

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141115.10 + phase: 0 /pseudo
(173 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
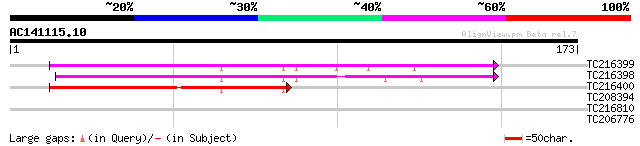
Score E
Sequences producing significant alignments: (bits) Value
TC216399 homologue to UP|Q700A9 (Q700A9) C2 domain-containing pr... 117 2e-27
TC216398 homologue to UP|Q6NPD6 (Q6NPD6) At2g22125, partial (60%) 110 3e-25
TC216400 homologue to UP|Q6NPD6 (Q6NPD6) At2g22125, partial (69%) 79 1e-15
TC208394 similar to UP|O81226 (O81226) Glutamine cyclotransferas... 28 2.2
TC216810 similar to PIR|T51557|T51557 Exportin1 (XPO1) protein -... 27 3.7
TC206776 similar to UP|Q7XXP9 (Q7XXP9) RNA helicase (Fragment), ... 27 4.8
>TC216399 homologue to UP|Q700A9 (Q700A9) C2 domain-containing protein
(Fragment), partial (92%)
Length = 1011
Score = 117 bits (294), Expect = 2e-27
Identities = 80/178 (44%), Positives = 99/178 (54%), Gaps = 41/178 (23%)
Frame = +1
Query: 13 MDALFLLIQGWSACPVEVSRDQSNAAAYAIPLLQNLIQFGPVLFFEKAEFI-------LV 65
+DALFLL Q WSACP EVSR QS AAA AIPLLQ LIQ GP F EKAEF+ LV
Sbjct: 148 LDALFLLRQAWSACPAEVSRAQSIAAADAIPLLQYLIQSGPPRFQEKAEFLLQCLPGTLV 327
Query: 66 MIVKRGNNMRQCVGNQG---KITL-------------EANQEWDERFTY---------MV 100
+I+KRGNNM+Q VGN K+TL N EW E F++ +
Sbjct: 328 VIIKRGNNMKQSVGNPSVYCKLTLGNTPPRQTQVVSTGPNPEWGESFSWTFESPPKGQKL 507
Query: 101 L*ECSSRT--------EASYLLQKQA*SGK-ADEHTLLPTSKSGQPRNLEVELKWSNK 149
C +++ + + + + G A E+ LLP SKSG PRNLE+E +WSNK
Sbjct: 508 HISCKNKSKVGKSKFGKVTIQIDRVVMLGSVAGEYALLPQSKSGPPRNLEIEFQWSNK 681
>TC216398 homologue to UP|Q6NPD6 (Q6NPD6) At2g22125, partial (60%)
Length = 859
Score = 110 bits (276), Expect = 3e-25
Identities = 78/178 (43%), Positives = 95/178 (52%), Gaps = 43/178 (24%)
Frame = +1
Query: 15 ALFLLIQGWSACPVEVSRDQSNAAAYAIPLLQNLIQFGPVLFFEKAEFI-------LVMI 67
ALFL WSACP EVSR QS AAA AIPLLQ LIQ GP F EKAEF+ LV+I
Sbjct: 28 ALFLXXXAWSACPAEVSRAQSIAAADAIPLLQYLIQXGPPRFQEKAEFLLQCLPGTLVVI 207
Query: 68 VKRGNNMRQCVGNQG---KITL-------------EANQEWDERFTYMVL*ECSSRTEAS 111
+K GNNM+Q VGN K+TL N EWDE FT+ E + +
Sbjct: 208 IKCGNNMKQSVGNPSVFCKLTLGNTPPRQTKVVSTGPNPEWDESFTWSF--ESPPKGQKL 381
Query: 112 YL-LQKQA*SGKAD-------------------EHTLLPTSKSGQPRNLEVELKWSNK 149
++ + ++ GK+ E+TLLP SKSG RNLE+E +WSNK
Sbjct: 382 HISCKNKSKMGKSSFGKVTIQIDRVVMLGAVSGEYTLLPESKSGPSRNLEIEFQWSNK 555
>TC216400 homologue to UP|Q6NPD6 (Q6NPD6) At2g22125, partial (69%)
Length = 857
Score = 78.6 bits (192), Expect = 1e-15
Identities = 49/84 (58%), Positives = 56/84 (66%), Gaps = 10/84 (11%)
Frame = +2
Query: 13 MDALFLLIQGWSACPVEVSRDQSNAAAYAIPLLQNLIQFGPVLFFEKAEFI-------LV 65
+DALFLL Q WSACP EVSR QS AAA AIPLLQ + P F EKAEF+ LV
Sbjct: 599 LDALFLLRQAWSACPAEVSRAQSIAAADAIPLLQT*SSW-PTSFHEKAEFLLQCLPGTLV 775
Query: 66 MIVKRGNNMRQCVGNQG---KITL 86
+I+K GNNM+Q VGN K+TL
Sbjct: 776 VIIKCGNNMKQSVGNPSVFCKLTL 847
>TC208394 similar to UP|O81226 (O81226) Glutamine cyclotransferase precursor
, partial (30%)
Length = 816
Score = 28.1 bits (61), Expect = 2.2
Identities = 13/35 (37%), Positives = 20/35 (57%)
Frame = -2
Query: 138 RNLEVELKWSNKPSYAYSIKLSIPSFKFDYLFILV 172
RNL+ W S+ Y + L+IPS+ F + I+V
Sbjct: 269 RNLDFIQLWPQFSSHKYPLLLTIPSYPFQTINIIV 165
>TC216810 similar to PIR|T51557|T51557 Exportin1 (XPO1) protein - Arabidopsis
thaliana (fragment) {Arabidopsis thaliana;} , partial
(22%)
Length = 1237
Score = 27.3 bits (59), Expect = 3.7
Identities = 13/35 (37%), Positives = 18/35 (51%)
Frame = -2
Query: 103 ECSSRTEASYLLQKQA*SGKADEHTLLPTSKSGQP 137
+C SR S + ++ SGK D TL P + S P
Sbjct: 291 KCQSRRRISLVQLSKSMSGKTDYRTLSPETSSASP 187
>TC206776 similar to UP|Q7XXP9 (Q7XXP9) RNA helicase (Fragment), partial
(29%)
Length = 768
Score = 26.9 bits (58), Expect = 4.8
Identities = 9/25 (36%), Positives = 14/25 (56%)
Frame = -2
Query: 142 VELKWSNKPSYAYSIKLSIPSFKFD 166
+ + W NK SY Y I + + KF+
Sbjct: 764 IHINWYNKHSYGYRINVEVGEIKFN 690
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.328 0.139 0.431
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,972,686
Number of Sequences: 63676
Number of extensions: 96439
Number of successful extensions: 582
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 573
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 577
length of query: 173
length of database: 12,639,632
effective HSP length: 91
effective length of query: 82
effective length of database: 6,845,116
effective search space: 561299512
effective search space used: 561299512
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 55 (25.8 bits)
Medicago: description of AC141115.10


