

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141322.10 + phase: 0 /pseudo
(409 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
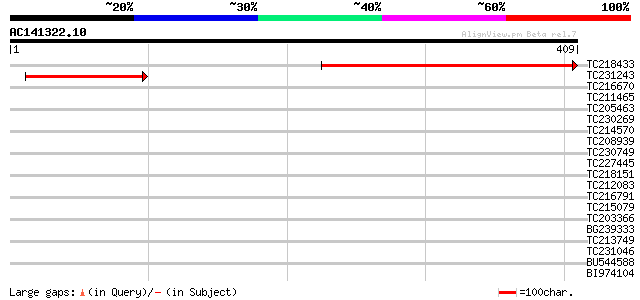
Score E
Sequences producing significant alignments: (bits) Value
TC218433 similar to UP|Q7XAL4 (Q7XAL4) Mitochondrial inner membr... 274 5e-74
TC231243 132 2e-31
TC216670 weakly similar to UP|Q39364 (Q39364) Non-green plastid ... 40 0.002
TC211465 similar to UP|CLPA_PEA (P35100) ATP-dependent Clp prote... 35 0.047
TC205463 UP|Q39804 (Q39804) BiP isoform B, complete 34 0.14
TC230269 weakly similar to UP|WSC2_YEAST (P53832) Cell wall inte... 34 0.14
TC214570 homologue to PIR|T05766|T05766 peptidylprolyl isomerase... 34 0.14
TC208939 UP|Q39830 (Q39830) BiP isoform A, complete 33 0.18
TC230749 similar to UP|Q9LYV6 (Q9LYV6) ABA-responsive protein-li... 33 0.23
TC227445 homologue to UP|O22639 (O22639) Endoplasmic reticulum H... 33 0.23
TC218151 homologue to UP|O22639 (O22639) Endoplasmic reticulum H... 33 0.23
TC212083 similar to UP|Q71BZ1 (Q71BZ1) Type-B response regulator... 33 0.30
TC216791 similar to UP|O23144 (O23144) Proton pump interactor (A... 32 0.67
TC215079 similar to UP|Q6H730 (Q6H730) Ribosomal protein L12-lik... 32 0.67
TC203366 similar to UP|Q93Z89 (Q93Z89) Matrix metalloproteinase ... 32 0.67
BG239333 31 1.2
TC213749 30 1.5
TC231046 30 2.0
BU544588 30 2.6
BI974104 29 3.3
>TC218433 similar to UP|Q7XAL4 (Q7XAL4) Mitochondrial inner membrane
translocating protein-like protein, partial (30%)
Length = 859
Score = 274 bits (701), Expect = 5e-74
Identities = 131/185 (70%), Positives = 156/185 (83%), Gaps = 1/185 (0%)
Frame = +3
Query: 226 YIDPVKTTSRDIVDDVRDIIDRIDNPIIDKIQSNMSRTFQETDAALTYREIRKRDPKFSL 285
Y DPVKT ++IV+D+R+ + D+PII KIQ FQETDAA++Y+EIR+RDP FSL
Sbjct: 3 YSDPVKTKGQEIVEDLRERYETSDSPIIHKIQDINDSMFQETDAAISYKEIRQRDPYFSL 182
Query: 286 PDFVGEVQEAIKPVLNAYIKGDFETLKKYCAPQLIERCKAEHGAYKDRGIFYDNKILHIS 345
P+FVGEVQEAIKPVLNAYIKGD ETLKKYC+P+LIERCKAEH AY+ GIF+DNKILH+S
Sbjct: 183 PEFVGEVQEAIKPVLNAYIKGDVETLKKYCSPELIERCKAEHNAYQSHGIFFDNKILHVS 362
Query: 346 DADVREVKILESSPFIIVVFQTQQIHCVRDRNGEITEGGKDTIHSVYYLWAL-QMDSEDH 404
D ++RE K++ SSP IIV+FQTQQI+CVRDRNG ITEGGKDTIH+V+Y WAL QMD ED
Sbjct: 363 DLEIRETKMMGSSPVIIVMFQTQQIYCVRDRNGAITEGGKDTIHTVFYFWALQQMDQEDR 542
Query: 405 AEDGI 409
EDGI
Sbjct: 543 GEDGI 557
>TC231243
Length = 551
Score = 132 bits (333), Expect = 2e-31
Identities = 64/88 (72%), Positives = 77/88 (86%)
Frame = +1
Query: 12 DRRGYSVFNEFSKKVKDETVKNPEFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQLYRQ 71
DRR YSVFNEFSK +K + V+N EFQ+SVKELKEKA+ELKG+KE LKE+TKQ TE+LY+Q
Sbjct: 277 DRRRYSVFNEFSKNIKGQAVRNQEFQQSVKELKEKADELKGVKEELKERTKQKTEKLYKQ 456
Query: 72 FDSVWKEAEAAAKKVSHNVKEKISAATD 99
D W EAEAAA+KVS+NVKEKISAA++
Sbjct: 457 VDEAWTEAEAAARKVSYNVKEKISAASE 540
>TC216670 weakly similar to UP|Q39364 (Q39364) Non-green plastid inner
envelope membrane protein precursor, partial (24%)
Length = 1453
Score = 39.7 bits (91), Expect = 0.002
Identities = 27/83 (32%), Positives = 46/83 (54%), Gaps = 5/83 (6%)
Frame = +1
Query: 24 KKVKDETVKNPEFQKSVKE----LKEKAEELKGI-KEGLKEKTKQTTEQLYRQFDSVWKE 78
+K +D + E Q++ K+ +E+AE+ +G+ +E + +K+T EQL D E
Sbjct: 400 EKERDVHAGSEESQEAWKQALDTFREQAEKFQGVSQEAYEVYSKKTAEQLKVLADKTKNE 579
Query: 79 AEAAAKKVSHNVKEKISAATDFS 101
AAK+++ KE +SAA D S
Sbjct: 580 LSVAAKEITDEGKEYLSAAADSS 648
>TC211465 similar to UP|CLPA_PEA (P35100) ATP-dependent Clp protease
ATP-binding subunit clpA homolog, chloroplast precursor
, partial (15%)
Length = 593
Score = 35.4 bits (80), Expect = 0.047
Identities = 16/38 (42%), Positives = 24/38 (63%)
Frame = -2
Query: 30 TVKNPEFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQ 67
T+ NPE QK + K+K +EL IKE KE+ ++ E+
Sbjct: 130 TISNPEIQKGKERKKDKKDELISIKERKKERERRERER 17
>TC205463 UP|Q39804 (Q39804) BiP isoform B, complete
Length = 2383
Score = 33.9 bits (76), Expect = 0.14
Identities = 26/89 (29%), Positives = 43/89 (48%)
Frame = +1
Query: 35 EFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQLYRQFDSVWKEAEAAAKKVSHNVKEKI 94
E ++ V+E +E AEE K +KE + + T +Y + + + + A K+ + KEKI
Sbjct: 1720 EIERMVREAEEFAEEDKKVKERIDARNSLET-YVYNMKNQI-SDKDKLADKLESDEKEKI 1893
Query: 95 SAATDFSTKQNADAKQGSQKSPEEEKNEE 123
A + D Q +K EEK +E
Sbjct: 1894 ETAVK-EALEWLDDNQSMEKEDYEEKLKE 1977
>TC230269 weakly similar to UP|WSC2_YEAST (P53832) Cell wall integrity and
stress response component 2 precursor, partial (6%)
Length = 696
Score = 33.9 bits (76), Expect = 0.14
Identities = 26/108 (24%), Positives = 51/108 (47%)
Frame = +2
Query: 24 KKVKDETVKNPEFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQLYRQFDSVWKEAEAAA 83
KKV +ETV+ +K K+ K+K +E + +EKT++ + + + E + +
Sbjct: 92 KKVDEETVEVEVEKKEKKKKKKKDKENSEVASSDEEKTEKKKNK--NKIEDGSPELDKSE 265
Query: 84 KKVSHNVKEKISAATDFSTKQNADAKQGSQKSPEEEKNEESPSGNASE 131
KK + +AA + S N + E++KN++ + +A E
Sbjct: 266 KKKKKKKDKAAAAAAEIS---NGKVDDSNADKSEKKKNKKKKNKDAKE 400
>TC214570 homologue to PIR|T05766|T05766 peptidylprolyl isomerase M4E13.20 -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(13%)
Length = 944
Score = 33.9 bits (76), Expect = 0.14
Identities = 24/104 (23%), Positives = 54/104 (51%), Gaps = 3/104 (2%)
Frame = +2
Query: 24 KKVKDETVKNPEFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQLYRQFDSVWKEAEAAA 83
KKV +ETV+ +K K+ K+K +E +EKT++ ++ + + E + +
Sbjct: 446 KKVDEETVEVEVEKKEKKKKKKKNKENSEAASSDEEKTEKKKKKHKDKVEDSSPELDKSE 625
Query: 84 KKVSHNVKEKISAAT---DFSTKQNADAKQGSQKSPEEEKNEES 124
KK ++ +AAT + +++A + +K +++KN+++
Sbjct: 626 KKKKKKKDKEAAAATAEISNGKEDDSNADKSEKKKHKKKKNKDA 757
>TC208939 UP|Q39830 (Q39830) BiP isoform A, complete
Length = 2452
Score = 33.5 bits (75), Expect = 0.18
Identities = 26/89 (29%), Positives = 43/89 (48%)
Frame = +3
Query: 35 EFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQLYRQFDSVWKEAEAAAKKVSHNVKEKI 94
E ++ V+E +E AEE K +KE + + T +Y + V + + A K+ + KEK+
Sbjct: 1686 EIERMVREAEEFAEEDKKVKERIDARNSLET-YVYNMKNQV-SDKDKLADKLESDEKEKV 1859
Query: 95 SAATDFSTKQNADAKQGSQKSPEEEKNEE 123
A + D Q +K EEK +E
Sbjct: 1860 ETAVK-EALEWLDDNQSVEKEEYEEKLKE 1943
>TC230749 similar to UP|Q9LYV6 (Q9LYV6) ABA-responsive protein-like
(At5g13200), partial (58%)
Length = 972
Score = 33.1 bits (74), Expect = 0.23
Identities = 26/97 (26%), Positives = 44/97 (44%), Gaps = 3/97 (3%)
Frame = +3
Query: 57 LKEKTKQTTEQLYRQFDSVWKEAEAAAKKVSHNVKEKISAATDFSTKQNADAK---QGSQ 113
L + + E + FDS ++AEA A V HN+K S ++ K N K +G
Sbjct: 225 LDKPSNSPMESILNMFDSWSRKAEATAHNVWHNLKTGPSVSSAALGKMNLTVKAISEGGF 404
Query: 114 KSPEEEKNEESPSGNASESLFGKFKSTFSSPMVSTSF 150
+S ++ P+ +S F + ST + P+ T +
Sbjct: 405 ESLYKQTFTTYPNEKLKKS-FACYLSTSTGPVAGTLY 512
>TC227445 homologue to UP|O22639 (O22639) Endoplasmic reticulum HSC70-cognate
binding protein precursor, complete
Length = 2254
Score = 33.1 bits (74), Expect = 0.23
Identities = 27/89 (30%), Positives = 42/89 (46%)
Frame = +2
Query: 35 EFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQLYRQFDSVWKEAEAAAKKVSHNVKEKI 94
E + V+E +E AEE K +KE + + T +Y + V + + A K+ + KEKI
Sbjct: 1739 EIDRMVREAEEFAEEDKKVKERIDARNSLET-YVYNMKNQV-SDKDKLADKLESDEKEKI 1912
Query: 95 SAATDFSTKQNADAKQGSQKSPEEEKNEE 123
A + D Q +K EEK +E
Sbjct: 1913 ETAVK-EALEWLDDNQSVEKEDYEEKLKE 1996
>TC218151 homologue to UP|O22639 (O22639) Endoplasmic reticulum HSC70-cognate
binding protein precursor, partial (35%)
Length = 922
Score = 33.1 bits (74), Expect = 0.23
Identities = 27/89 (30%), Positives = 42/89 (46%)
Frame = +1
Query: 35 EFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQLYRQFDSVWKEAEAAAKKVSHNVKEKI 94
E + V+E +E AEE K +KE + + T +Y + V + + A K+ + KEKI
Sbjct: 421 EIDRMVREAEEFAEEDKKVKERIDARNSLET-YVYNMKNQV-SDKDKLADKLESDEKEKI 594
Query: 95 SAATDFSTKQNADAKQGSQKSPEEEKNEE 123
A + D Q +K EEK +E
Sbjct: 595 ETAVK-EALEWLDDNQSVEKEDYEEKLKE 678
>TC212083 similar to UP|Q71BZ1 (Q71BZ1) Type-B response regulator, partial
(35%)
Length = 708
Score = 32.7 bits (73), Expect = 0.30
Identities = 17/65 (26%), Positives = 32/65 (49%)
Frame = +1
Query: 59 EKTKQTTEQLYRQFDSVWKEAEAAAKKVSHNVKEKISAATDFSTKQNADAKQGSQKSPEE 118
E K + + R+ + WK+AE + + + K S D+S+ N + + S+K +E
Sbjct: 190 EALKNIWQHVVRKRKNEWKDAEQSGSAEEGDRQPKASDEADYSSSANEGSWRNSKKRRDE 369
Query: 119 EKNEE 123
E+ E
Sbjct: 370 EEEAE 384
>TC216791 similar to UP|O23144 (O23144) Proton pump interactor
(AT4g27500/F27G19_100), partial (25%)
Length = 1385
Score = 31.6 bits (70), Expect = 0.67
Identities = 20/77 (25%), Positives = 31/77 (39%)
Frame = +3
Query: 21 EFSKKVKDETVKNPEFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQLYRQFDSVWKEAE 80
EF K+ K PE + + ++ EE+ K+ L+ K K + + KEAE
Sbjct: 579 EFKNPQKEAPAKEPEIDPAKLKEMKREEEIAKAKQALERKKKLAEKAAAKAAIRAQKEAE 758
Query: 81 AAAKKVSHNVKEKISAA 97
K K+K A
Sbjct: 759 KKLKDREKKAKKKSGTA 809
>TC215079 similar to UP|Q6H730 (Q6H730) Ribosomal protein L12-like protein,
partial (9%)
Length = 711
Score = 31.6 bits (70), Expect = 0.67
Identities = 28/86 (32%), Positives = 43/86 (49%), Gaps = 12/86 (13%)
Frame = +2
Query: 13 RRGYSVFNEFSKKVK---------DETVKNPEFQKSVKELKEKAEELKGIKEGLK---EK 60
R V NE KK+K + ++ +F++++ LKE+AE+L+ K +K EK
Sbjct: 176 REALRVANEERKKLKAQAAQLRAQERDIERQQFRETL--LKERAEKLENWKMKVKMHEEK 349
Query: 61 TKQTTEQLYRQFDSVWKEAEAAAKKV 86
+ E L+RQ S W E KKV
Sbjct: 350 KAEKKEFLHRQ-SSAWIEEGNLEKKV 424
>TC203366 similar to UP|Q93Z89 (Q93Z89) Matrix metalloproteinase MMP2,
partial (18%)
Length = 576
Score = 31.6 bits (70), Expect = 0.67
Identities = 18/60 (30%), Positives = 33/60 (55%)
Frame = +1
Query: 11 VDRRGYSVFNEFSKKVKDETVKNPEFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQLYR 70
+DR+ ++F KVKD T+KN + K+L+E E+ + + +++KT +YR
Sbjct: 106 IDRKTITMF---WGKVKDATIKNFPSRDERKKLREYYEKFREFMDRVEQKTTPPPPPMYR 276
>BG239333
Length = 376
Score = 30.8 bits (68), Expect = 1.2
Identities = 17/46 (36%), Positives = 28/46 (59%)
Frame = +2
Query: 23 SKKVKDETVKNPEFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQL 68
+K++K+ N E Q VKELK + EL+ K LKE+ ++ +Q+
Sbjct: 149 AKRLKE---MNDELQAKVKELKGEKNELRDEKNRLKEEKEKLEQQV 277
>TC213749
Length = 669
Score = 30.4 bits (67), Expect = 1.5
Identities = 25/94 (26%), Positives = 43/94 (45%), Gaps = 16/94 (17%)
Frame = +1
Query: 19 FNEFSKKVKDET------VKNPEFQKSVKELKEKAEELKGIKEGLKEKTKQTTE------ 66
F E + K+++E + NP+ Q S +ELKE ++ ++ L++K E
Sbjct: 121 FKESNGKLENEIRNQKLIISNPDAQHSEEELKEARNKVLALEVELEKKNSNCKELEAKCI 300
Query: 67 QLYRQFDSVWKEAE----AAAKKVSHNVKEKISA 96
+L Q +S+ KE K HNV +S+
Sbjct: 301 ELQFQLESMSKECSNHDIIEKDKPLHNVSSSLSS 402
>TC231046
Length = 959
Score = 30.0 bits (66), Expect = 2.0
Identities = 23/84 (27%), Positives = 38/84 (44%), Gaps = 9/84 (10%)
Frame = +1
Query: 25 KVKDETVKNPEFQKSVKELKEKAEELKG-IKEGLKEKTKQTTEQLY--------RQFDSV 75
K+K+E ++ E K E ++ A E K I E + + E+ + F+ V
Sbjct: 361 KLKEEPIEPSEEGKPSGESEKNASETKTLIVESINSEVAPVPEERVIEEIPTAAKSFEKV 540
Query: 76 WKEAEAAAKKVSHNVKEKISAATD 99
KEA ++ ++ SHN E A D
Sbjct: 541 TKEAASSTEEKSHNGVENAPKAVD 612
>BU544588
Length = 414
Score = 29.6 bits (65), Expect = 2.6
Identities = 24/89 (26%), Positives = 40/89 (43%)
Frame = -2
Query: 35 EFQKSVKELKEKAEELKGIKEGLKEKTKQTTEQLYRQFDSVWKEAEAAAKKVSHNVKEKI 94
E ++ V+E +E AEE K +KE + + + E + + + K+ + KEKI
Sbjct: 404 EIERMVREAEEFAEEDKKVKE--RXXARNSLETYVYNMKNQIGDKDKLXXKLXXDEKEKI 231
Query: 95 SAATDFSTKQNADAKQGSQKSPEEEKNEE 123
A + D Q +K EEK +E
Sbjct: 230 ENAVK-EALEWLDDNQSVEKEEYEEKLKE 147
>BI974104
Length = 421
Score = 29.3 bits (64), Expect = 3.3
Identities = 14/39 (35%), Positives = 24/39 (60%)
Frame = +2
Query: 11 VDRRGYSVFNEFSKKVKDETVKNPEFQKSVKELKEKAEE 49
VD +++F F+ ++D +K+PE S++ELK EE
Sbjct: 185 VDNSNFNIFLTFTNHIRD--IKDPEHPYSLEELKVITEE 295
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.321 0.136 0.390
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,940,126
Number of Sequences: 63676
Number of extensions: 150173
Number of successful extensions: 787
Number of sequences better than 10.0: 47
Number of HSP's better than 10.0 without gapping: 775
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 784
length of query: 409
length of database: 12,639,632
effective HSP length: 100
effective length of query: 309
effective length of database: 6,272,032
effective search space: 1938057888
effective search space used: 1938057888
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC141322.10


