3R2) Calmodulin-like pr... 125 7e-30
TC228597 weakly similar to UP|Q93WY1 (Q93WY1) Calmodulin-like pr... 117 2e-27
TC210189 weakly similar to UP|Q8L3R2 (Q8L3R2) Calmodulin-like pr... 113 3e-26
TC229519 weakly similar to UP|Q93YA8 (Q93YA8) Calcium binding pr... 103 3e-23
TC226232 weakly similar to UP|Q93YA8 (Q93YA8) Calcium binding pr... 103 3e-23
TC226233 similar to UP|TCH2_ARATH (P25070) Calmodulin-related pr... 102 5e-23
CA801942 102 8e-23
TC216101 similar to UP|Q9SCA1 (Q9SCA1) Calcium-binding protein, ... 99 5e-22
TC234321 similar to UP|Q40982 (Q40982) Calmodulin-like protein, ... 95 1e-20
TC212917 weakly similar to UP|Q9LX27 (Q9LX27) Calmodulin-like pr... 92 6e-20
TC216099 similar to UP|Q9SCA1 (Q9SCA1) Calcium-binding protein, ... 91 2e-19
TC227424 UP|Q39890 (Q39890) Calmodulin, complete 90 3e-19
TC209277 homologue to UP|Q43447 (Q43447) Calmodulin, complete 90 3e-19
TC225610 homologue to UP|Q43447 (Q43447) Calmodulin, complete 90 4e-19
TC226142 similar to UP|Q93YA8 (Q93YA8) Calcium binding protein, ... 89 7e-19
>BE661046
Length = 486
Score = 210 bits (534), Expect = 2e-55
Identities = 99/139 (71%), Positives = 122/139 (87%)
Frame = +3
Query: 1 MKNAGFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLE 60
M+ A +E VLRYFDEDGDGK+SP+ELR R+ +MG E++LKEAEMAIEA+DSDGDG L L+
Sbjct: 54 MRGAEYERVLRYFDEDGDGKISPSELRNRIAMMGGEVMLKEAEMAIEALDSDGDGLLCLD 233
Query: 61 ELIALMEEGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKH 120
+L+ LME GEE+KLKDLREAF+MYD+E+CGFITPK+LKRMLKK+GESKS+ ECK MI
Sbjct: 234 DLMNLMEAAGEEEKLKDLREAFDMYDTERCGFITPKALKRMLKKLGESKSMVECKVMISR 413
Query: 121 FDLDGDGLLSFDEFITMMQ 139
FDL+GDG+LSF+EF MM+
Sbjct: 414 FDLNGDGMLSFEEFRIMMK 470
>TC215516 weakly similar to UP|Q8VWY6 (Q8VWY6) Pollen-specific
calcium-binding protein, partial (51%)
Length = 649
Score = 189 bits (480), Expect = 4e-49
Identities = 92/138 (66%), Positives = 112/138 (80%), Gaps = 1/138 (0%)
Frame = +1
Query: 3 NAGFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEEL 62
N FE VL+YFDEDGDGK+SP ELR RL ++G E+L K+AE IE +DSDGDG+LSLE+
Sbjct: 109 NTEFERVLKYFDEDGDGKISPCELRNRLGMIGGELLAKDAEKLIEELDSDGDGFLSLEDF 288
Query: 63 IALMEEGGEEQKLKDLREAFEMY-DSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHF 121
+ LME GE++KLKDL EAFEMY D+E GFITPKSL+RML ++GESKS+++C MI HF
Sbjct: 289 VKLMEAAGEDEKLKDLEEAFEMYNDTEMSGFITPKSLQRMLGRLGESKSMEQCTTMIGHF 468
Query: 122 DLDGDGLLSFDEFITMMQ 139
DL+GDGLL FDEF MMQ
Sbjct: 469 DLNGDGLLCFDEFRVMMQ 522
>TC215517
Length = 660
Score = 186 bits (472), Expect = 3e-48
Identities = 91/132 (68%), Positives = 111/132 (83%), Gaps = 1/132 (0%)
Frame = +2
Query: 9 VLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALMEE 68
VL+YFDEDGDGK+SP+ELR RL +MG +L K+AE IE +DSDGDG+LSLE+ + +ME
Sbjct: 5 VLKYFDEDGDGKISPSELRNRLGMMGGVLLFKDAEKLIEELDSDGDGFLSLEDFVKIMEA 184
Query: 69 GGEEQKLKDLREAFEMY-DSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDG 127
GEE+KLKDL EAFEMY DSE GFITPKSL++ML ++GESKS+++C AMI HFDL+GDG
Sbjct: 185 AGEEEKLKDLAEAFEMYHDSEMFGFITPKSLQKMLGRLGESKSMEQCTAMIGHFDLNGDG 364
Query: 128 LLSFDEFITMMQ 139
LLSFDEF MMQ
Sbjct: 365 LLSFDEFRVMMQ 400
>TC210461
Length = 495
Score = 164 bits (415), Expect = 1e-41
Identities = 82/99 (82%), Positives = 89/99 (89%)
Frame = +2
Query: 41 EAEMAIEAMDSDGDGYLSLEELIALMEEGGEEQKLKDLREAFEMYDSEKCGFITPKSLKR 100
EAEMAI A+DSDGDG LSLE+LIALME GGEEQKL DL+ AFEMYD+E CGFITPKSLKR
Sbjct: 2 EAEMAIAALDSDGDGLLSLEDLIALMEAGGEEQKLNDLKVAFEMYDTEGCGFITPKSLKR 181
Query: 101 MLKKMGESKSIDECKAMIKHFDLDGDGLLSFDEFITMMQ 139
MLKKMGESKSIDECKAMIK FDL+GDG+LS +EF MMQ
Sbjct: 182 MLKKMGESKSIDECKAMIKQFDLNGDGVLSIEEFRIMMQ 298
Score = 44.7 bits (104), Expect = 1e-05
Identities = 21/57 (36%), Positives = 35/57 (60%)
Frame = +2
Query: 13 FDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALMEEG 69
+D +G G ++P L++ L+ MGE + E + I+ D +GDG LS+EE +M+ G
Sbjct: 134 YDTEGCGFITPKSLKRMLKKMGESKSIDECKAMIKQFDLNGDGVLSIEEFRIMMQ*G 304
>TC224028 similar to UP|Q9AXG2 (Q9AXG2) Regulator of gene silencing, partial
(34%)
Length = 843
Score = 155 bits (391), Expect = 8e-39
Identities = 72/137 (52%), Positives = 101/137 (73%)
Frame = +1
Query: 2 KNAGFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEE 61
K + ++ V FDE+GD K+SP+ELRQ + +G E+ K+AE+A+ +D DGDG + E+
Sbjct: 289 KLSQYKRVFNQFDENGDSKISPSELRQCVEAIGGELSEKDAEVAVTLLDRDGDGLVGFED 468
Query: 62 LIALMEEGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHF 121
+ +EEG EE+K DL+EAF+ Y+ + G ITP+SLKRML ++GES+S+DECK MI F
Sbjct: 469 FVRFLEEGKEEEKEDDLKEAFKRYEMDGSGCITPRSLKRMLSRLGESRSLDECKVMIARF 648
Query: 122 DLDGDGLLSFDEFITMM 138
DLDGDG+L+FDEF MM
Sbjct: 649 DLDGDGVLTFDEFKVMM 699
>TC228598 weakly similar to UP|Q8L3R2 (Q8L3R2) Calmodulin-like protein,
partial (40%)
Length = 842
Score = 125 bits (314), Expect = 7e-30
Identities = 68/139 (48%), Positives = 85/139 (60%), Gaps = 5/139 (3%)
Frame = +3
Query: 5 GFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIA 64
G R+FD DGDGK+S ELR +GE + +EAE I +DSDGD L ++
Sbjct: 207 GLMEAFRHFDNDGDGKISAYELRSYFGSIGEHMSHEEAEGVIHDLDSDGDNLLDFKDFTK 386
Query: 65 LMEE---GGEEQKLKDLREAFEMY--DSEKCGFITPKSLKRMLKKMGESKSIDECKAMIK 119
LM+ G E DLR AFEM+ + E CG ITPK L+RML ++G+ KS DEC AMI
Sbjct: 387 LMKRDVGGDEHDDEGDLRRAFEMFVWEKEGCGCITPKGLQRMLHRLGDDKSYDECVAMID 566
Query: 120 HFDLDGDGLLSFDEFITMM 138
FD+D +GLL FDEF MM
Sbjct: 567 AFDIDHNGLLDFDEFYQMM 623
Score = 25.4 bits (54), Expect = 9.2
Identities = 10/26 (38%), Positives = 15/26 (57%)
Frame = +3
Query: 105 MGESKSIDECKAMIKHFDLDGDGLLS 130
M ++ I+ +HFD DGDG +S
Sbjct: 183 MSNNREINGLMEAFRHFDNDGDGKIS 260
>TC228597 weakly similar to UP|Q93WY1 (Q93WY1) Calmodulin-like protein
(Fragment), partial (52%)
Length = 774
Score = 117 bits (292), Expect = 2e-27
Identities = 63/138 (45%), Positives = 83/138 (59%), Gaps = 4/138 (2%)
Frame = +2
Query: 5 GFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIA 64
G R+FD DGDGK+S ELR +G+ + +EAE I +DSDGD L ++
Sbjct: 182 GLMEAFRHFDNDGDGKISAYELRSYFGSIGDHMSHEEAEGVIHDLDSDGDNLLDFKDFTK 361
Query: 65 LMEE--GGEEQKLKDLREAFEMY--DSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKH 120
LM+ G + DLR AFEM+ + E G ITPK L+RML ++G+ KS DEC MI
Sbjct: 362 LMKRDVGDDHDDEGDLRRAFEMFVWEKEGSGCITPKGLQRMLHRLGDDKSYDECVTMIDA 541
Query: 121 FDLDGDGLLSFDEFITMM 138
FD+D +G+L FDEF MM
Sbjct: 542 FDIDHNGVLDFDEFYQMM 595
>TC210189 weakly similar to UP|Q8L3R2 (Q8L3R2) Calmodulin-like protein,
partial (60%)
Length = 723
Score = 113 bits (283), Expect = 3e-26
Identities = 62/131 (47%), Positives = 82/131 (62%), Gaps = 1/131 (0%)
Frame = +2
Query: 9 VLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALMEE 68
V + D D DGK+S +EL +GE + K AE I DSDGD L + LM++
Sbjct: 200 VFDHLDIDKDGKISSSELMDYFASVGESLSHKVAERVINEFDSDGDELLDFGDFEKLMKQ 379
Query: 69 GGEEQKLKDLREAFEMYDSEK-CGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDG 127
E+ LR AFEM++ EK CG ITPK L++ML+++G+ KS DEC AMI+ FDLDG+G
Sbjct: 380 EDSEELEDVLRSAFEMFEVEKGCGCITPKGLQQMLRQLGDVKSHDECAAMIQAFDLDGNG 559
Query: 128 LLSFDEFITMM 138
L F+EF MM
Sbjct: 560 FLDFNEFQQMM 592
Score = 42.0 bits (97), Expect = 1e-04
Identities = 24/69 (34%), Positives = 38/69 (54%)
Frame = +2
Query: 71 EEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLS 130
++++LKD+ F+ D +K G I+ L +GES S + +I FD DGD LL
Sbjct: 179 DDERLKDV---FDHLDIDKDGKISSSELMDYFASVGESLSHKVAERVINEFDSDGDELLD 349
Query: 131 FDEFITMMQ 139
F +F +M+
Sbjct: 350 FGDFEKLMK 376
>TC229519 weakly similar to UP|Q93YA8 (Q93YA8) Calcium binding protein,
partial (67%)
Length = 1060
Score = 103 bits (257), Expect = 3e-23
Identities = 50/131 (38%), Positives = 82/131 (62%), Gaps = 1/131 (0%)
Frame = +3
Query: 9 VLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALMEE 68
+ FD++GDGK+S EL+ L +G + +E + +E +D +GDG++ L+E
Sbjct: 114 IFNKFDKNGDGKISVTELKDMLAALGSKTTDEELKRMMEELDQNGDGFIDLKEFADFHCN 293
Query: 69 GGE-EQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDG 127
GG + K+LR+AF++YD +K G I+ K L +L+ +GE S+ +C+ MI + D DGDG
Sbjct: 294 GGAGKDDSKELRDAFDLYDVDKNGLISAKELHDVLRNLGEKCSLSDCRRMISNVDADGDG 473
Query: 128 LLSFDEFITMM 138
++F+EF MM
Sbjct: 474 NVNFEEFKKMM 506
>TC226232 weakly similar to UP|Q93YA8 (Q93YA8) Calcium binding protein,
partial (67%)
Length = 876
Score = 103 bits (256), Expect = 3e-23
Identities = 50/126 (39%), Positives = 83/126 (65%)
Frame = +2
Query: 13 FDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALMEEGGEE 72
FD++GDGK+S AEL++ + +G + +E + + +D +GDGY+ L+E GG+
Sbjct: 305 FDKNGDGKISCAELKEMMVALGSKTTSEEVKRMMAELDRNGDGYIDLKEFGEFHCGGGDG 484
Query: 73 QKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSFD 132
++LREAFE+YD +K G I+ K L +++++GE S+ +C+ MI + D DGDG ++F+
Sbjct: 485 ---RELREAFELYDLDKNGLISAKELHSVMRRLGEKCSLSDCRRMIGNVDADGDGNVNFE 655
Query: 133 EFITMM 138
EF MM
Sbjct: 656 EFKKMM 673
>TC226233 similar to UP|TCH2_ARATH (P25070) Calmodulin-related protein 2,
touch-induced, partial (55%)
Length = 800
Score = 102 bits (255), Expect = 5e-23
Identities = 50/126 (39%), Positives = 82/126 (64%)
Frame = +3
Query: 13 FDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALMEEGGEE 72
FD++GDGK+S AEL++ + +G + E + + +D +GDGY+ L+E GG +
Sbjct: 237 FDKNGDGKISCAELKEMMAALGSKTTSDEVKRMMAELDRNGDGYIDLKEFGEFHCGGGGD 416
Query: 73 QKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSFD 132
+ +LREAFE+YD +K G I+ K L +++++GE S+ +C+ MI + D DGDG ++F+
Sbjct: 417 GR--ELREAFELYDLDKNGLISAKELHSVMRRLGEKCSLSDCRRMIGNVDADGDGNVNFE 590
Query: 133 EFITMM 138
EF MM
Sbjct: 591 EFKKMM 608
>CA801942
Length = 446
Score = 102 bits (253), Expect = 8e-23
Identities = 49/128 (38%), Positives = 81/128 (63%), Gaps = 1/128 (0%)
Frame = +2
Query: 7 EHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALM 66
+ + FD++GDGK+S AEL+ L +G + +E + IE +D +GDG++ L+E
Sbjct: 62 QQIFNKFDKNGDGKISMAELKDMLSALGSKTTDEELKRMIEELDQNGDGFIDLKEFADFH 241
Query: 67 EEGGE-EQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDG 125
GG + K+LR+AF++YD +K G I+ K L +L+ +GE S+ +C+ MI + D DG
Sbjct: 242 CNGGAGKDDSKELRDAFDLYDVDKNGLISAKELHHVLRNLGEKCSLSDCRRMISNVDGDG 421
Query: 126 DGLLSFDE 133
DG ++F+E
Sbjct: 422 DGNVNFEE 445
>TC216101 similar to UP|Q9SCA1 (Q9SCA1) Calcium-binding protein, partial
(65%)
Length = 796
Score = 99.4 bits (246), Expect = 5e-22
Identities = 54/139 (38%), Positives = 87/139 (61%), Gaps = 6/139 (4%)
Frame = +1
Query: 7 EHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEEL---- 62
+ V + FD +GDG++S EL L +G I K+ IE +D +GDG + ++E
Sbjct: 238 KRVFQMFDRNGDGRISLKELSDSLENLGILIPDKDLAQMIERIDVNGDGCVDMDEFGDLY 417
Query: 63 IALMEEGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMG--ESKSIDECKAMIKH 120
++MEE EE+ D+REAF ++D + GFIT + L+R+L +G + +++DECK MI
Sbjct: 418 ESIMEERDEEE---DMREAFNVFDQNRDGFITVEELRRVLASLGLKQGRTLDECKKMITK 588
Query: 121 FDLDGDGLLSFDEFITMMQ 139
D+DGDG++++ EF MM+
Sbjct: 589 VDVDGDGMVNYKEFRQMMK 645
Score = 38.1 bits (87), Expect = 0.001
Identities = 19/58 (32%), Positives = 31/58 (52%)
Frame = +1
Query: 77 DLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSFDEF 134
+L+ F+M+D G I+ K L L+ +G + MI+ D++GDG + DEF
Sbjct: 232 ELKRVFQMFDRNGDGRISLKELSDSLENLGILIPDKDLAQMIERIDVNGDGCVDMDEF 405
>TC234321 similar to UP|Q40982 (Q40982) Calmodulin-like protein, partial
(96%)
Length = 626
Score = 94.7 bits (234), Expect = 1e-20
Identities = 44/127 (34%), Positives = 81/127 (63%), Gaps = 1/127 (0%)
Frame = +3
Query: 13 FDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALMEEGGEE 72
FD+DGDG ++ EL +R + E ++E ++ + +D DG+G + E + LM +E
Sbjct: 126 FDKDGDGCITIEELSTAIRSLDENPTVEELQIMMNEVDMDGNGTIEFGEFLNLMARKMKE 305
Query: 73 QKLKD-LREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSF 131
+ ++ L+EAF ++D + G+I+P L+ +++ +GE + +E + M+K DLDGDGL+ +
Sbjct: 306 TEAEEELKEAFRVFDKDHDGYISPSELRSVMRTIGEKVTDEEVEQMVKEADLDGDGLIDY 485
Query: 132 DEFITMM 138
+EF+ MM
Sbjct: 486 EEFVRMM 506
Score = 56.2 bits (134), Expect = 5e-09
Identities = 24/60 (40%), Positives = 38/60 (63%)
Frame = +3
Query: 7 EHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALM 66
+ R FD+D DG +SP+ELR +R +GE++ +E E ++ D DGDG + EE + +M
Sbjct: 327 KEAFRVFDKDHDGYISPSELRSVMRTIGEKVTDEEVEQMVKEADLDGDGLIDYEEFVRMM 506
Score = 51.6 bits (122), Expect = 1e-07
Identities = 21/73 (28%), Positives = 45/73 (60%)
Frame = +3
Query: 66 MEEGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDG 125
M++ E+++ + EAF ++D + G IT + L ++ + E+ +++E + M+ D+DG
Sbjct: 69 MKDALREEQIGEFLEAFCLFDKDGDGCITIEELSTAIRSLDENPTVEELQIMMNEVDMDG 248
Query: 126 DGLLSFDEFITMM 138
+G + F EF+ +M
Sbjct: 249 NGTIEFGEFLNLM 287
>TC212917 weakly similar to UP|Q9LX27 (Q9LX27) Calmodulin-like protein,
partial (49%)
Length = 1086
Score = 92.4 bits (228), Expect = 6e-20
Identities = 52/139 (37%), Positives = 82/139 (58%), Gaps = 13/139 (9%)
Frame = +1
Query: 13 FDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALMEE--GG 70
FD++GDG ++ ELR+ LR +G + KE + + DS+ DG + EE L E GG
Sbjct: 343 FDKNGDGFITKQELRESLRNIGIFMADKEVDDIVVKYDSNSDGLIDFEEFCLLTSECVGG 522
Query: 71 EEQKLK---------DLREAFEMYDSEKCGFITPKSLKRMLKKMG--ESKSIDECKAMIK 119
+ + + DL+EAF+++D + G I+ + L +L +G E + I+ECK MIK
Sbjct: 523 DHHEKEGGVMGNEEVDLKEAFDVFDKDNDGLISVEELALVLTSLGLREGRKIEECKEMIK 702
Query: 120 HFDLDGDGLLSFDEFITMM 138
D+DGDG+++F+EF MM
Sbjct: 703 KVDMDGDGMVNFNEFKRMM 759
Score = 50.4 bits (119), Expect = 3e-07
Identities = 23/69 (33%), Positives = 40/69 (57%)
Frame = +1
Query: 66 MEEGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDG 125
M E G ++K ++LR+ F +D GFIT + L+ L+ +G + E ++ +D +
Sbjct: 286 MAESGSQKKKEELRKLFSTFDKNGDGFITKQELRESLRNIGIFMADKEVDDIVVKYDSNS 465
Query: 126 DGLLSFDEF 134
DGL+ F+EF
Sbjct: 466 DGLIDFEEF 492
Score = 39.3 bits (90), Expect = 6e-04
Identities = 22/61 (36%), Positives = 34/61 (55%), Gaps = 2/61 (3%)
Frame = +1
Query: 13 FDEDGDGKVSPAELRQRLRIMG--EEILLKEAEMAIEAMDSDGDGYLSLEELIALMEEGG 70
FD+D DG +S EL L +G E ++E + I+ +D DGDG ++ E +M GG
Sbjct: 592 FDKDNDGLISVEELALVLTSLGLREGRKIEECKEMIKKVDMDGDGMVNFNEFKRMMMNGG 771
Query: 71 E 71
+
Sbjct: 772 K 774
>TC216099 similar to UP|Q9SCA1 (Q9SCA1) Calcium-binding protein, partial
(63%)
Length = 946
Score = 90.9 bits (224), Expect = 2e-19
Identities = 48/142 (33%), Positives = 81/142 (56%), Gaps = 3/142 (2%)
Frame = +3
Query: 1 MKNAGFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLE 60
M+ + V + FD +GDG+++ EL L +G I K+ I+ +D +GDG + ++
Sbjct: 54 MEAQELKRVFQMFDRNGDGRITKKELNDSLENLGIFISDKDLSQMIQRIDVNGDGCVDMD 233
Query: 61 ELIALMEE-GGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMG--ESKSIDECKAM 117
E L + E +D+REAF ++D GFIT L+ +L +G + +++ +CKAM
Sbjct: 234 EFGELYQTIMDERDNEEDMREAFNVFDQNADGFITVDELRTVLSSLGLKQGRTVQDCKAM 413
Query: 118 IKHFDLDGDGLLSFDEFITMMQ 139
I D+DGDG++ + EF MM+
Sbjct: 414 ISKVDVDGDGMVDYKEFKQMMK 479
>TC227424 UP|Q39890 (Q39890) Calmodulin, complete
Length = 1424
Score = 90.1 bits (222), Expect = 3e-19
Identities = 43/134 (32%), Positives = 84/134 (62%), Gaps = 1/134 (0%)
Frame = +3
Query: 6 FEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIAL 65
F+ FD+DGDG ++ EL +R + + +E + I +D+DG+G + +E ++L
Sbjct: 693 FKEAFGLFDKDGDGCITVEELATVIRSLDQNPTEEELQDMISEVDADGNGTIEFDEFLSL 872
Query: 66 MEEGGEEQKLKD-LREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLD 124
M + ++ ++ L+EAF+++D ++ G+I+ L+ ++ +GE + +E + MIK DLD
Sbjct: 873 MAKKVKDTDAEEELKEAFKVFDKDQNGYISASELRHVMINLGEKLTDEEVEQMIKEADLD 1052
Query: 125 GDGLLSFDEFITMM 138
GDG ++++EF+ MM
Sbjct: 1053GDGQVNYEEFVKMM 1094
Score = 53.9 bits (128), Expect = 2e-08
Identities = 22/67 (32%), Positives = 44/67 (64%)
Frame = +3
Query: 72 EQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSF 131
E+++ D +EAF ++D + G IT + L +++ + ++ + +E + MI D DG+G + F
Sbjct: 675 EEQIVDFKEAFGLFDKDGDGCITVEELATVIRSLDQNPTEEELQDMISEVDADGNGTIEF 854
Query: 132 DEFITMM 138
DEF+++M
Sbjct: 855 DEFLSLM 875
>TC209277 homologue to UP|Q43447 (Q43447) Calmodulin, complete
Length = 873
Score = 90.1 bits (222), Expect = 3e-19
Identities = 45/127 (35%), Positives = 80/127 (62%), Gaps = 1/127 (0%)
Frame = +3
Query: 13 FDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALMEEGGEE 72
FD+DGDG ++ EL +R + + +E + I +D+DG+G + E + LM + +E
Sbjct: 126 FDKDGDGCITVEELATVIRSLDQNPTEEELQDMINEVDTDGNGTIEFVEFLNLMAKKMKE 305
Query: 73 QKLK-DLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSF 131
+ DL+EAF+++D ++ G+I+ L+ ++ +GE + +E + MIK DLDGDG +++
Sbjct: 306 TDAEEDLKEAFKVFDKDQNGYISASELRHVMINLGEKLTDEEVEQMIKEADLDGDGQVNY 485
Query: 132 DEFITMM 138
DEF+ MM
Sbjct: 486 DEFVKMM 506
Score = 49.3 bits (116), Expect = 6e-07
Identities = 20/67 (29%), Positives = 43/67 (63%)
Frame = +3
Query: 72 EQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSF 131
E+++ +++EAF ++D + G IT + L +++ + ++ + +E + MI D DG+G + F
Sbjct: 87 EEQIGEIKEAFGLFDKDGDGCITVEELATVIRSLDQNPTEEELQDMINEVDTDGNGTIEF 266
Query: 132 DEFITMM 138
EF+ +M
Sbjct: 267 VEFLNLM 287
Score = 45.4 bits (106), Expect = 9e-06
Identities = 20/60 (33%), Positives = 37/60 (61%)
Frame = +3
Query: 7 EHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALM 66
+ + FD+D +G +S +ELR + +GE++ +E E I+ D DGDG ++ +E + +M
Sbjct: 327 KEAFKVFDKDQNGYISASELRHVMINLGEKLTDEEVEQMIKEADLDGDGQVNYDEFVKMM 506
>TC225610 homologue to UP|Q43447 (Q43447) Calmodulin, complete
Length = 701
Score = 89.7 bits (221), Expect = 4e-19
Identities = 45/127 (35%), Positives = 79/127 (61%), Gaps = 1/127 (0%)
Frame = +3
Query: 13 FDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALMEEGGEE 72
FD+DGDG ++ EL +R + + +E + I +D+DG+G + E + LM + +E
Sbjct: 147 FDKDGDGCITVEELATVIRSLDQNPTEEELQDMINEVDTDGNGTIEFVEFLNLMAKKMKE 326
Query: 73 QKLK-DLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSF 131
+ DL+EAF+++D ++ G+I+ L+ ++ +GE + +E + MIK DLDGDG + +
Sbjct: 327 TDAEEDLKEAFKVFDKDQNGYISASELRHVMINLGEKLTDEEVEQMIKEADLDGDGQVGY 506
Query: 132 DEFITMM 138
DEF+ MM
Sbjct: 507 DEFVKMM 527
Score = 49.3 bits (116), Expect = 6e-07
Identities = 20/67 (29%), Positives = 43/67 (63%)
Frame = +3
Query: 72 EQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGDGLLSF 131
E+++ +++EAF ++D + G IT + L +++ + ++ + +E + MI D DG+G + F
Sbjct: 108 EEQIGEIKEAFGLFDKDGDGCITVEELATVIRSLDQNPTEEELQDMINEVDTDGNGTIEF 287
Query: 132 DEFITMM 138
EF+ +M
Sbjct: 288 VEFLNLM 308
Score = 45.1 bits (105), Expect = 1e-05
Identities = 20/60 (33%), Positives = 36/60 (59%)
Frame = +3
Query: 7 EHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALM 66
+ + FD+D +G +S +ELR + +GE++ +E E I+ D DGDG + +E + +M
Sbjct: 348 KEAFKVFDKDQNGYISASELRHVMINLGEKLTDEEVEQMIKEADLDGDGQVGYDEFVKMM 527
>TC226142 similar to UP|Q93YA8 (Q93YA8) Calcium binding protein, partial
(89%)
Length = 572
Score = 89.0 bits (219), Expect = 7e-19
Identities = 46/132 (34%), Positives = 75/132 (55%)
Frame = +3
Query: 7 EHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEELIALM 66
+ V FD + DGK+S EL LR +G + ++ + ++ +D+D DG+++L E A
Sbjct: 165 KRVFSRFDANCDGKISVTELDNVLRSLGSGVPPEDIQRVMDDLDTDHDGFINLSEFAAFC 344
Query: 67 EEGGEEQKLKDLREAFEMYDSEKCGFITPKSLKRMLKKMGESKSIDECKAMIKHFDLDGD 126
+ +L +AF +YD +K G I+ L ++L ++G S++EC MIK D DGD
Sbjct: 345 RSDTADGGDAELHDAFNLYDHDKNGHISATELCQVLNRLGMKCSVEECHNMIKSVDSDGD 524
Query: 127 GLLSFDEFITMM 138
G ++F EF MM
Sbjct: 525 GNVNFPEFKRMM 560
Score = 39.7 bits (91), Expect = 5e-04
Identities = 20/64 (31%), Positives = 34/64 (52%)
Frame = +3
Query: 3 NAGFEHVLRYFDEDGDGKVSPAELRQRLRIMGEEILLKEAEMAIEAMDSDGDGYLSLEEL 62
+A +D D +G +S EL Q L +G + ++E I+++DSDGDG ++ E
Sbjct: 369 DAELHDAFNLYDHDKNGHISATELCQVLNRLGMKCSVEECHNMIKSVDSDGDGNVNFPEF 548
Query: 63 IALM 66
+M
Sbjct: 549 KRMM 560
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.317 0.138 0.379
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,097,886
Number of Sequences: 63676
Number of extensions: 37130
Number of successful extensions: 523
Number of sequences better than 10.0: 131
Number of HSP's better than 10.0 without gapping: 313
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 421
length of query: 139
length of database: 12,639,632
effective HSP length: 87
effective length of query: 52
effective length of database: 7,099,820
effective search space: 369190640
effective search space used: 369190640
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 54 (25.4 bits)
Medicago: description of AC141863.5


