

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141114.8 - phase: 0
(239 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
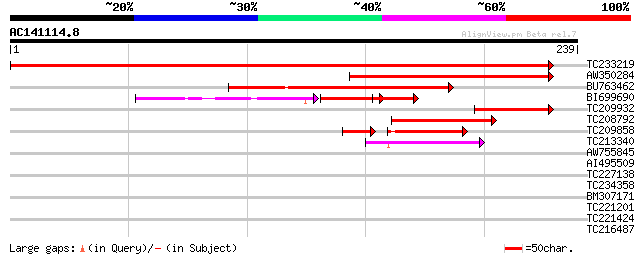
Score E
Sequences producing significant alignments: (bits) Value
TC233219 weakly similar to UP|Q8LKI9 (Q8LKI9) NBS/LRR resistance... 363 e-101
AW350284 similar to GP|22946224|gb| CG31868-PA {Drosophila melan... 157 3e-39
BU763462 weakly similar to GP|9757923|dbj| laccase (diphenol oxi... 85 3e-17
BI699690 50 3e-13
TC209932 53 1e-07
TC208792 homologue to UP|Q9S834 (Q9S834) ATP-dependent Clp prote... 48 5e-06
TC209858 39 3e-05
TC213340 45 4e-05
AW755845 39 0.002
AI495509 37 0.008
TC227138 similar to GB|AAO44062.1|28466907|BT004796 At1g51160 {A... 32 0.010
TC234358 36 0.014
BM307171 32 0.20
TC221201 29 2.2
TC221424 28 4.8
TC216487 homologue to UP|STAD_SOYBN (Q42807) Acyl-[acyl-carrier-... 27 6.3
>TC233219 weakly similar to UP|Q8LKI9 (Q8LKI9) NBS/LRR resistance
protein-like protein (Fragment), partial (74%)
Length = 781
Score = 363 bits (932), Expect = e-101
Identities = 174/229 (75%), Positives = 199/229 (85%)
Frame = +3
Query: 1 MYEGVSTSVRTQDGTTEVFPITIGLHQGSTLSPYLFTLVLDVLTEHIQELAPRCMLFADD 60
MY+ VSTSVRTQ G ++ FPITIGLHQGSTLSPYLFTL+LDVLTE IQE+ RCMLFADD
Sbjct: 63 MYDRVSTSVRTQGGESDDFPITIGLHQGSTLSPYLFTLILDVLTEQIQEIVSRCMLFADD 242
Query: 61 VVLVGESREEVNGRLETWRQALEAYGFRLSRSKTEYMEWNFSGRRSRSTLEVKVGDHIIP 120
+VL+GESRE++N RLETWR+ALE +GFRLSRSK+EYME F+ RR S EVK+GDHIIP
Sbjct: 243 IVLLGESREKLNERLETWRRALETHGFRLSRSKSEYMECKFNKRRRVSNSEVKIGDHIIP 422
Query: 121 QVTRFKYLGSFVQNDGEIEADVSHRIQAGWLKWRRASGVLCDKKVPLKLKGKFYRTAIRP 180
QVTRFKYLGS +Q+DGEIE DV+HRIQA W+KWR+ASGVLCD KVP+KLKGKFYRTA+RP
Sbjct: 423 QVTRFKYLGSVIQDDGEIEGDVNHRIQA*WMKWRKASGVLCDAKVPIKLKGKFYRTAVRP 602
Query: 181 ALLYGTECWAVKSQHENQVSVTEMRMLRWMSGKTRQDRIRNDTIREGRG 229
A+LYGTECWAVKSQHEN+V V RMLRWM GKTRQD+IRN+ E G
Sbjct: 603 AILYGTECWAVKSQHENKVGVAXXRMLRWMCGKTRQDKIRNEAXXERVG 749
>AW350284 similar to GP|22946224|gb| CG31868-PA {Drosophila melanogaster},
partial (1%)
Length = 767
Score = 157 bits (398), Expect = 3e-39
Identities = 70/86 (81%), Positives = 78/86 (90%)
Frame = -1
Query: 144 HRIQAGWLKWRRASGVLCDKKVPLKLKGKFYRTAIRPALLYGTECWAVKSQHENQVSVTE 203
HRIQAGW+KWR+ SGVLCD KVP+KLKGKFYRTA+RPA+LYGTECWAVKSQHEN+V V E
Sbjct: 590 HRIQAGWMKWRKTSGVLCDAKVPIKLKGKFYRTAVRPAILYGTECWAVKSQHENKVGVAE 411
Query: 204 MRMLRWMSGKTRQDRIRNDTIREGRG 229
MRMLRWM GKTRQD+IRN+ IRE G
Sbjct: 410 MRMLRWMCGKTRQDKIRNEAIRERVG 333
>BU763462 weakly similar to GP|9757923|dbj| laccase (diphenol oxidase)
{Arabidopsis thaliana}, partial (4%)
Length = 421
Score = 85.1 bits (209), Expect = 3e-17
Identities = 45/95 (47%), Positives = 60/95 (62%)
Frame = +1
Query: 93 KTEYMEWNFSGRRSRSTLEVKVGDHIIPQVTRFKYLGSFVQNDGEIEADVSHRIQAGWLK 152
KT+YM NFS + + LEVK+GD IP V++FKY +QN I D +H++Q G LK
Sbjct: 1 KTKYMHSNFSKK*EENELEVKMGD-AIP*VSKFKYFEPILQNSW*INEDGTHKMQLGCLK 177
Query: 153 WRRASGVLCDKKVPLKLKGKFYRTAIRPALLYGTE 187
+AS + CD KVP +K +FY T I P +LYG E
Sbjct: 178 EAKASRINCDHKVPTNIKAQFYCTVIHPNILYGNE 282
>BI699690
Length = 407
Score = 50.4 bits (119), Expect(3) = 3e-13
Identities = 33/81 (40%), Positives = 48/81 (58%), Gaps = 4/81 (4%)
Frame = -1
Query: 54 CMLFADDVVLVGESREEVNGRLETWRQALEAYGFRLSRSKTEYMEWNFSGRRSRSTLEVK 113
CML ADD+ + ES G + W+Q+ F LSRSKT+YM +FS + + + +
Sbjct: 347 CMLSADDIFFIEESHNMRFGH-KLWKQS-----FHLSRSKTKYMYCSFS--KKQEEYK*E 192
Query: 114 VGDHIIPQVT----RFKYLGS 130
G+ +IPQV+ +FKYLGS
Sbjct: 191 DGEDVIPQVSKFKFKFKYLGS 129
Score = 32.7 bits (73), Expect(3) = 3e-13
Identities = 13/19 (68%), Positives = 16/19 (83%)
Frame = -3
Query: 154 RRASGVLCDKKVPLKLKGK 172
+R GV+CD KVP+KLKGK
Sbjct: 57 KRQLGVICDHKVPIKLKGK 1
Score = 28.1 bits (61), Expect(3) = 3e-13
Identities = 13/27 (48%), Positives = 19/27 (70%)
Frame = -2
Query: 132 VQNDGEIEADVSHRIQAGWLKWRRASG 158
++N G++ DVSH IQA LK ++A G
Sbjct: 124 MKNYGKMNEDVSHEIQAKRLKCKKAIG 44
>TC209932
Length = 829
Score = 52.8 bits (125), Expect = 1e-07
Identities = 24/33 (72%), Positives = 27/33 (81%)
Frame = +2
Query: 197 NQVSVTEMRMLRWMSGKTRQDRIRNDTIREGRG 229
N+V V EMRMLRWM GKTRQD+IRN+ IRE G
Sbjct: 224 NKVGVAEMRMLRWMCGKTRQDKIRNEAIRERVG 322
>TC208792 homologue to UP|Q9S834 (Q9S834) ATP-dependent Clp protease subunit
ClpP (NClpP1) (ATP-dependent Clp protease proteolytic
subunit ClpP5) , partial (14%)
Length = 739
Score = 47.8 bits (112), Expect = 5e-06
Identities = 23/44 (52%), Positives = 30/44 (67%)
Frame = -2
Query: 162 DKKVPLKLKGKFYRTAIRPALLYGTECWAVKSQHENQVSVTEMR 205
+KKVPLKLKGK T+IR ++YGT+ W VK Q + +V E R
Sbjct: 597 NKKVPLKLKGKLQDTSIRSMMVYGTKEWVVKGQQ*RKHNVAETR 466
>TC209858
Length = 1340
Score = 38.5 bits (88), Expect(2) = 3e-05
Identities = 20/34 (58%), Positives = 24/34 (69%)
Frame = -3
Query: 160 LCDKKVPLKLKGKFYRTAIRPALLYGTECWAVKS 193
LC + VPLK KGKFY IR A+ GT+C A+KS
Sbjct: 285 LC-QNVPLKTKGKFYHPNIRHAM*CGTKCRAIKS 187
Score = 27.7 bits (60), Expect = 4.8
Identities = 27/90 (30%), Positives = 41/90 (45%), Gaps = 2/90 (2%)
Frame = -1
Query: 123 TRFKYLGSFVQNDGEIEADVSHRIQAGWLKWRRASGVLCDKKVPLKLKGKFYRTAIRPAL 182
++ YLGS +Q++GEI R + + + LK K F T +
Sbjct: 386 SQLNYLGSIMQDNGEI----CGRKP*DSCRKNEMENFIYARMYHLKPKENF--TIQI*DM 225
Query: 183 LYGTECWAV--KSQHENQVSVTEMRMLRWM 210
E AV K+ +NQ++V EMR+L WM
Sbjct: 224 QCSVELNAVP*KANKKNQLNVVEMRLLHWM 135
Score = 25.8 bits (55), Expect(2) = 3e-05
Identities = 9/14 (64%), Positives = 12/14 (85%)
Frame = -2
Query: 141 DVSHRIQAGWLKWR 154
DV+HRI AG +KW+
Sbjct: 334 DVNHRIHAGRMKWK 293
>TC213340
Length = 501
Score = 44.7 bits (104), Expect = 4e-05
Identities = 23/53 (43%), Positives = 31/53 (58%), Gaps = 3/53 (5%)
Frame = -1
Query: 151 LKWRRASG---VLCDKKVPLKLKGKFYRTAIRPALLYGTECWAVKSQHENQVS 200
L W+++ G V+ KV LK +FY T I P +LY +ECW K HE +VS
Sbjct: 429 LMWKKSIGGLFVIEICKVLTNLKREFYCTVIEPTILYSSECWGSKG*HEKKVS 271
>AW755845
Length = 152
Score = 38.9 bits (89), Expect = 0.002
Identities = 15/30 (50%), Positives = 22/30 (73%)
Frame = -2
Query: 122 VTRFKYLGSFVQNDGEIEADVSHRIQAGWL 151
VT F+YLGS +Q GE + DV+ +I +GW+
Sbjct: 94 VT*FRYLGSIIQRKGETKGDVNQKIHSGWM 5
>AI495509
Length = 394
Score = 37.0 bits (84), Expect = 0.008
Identities = 13/27 (48%), Positives = 22/27 (81%)
Frame = +2
Query: 178 IRPALLYGTECWAVKSQHENQVSVTEM 204
I+ +L+GTECW VKSQ +N+++V ++
Sbjct: 314 IQXVMLFGTECWTVKSQQDNKLNVADI 394
>TC227138 similar to GB|AAO44062.1|28466907|BT004796 At1g51160 {Arabidopsis
thaliana;} , partial (97%)
Length = 1259
Score = 32.0 bits (71), Expect(2) = 0.010
Identities = 18/48 (37%), Positives = 28/48 (57%)
Frame = -3
Query: 117 HIIPQVTRFKYLGSFVQNDGEIEADVSHRIQAGWLKWRRASGVLCDKK 164
H+I +VTRF +L + EI+ +++ RIQ W + V+CDKK
Sbjct: 1209 HVI-RVTRFTHLQFIIYIGREIQGNINCRIQIKW----SITSVICDKK 1081
Score = 23.5 bits (49), Expect(2) = 0.010
Identities = 9/16 (56%), Positives = 11/16 (68%)
Frame = -2
Query: 104 RRSRSTLEVKVGDHII 119
RR EVK+GDHI+
Sbjct: 1258 RRINLXFEVKIGDHIL 1211
>TC234358
Length = 432
Score = 36.2 bits (82), Expect = 0.014
Identities = 25/69 (36%), Positives = 38/69 (54%)
Frame = -2
Query: 57 FADDVVLVGESREEVNGRLETWRQALEAYGFRLSRSKTEYMEWNFSGRRSRSTLEVKVGD 116
F DDVVL+ +SRE V + + W + + L RSKT Y+ + L+VK+
Sbjct: 317 FLDDVVLIEDSREVVYFKHKLWIYNFDTRSYCLRRSKTVYI---LQFQ*EEYKLKVKI-R 150
Query: 117 HIIPQVTRF 125
+I+ QV+RF
Sbjct: 149 NILSQVSRF 123
>BM307171
Length = 422
Score = 32.3 bits (72), Expect = 0.20
Identities = 12/24 (50%), Positives = 19/24 (79%)
Frame = -1
Query: 122 VTRFKYLGSFVQNDGEIEADVSHR 145
VT+F+YLGS +Q +GE + DV+ +
Sbjct: 299 VTKFRYLGSIIQRNGETKGDVNSK 228
>TC221201
Length = 880
Score = 28.9 bits (63), Expect = 2.2
Identities = 17/45 (37%), Positives = 23/45 (50%)
Frame = +3
Query: 87 FRLSRSKTEYMEWNFSGRRSRSTLEVKVGDHIIPQVTRFKYLGSF 131
F L SKT Y + F + S LEV + + II +F+YL F
Sbjct: 396 FCLGWSKTLYT*YKFQKTQLISVLEVPIDESIISIAVQFRYLQYF 530
>TC221424
Length = 918
Score = 27.7 bits (60), Expect = 4.8
Identities = 11/19 (57%), Positives = 15/19 (78%)
Frame = +2
Query: 153 WRRASGVLCDKKVPLKLKG 171
WR+AS V+C+ KV +LKG
Sbjct: 17 WRKASRVICNPKVLTELKG 73
>TC216487 homologue to UP|STAD_SOYBN (Q42807) Acyl-[acyl-carrier-protein]
desaturase, chloroplast precursor (Stearoyl-ACP
desaturase) , complete
Length = 1588
Score = 27.3 bits (59), Expect = 6.3
Identities = 29/130 (22%), Positives = 47/130 (35%), Gaps = 26/130 (20%)
Frame = +3
Query: 41 DVLTEHIQELAPRCMLFADD--VVLVGES-------------------REEVNGRLETWR 79
D E ++EL R DD VVLVG+ R+E L +W
Sbjct: 387 DGFEEQVKELRERAKEIPDDYFVVLVGDMITEEALPTYQTMLNTLDGVRDETGASLTSWA 566
Query: 80 QALEAYGFRLSR-----SKTEYMEWNFSGRRSRSTLEVKVGDHIIPQVTRFKYLGSFVQN 134
A+ +R +K Y+ ++ T++ +G + P+ YLG +
Sbjct: 567 IWTRAWTAEENRHGDLLNKYLYLSGRVDMKQIEKTIQYLIGSGMDPRTENSPYLGFIYTS 746
Query: 135 DGEIEADVSH 144
E +SH
Sbjct: 747 FQERATFISH 776
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.321 0.136 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,285,041
Number of Sequences: 63676
Number of extensions: 133465
Number of successful extensions: 618
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 613
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 615
length of query: 239
length of database: 12,639,632
effective HSP length: 94
effective length of query: 145
effective length of database: 6,654,088
effective search space: 964842760
effective search space used: 964842760
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 57 (26.6 bits)
Medicago: description of AC141114.8


