

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141323.9 - phase: 0
(523 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
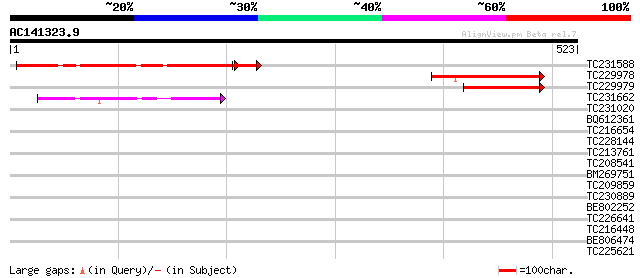
Score E
Sequences producing significant alignments: (bits) Value
TC231588 similar to UP|Q9FIN8 (Q9FIN8) Arabidopsis thaliana geno... 268 2e-75
TC229978 similar to UP|Q9FIN8 (Q9FIN8) Arabidopsis thaliana geno... 196 3e-50
TC229979 similar to UP|Q9FIN8 (Q9FIN8) Arabidopsis thaliana geno... 153 2e-37
TC231662 weakly similar to GB|CAA80137.1|3881488|CEZK1098 C. ele... 83 3e-16
TC231020 34 0.18
BQ612361 34 0.18
TC216654 32 0.90
TC228144 homologue to UP|Q76DY3 (Q76DY3) AG-motif binding protei... 31 1.5
TC213761 30 2.0
TC208541 homologue to UP|Q76NE1 (Q76NE1) ORF55b, partial (22%) 30 3.4
BM269751 29 4.5
TC209859 similar to UP|Q94BY1 (Q94BY1) AT3g52150/F4F15_260, part... 29 4.5
TC230889 29 5.8
BE802252 similar to SP|P00865|RBS1 Ribulose bisphosphate carboxy... 28 7.6
TC226641 weakly similar to UP|ORM1_YEAST (P53224) ORM1 protein, ... 28 7.6
TC216448 28 7.6
BE806474 28 10.0
TC225621 similar to UP|Q6NQE2 (Q6NQE2) At4g27270, partial (86%) 28 10.0
>TC231588 similar to UP|Q9FIN8 (Q9FIN8) Arabidopsis thaliana genomic DNA,
chromosome 5, TAC clone:K16H17 (At5g24340), partial
(18%)
Length = 678
Score = 268 bits (684), Expect(2) = 2e-75
Identities = 143/206 (69%), Positives = 161/206 (77%)
Frame = +2
Query: 7 KPLEIHFVTTTDSPEFTHLTRSLTQTSLVGLDAEWKPVRTHQNSFPTVSLLQIACQLGDD 66
K L++H VT TDS EF L+ +LT+TS+VGLDAEWKPVR FP V++LQIAC D
Sbjct: 32 KLLKVHLVTCTDSAEFALLSSALTRTSVVGLDAEWKPVR---RLFPRVAVLQIAC---GD 193
Query: 67 EVVFLLDLISLPLSSIWEPLREMLVSADILKLGFRFKQDLVYLSSTFCEQGCNPGFDKVE 126
VFLLDL+SLPLSS+W PLRE+L+S DILKLGF FKQDLVYLSSTF G GFDKVE
Sbjct: 194 SAVFLLDLLSLPLSSLWAPLRELLLSPDILKLGFGFKQDLVYLSSTFASHG---GFDKVE 364
Query: 127 PYLDITSVYNHLQFKKNGRIASKQNKSLSTICGELLGITLSKELQCSDWSQRPLTEEQMT 186
PYLDI SVYNHLQ K + KQ+KSLSTIC E+LG +LSKELQCSDWS RPLTEEQ+T
Sbjct: 365 PYLDIKSVYNHLQHNK--KHVPKQSKSLSTICAEVLGFSLSKELQCSDWSHRPLTEEQIT 538
Query: 187 YAAMDAHCLLGIFKVFQATVAKEGEL 212
YAAMDAHCLL IF+VFQA V K L
Sbjct: 539 YAAMDAHCLLDIFEVFQAKVVKRRRL 616
Score = 33.5 bits (75), Expect(2) = 2e-75
Identities = 15/27 (55%), Positives = 22/27 (80%)
Frame = +3
Query: 206 VAKEGELVNKTNILSIRSANLGLKELF 232
++KEG+L+ +T +LS A+LGLKELF
Sbjct: 597 LSKEGDLILETTVLSNPDASLGLKELF 677
>TC229978 similar to UP|Q9FIN8 (Q9FIN8) Arabidopsis thaliana genomic DNA,
chromosome 5, TAC clone:K16H17 (At5g24340), partial
(25%)
Length = 648
Score = 196 bits (497), Expect = 3e-50
Identities = 93/108 (86%), Positives = 100/108 (92%), Gaps = 4/108 (3%)
Frame = +2
Query: 390 AQKEKRVLLTRDAKLLRHDY----QIYKVKSLLKNEQLLEIIETFQLNINEDQLMSRCTK 445
AQKEKRV+LT DAKLLRHDY QIY+VKSLLKNEQLLE+IE FQ+ INED+LMSRCTK
Sbjct: 2 AQKEKRVILTWDAKLLRHDYLTQNQIYRVKSLLKNEQLLEVIEAFQIKINEDKLMSRCTK 181
Query: 446 CNGRFIQKPLSTEEAIEAAKGFQKIPNCLFNKNLEFWQCMDCHQLYWE 493
CNG FIQKPL+TEEAIEAAKGFQ+IPNCLFNKNLEFWQCMDCHQLYWE
Sbjct: 182 CNGTFIQKPLTTEEAIEAAKGFQRIPNCLFNKNLEFWQCMDCHQLYWE 325
>TC229979 similar to UP|Q9FIN8 (Q9FIN8) Arabidopsis thaliana genomic DNA,
chromosome 5, TAC clone:K16H17 (At5g24340), partial
(18%)
Length = 535
Score = 153 bits (387), Expect = 2e-37
Identities = 68/75 (90%), Positives = 72/75 (95%)
Frame = +2
Query: 419 KNEQLLEIIETFQLNINEDQLMSRCTKCNGRFIQKPLSTEEAIEAAKGFQKIPNCLFNKN 478
KNEQLLE+IE FQ+ INEDQLMSRCTKCNG FIQKPL+TEEAIEAAKGFQ+IPNCLFNKN
Sbjct: 2 KNEQLLEVIEAFQIKINEDQLMSRCTKCNGTFIQKPLTTEEAIEAAKGFQRIPNCLFNKN 181
Query: 479 LEFWQCMDCHQLYWE 493
LEFWQCMDCHQLYWE
Sbjct: 182 LEFWQCMDCHQLYWE 226
>TC231662 weakly similar to GB|CAA80137.1|3881488|CEZK1098 C. elegans MUT-7
protein (corresponding sequence ZK1098.8)
{Caenorhabditis elegans;} , partial (5%)
Length = 794
Score = 83.2 bits (204), Expect = 3e-16
Identities = 60/177 (33%), Positives = 93/177 (51%), Gaps = 3/177 (1%)
Frame = +2
Query: 26 TRSLTQTSLVGLDAEWKPVRTHQNSFPTVSLLQIACQLGDDEVVFLLDLISLPLSS---I 82
TR + ++GLD EWKP + VS++QIA +++VF+ DLI L +
Sbjct: 83 TRHIKGFKVIGLDCEWKPNYVKGSKPNKVSIMQIA----SEKMVFIFDLIKLHKEVPDIL 250
Query: 83 WEPLREMLVSADILKLGFRFKQDLVYLSSTFCEQGCNPGFDKVEPYLDITSVYNHLQFKK 142
+ L +L+S ILKLG+ F+ D L+ ++ E C F E LDI +V+
Sbjct: 251 DDCLSCILLSPRILKLGYNFQCDAKQLAYSYEELRC---FKNYEMLLDIQNVFK------ 403
Query: 143 NGRIASKQNKSLSTICGELLGITLSKELQCSDWSQRPLTEEQMTYAAMDAHCLLGIF 199
+ L+ + ++LG +L+K + S+W QRPLT Q+ YAA+DA L+ IF
Sbjct: 404 ------EPRGGLAGLAEKILGASLNKTRRNSNWEQRPLTPNQLEYAALDAVVLVHIF 556
>TC231020
Length = 704
Score = 33.9 bits (76), Expect = 0.18
Identities = 47/174 (27%), Positives = 78/174 (44%), Gaps = 5/174 (2%)
Frame = +2
Query: 24 HLTRSLTQTSLVGLDAEWKPVRTHQNSFPTVSLLQIACQLGDDEVVFLLDLISLPLSSIW 83
H R LT VGLD EW+P T +N V+ LQ+ E + ++ P SI
Sbjct: 86 HQQRVLT----VGLDIEWRP-NTQRNMQNPVATLQLCVA----ERCLVFQILHSP--SIP 232
Query: 84 EPLREMLVSADILKLGFRFKQDLVYLSSTFCEQGCNPGFDKVEPY-LDITSVYNHLQF-- 140
L L +I +G ++D+ L +E Y L++ +V + F
Sbjct: 233 PSLVSFLADPNITFVGVGIQEDVEKL---------------LEDYNLNVANVRDLRSFAA 367
Query: 141 KKNGRIASKQNKSLSTICGELLGITLSKELQC--SDWSQRPLTEEQMTYAAMDA 192
++ G + K+ L ++ +LG+ ++K + S W LT +Q+ YAA+DA
Sbjct: 368 ERLGDLELKR-AGLKSLGLRVLGLEVAKPKRVTRSRWDNPWLTAQQVQYAAVDA 526
>BQ612361
Length = 432
Score = 33.9 bits (76), Expect = 0.18
Identities = 19/63 (30%), Positives = 30/63 (47%), Gaps = 2/63 (3%)
Frame = +2
Query: 134 VYNHLQFKKNGRIASKQNKSLSTICGELLGITLSKELQCSDWSQRPLTEEQM--TYAAMD 191
V N ++ R+ SL + + G+T +KE Q +DW RPL + + T A M
Sbjct: 134 VCNMFDTGQSSRVLKLDRYSLQYLLQQFCGVTANKEYQSADWRLRPLPDVMLRFTSAVMM 313
Query: 192 AHC 194
+ C
Sbjct: 314 SAC 322
>TC216654
Length = 1170
Score = 31.6 bits (70), Expect = 0.90
Identities = 21/53 (39%), Positives = 27/53 (50%), Gaps = 3/53 (5%)
Frame = +1
Query: 474 LFNKNLEFWQCMDCHQLYWE---VLFTCSVCCELLFTIALSFILNKCVCHKIN 523
LF KN C +C Y+ +LFTCS C LLF F+LN + KI+
Sbjct: 979 LFRKNPP*GSC-NCSNTYFTYSVILFTCSGQCMLLFPPIC*FVLNPLLNEKIS 1134
>TC228144 homologue to UP|Q76DY3 (Q76DY3) AG-motif binding protein-1, partial
(27%)
Length = 1018
Score = 30.8 bits (68), Expect = 1.5
Identities = 20/56 (35%), Positives = 28/56 (49%), Gaps = 2/56 (3%)
Frame = +1
Query: 460 AIEAAKGFQKIPNCLFNKNLEFWQCMDCHQLYWEVLFTCS--VCCELLFTIALSFI 513
AI +A + IPN + LE+ C L W+ LF C C +LF I +SF+
Sbjct: 529 AINSAASPELIPNTNSSLTLEYM*CEGGLILCWKFLFHCCPLESCLMLFLIDISFL 696
>TC213761
Length = 708
Score = 30.4 bits (67), Expect = 2.0
Identities = 17/49 (34%), Positives = 26/49 (52%), Gaps = 2/49 (4%)
Frame = -1
Query: 472 NCL--FNKNLEFWQCMDCHQLYWEVLFTCSVCCELLFTIALSFILNKCV 518
NC F + LE W+C+ CH + ++F+C +F I F L KC+
Sbjct: 192 NCTDPFLQVLEVWKCICCHSI---LIFSCLWTTFSIFVIRQIFNLCKCL 55
>TC208541 homologue to UP|Q76NE1 (Q76NE1) ORF55b, partial (22%)
Length = 928
Score = 29.6 bits (65), Expect = 3.4
Identities = 11/27 (40%), Positives = 16/27 (58%)
Frame = +1
Query: 487 CHQLYWEVLFTCSVCCELLFTIALSFI 513
C +YW +L S C +LF + +SFI
Sbjct: 451 CSDIYWFMLCLISCCVAILFDLCISFI 531
>BM269751
Length = 394
Score = 29.3 bits (64), Expect = 4.5
Identities = 14/32 (43%), Positives = 19/32 (58%)
Frame = +3
Query: 286 FLLKIVRKHGDRILLKESDRAPKTSKKKRKKQ 317
FL K + HG LLK+ + K KKK+KK+
Sbjct: 45 FLNK*IMAHGHYTLLKKKKKKKKKKKKKKKKK 140
>TC209859 similar to UP|Q94BY1 (Q94BY1) AT3g52150/F4F15_260, partial (20%)
Length = 566
Score = 29.3 bits (64), Expect = 4.5
Identities = 11/28 (39%), Positives = 19/28 (67%)
Frame = -2
Query: 7 KPLEIHFVTTTDSPEFTHLTRSLTQTSL 34
KP+ +H +TTT+ EFT + + T +S+
Sbjct: 499 KPVVVHVLTTTEQHEFTSMNKCSTPSSI 416
>TC230889
Length = 852
Score = 28.9 bits (63), Expect = 5.8
Identities = 21/78 (26%), Positives = 34/78 (42%), Gaps = 1/78 (1%)
Frame = +2
Query: 291 VRKHGDRILLKESDRAPKTSKKKRKKQLPINGIPKEKHLENF-DEWQGTAPWDPLVGGDG 349
V +HG+ + P ++ K Q G ++K ++ + +EW T W + DG
Sbjct: 455 VEEHGESKQTGMISQLPNDAQDMSKTQ---QGKRRKKVVKRWREEWADTYKWAYVDMKDG 625
Query: 350 FPKFLCDVMVEGLAKHLR 367
P+ C V E KH R
Sbjct: 626 TPRIFCSVCREYGRKHRR 679
>BE802252 similar to SP|P00865|RBS1 Ribulose bisphosphate carboxylase small
chain 1 chloroplast precursor (EC 4.1.1.39), partial
(94%)
Length = 532
Score = 28.5 bits (62), Expect = 7.6
Identities = 14/63 (22%), Positives = 33/63 (52%)
Frame = +2
Query: 204 ATVAKEGELVNKTNILSIRSANLGLKELFRKHDTSDKVHSTQFCEALAIVQATSCSDVVR 263
A +AKE + + + + NL ++R+H+TS + + ++C ++ + C+D
Sbjct: 233 AQLAKEVKYLLRKGWIPCLEFNLDHSSVYREHNTSPRYYDRRYC-TMSNLTMLGCTDASH 409
Query: 264 VIS 266
V++
Sbjct: 410 VLN 418
>TC226641 weakly similar to UP|ORM1_YEAST (P53224) ORM1 protein, partial (21%)
Length = 1183
Score = 28.5 bits (62), Expect = 7.6
Identities = 10/19 (52%), Positives = 14/19 (73%)
Frame = +3
Query: 501 CCELLFTIALSFILNKCVC 519
CC+LLF + L +LNK +C
Sbjct: 1038 CCQLLFKLLLKVLLNK*IC 1094
>TC216448
Length = 739
Score = 28.5 bits (62), Expect = 7.6
Identities = 12/43 (27%), Positives = 18/43 (40%)
Frame = +2
Query: 478 NLEFWQCMDCHQLYWEVLFTCSVCCELLFTIALSFILNKCVCH 520
N W +D H W ++F VC + + F+L CH
Sbjct: 347 NYFLWHFIDLHSNVWLLMFPLEVCLNFMAAGSWCFMLILFCCH 475
>BE806474
Length = 429
Score = 28.1 bits (61), Expect = 10.0
Identities = 19/50 (38%), Positives = 25/50 (50%), Gaps = 6/50 (12%)
Frame = +1
Query: 472 NCLFNKNLEFWQCMDCHQLYWE------VLFTCSVCCELLFTIALSFILN 515
N L N NLE ++C+D L WE LF + LL T+ S I+N
Sbjct: 115 NVLINLNLEAYKCLD*FLLLWESMVFHDFLFFA*LELLLLTTLTQSNIIN 264
>TC225621 similar to UP|Q6NQE2 (Q6NQE2) At4g27270, partial (86%)
Length = 771
Score = 28.1 bits (61), Expect = 10.0
Identities = 21/71 (29%), Positives = 29/71 (40%)
Frame = -2
Query: 114 CEQGCNPGFDKVEPYLDITSVYNHLQFKKNGRIASKQNKSLSTICGELLGITLSKELQCS 173
C Q P +K+EP L + +HLQ K + S + S C + CS
Sbjct: 512 CRQSIYPD-EKLEPILTH*AALSHLQHKFLHHMGSHLSPFPSQTCHHQM---------CS 363
Query: 174 DWSQRPLTEEQ 184
W QR + EQ
Sbjct: 362 QWGQRSIHGEQ 330
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.321 0.136 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 25,349,918
Number of Sequences: 63676
Number of extensions: 383686
Number of successful extensions: 2150
Number of sequences better than 10.0: 43
Number of HSP's better than 10.0 without gapping: 2122
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2141
length of query: 523
length of database: 12,639,632
effective HSP length: 102
effective length of query: 421
effective length of database: 6,144,680
effective search space: 2586910280
effective search space used: 2586910280
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 61 (28.1 bits)
Medicago: description of AC141323.9


