

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141111.6 - phase: 0 /pseudo
(379 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
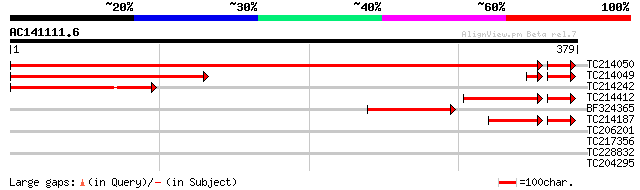
Score E
Sequences producing significant alignments: (bits) Value
TC214050 similar to UP|Q8VYW9 (Q8VYW9) Aminotransferase 1, complete 665 0.0
TC214049 similar to UP|Q8VYW9 (Q8VYW9) Aminotransferase 1, parti... 256 2e-68
TC214242 similar to UP|Q6V1W7 (Q6V1W7) Aminotransferase 1, parti... 186 1e-47
TC214412 homologue to UP|Q6V1W5 (Q6V1W5) Aminotransferase 1, par... 99 6e-26
BF324365 100 1e-21
TC214187 UP|Q93YG3 (Q93YG3) Chlorophyll a/b binding protein type... 75 9e-19
TC206201 similar to UP|Q6ISX8 (Q6ISX8) Pathogen-inducible trypsi... 28 5.2
TC217356 similar to UP|Q84JM5 (Q84JM5) Angustifolia, partial (57%) 28 6.8
TC228832 UP|O81698 (O81698) Proline-rich protein, complete 28 8.9
TC204295 similar to UP|Q9LJE5 (Q9LJE5) Gb|AAD10646.1 (AT3g13460/... 28 8.9
>TC214050 similar to UP|Q8VYW9 (Q8VYW9) Aminotransferase 1, complete
Length = 1676
Score = 665 bits (1716), Expect(2) = 0.0
Identities = 321/356 (90%), Positives = 342/356 (95%)
Frame = +3
Query: 1 MDYVNGPGRNHLFVPGPVNIPDQVIRAMSRNNEDYRSPAIPALTKTLLEDVKKIFKTTTG 60
MDY N PGRNHLFVPGPVNIPDQ+IRAM+RNNEDYRSPAIPA+TKTLLEDVKKIFKT TG
Sbjct: 357 MDYFNAPGRNHLFVPGPVNIPDQIIRAMNRNNEDYRSPAIPAMTKTLLEDVKKIFKTITG 536
Query: 61 TPFLIPTTGTGAWESALTNTLSPGDRIVSFVIGQFSLLWVDQQQRLKFNVDVVESEWGRG 120
PFLIPTTGTGAWESALTNTLSPGDRIVSF+IGQFSLLW+DQQQRLKFNVDVVESEWG+G
Sbjct: 537 IPFLIPTTGTGAWESALTNTLSPGDRIVSFLIGQFSLLWIDQQQRLKFNVDVVESEWGQG 716
Query: 121 ADLDILESKLASDSAHTIKAICIVHNETATGVTNNLAKVRQLLDAYQHPALLLVDGVSSI 180
A LD+LESK+ASD++HTIKAICIVHNETATGVTN+LAKVRQ+LD+YQHPALL+VDGVSSI
Sbjct: 717 AKLDVLESKIASDTSHTIKAICIVHNETATGVTNDLAKVRQILDSYQHPALLIVDGVSSI 896
Query: 181 CALDFRMDEWGVDVAITGSQKALSLPTGIGFVVASPKAIEASKSAKSLRVFFDWSDYLKF 240
CALDFRMDEWGVDVAITGSQKALSLPTGIG VVA PKAIEASK AKSLRVFFDW DYLKF
Sbjct: 897 CALDFRMDEWGVDVAITGSQKALSLPTGIGIVVAGPKAIEASKHAKSLRVFFDWKDYLKF 1076
Query: 241 YKMGTYWPYTPSIQLLYGLRAALDLIFEEGLENIIARHNRLGTATRLAVEAWGLKNCTQE 300
Y++GTYWPYTPSI LLYGLRAALDLIFEEGLEN+IARH+RLG ATRLAVEAWGLKNCTQ+
Sbjct: 1077YQLGTYWPYTPSIHLLYGLRAALDLIFEEGLENVIARHSRLGKATRLAVEAWGLKNCTQK 1256
Query: 301 EEWFSDTVTAVVVPPYIDGAEIVRRAWKRYNLSLGLGLNKVAGKVFRIGHLGNLNE 356
EEW+SDTVTAV+VP YID EIVRRAWKRYNLSLGLGLNKVAGKVFRIGHLGNLNE
Sbjct: 1257EEWYSDTVTAVLVPAYIDSTEIVRRAWKRYNLSLGLGLNKVAGKVFRIGHLGNLNE 1424
Score = 36.6 bits (83), Expect(2) = 0.0
Identities = 17/19 (89%), Positives = 18/19 (94%)
Frame = +1
Query: 360 EVELLLPVHTYRTIFLSSL 378
EVELLLPVHTYRT+FL SL
Sbjct: 1495 EVELLLPVHTYRTLFL*SL 1551
>TC214049 similar to UP|Q8VYW9 (Q8VYW9) Aminotransferase 1, partial (47%)
Length = 866
Score = 256 bits (653), Expect = 2e-68
Identities = 121/133 (90%), Positives = 129/133 (96%)
Frame = +2
Query: 1 MDYVNGPGRNHLFVPGPVNIPDQVIRAMSRNNEDYRSPAIPALTKTLLEDVKKIFKTTTG 60
MDY N PGRNHLFVPGPVNIPDQ+IRAM+RNNEDYRSPAIPA+TKTLLEDVKKIFKTTTG
Sbjct: 107 MDYFNAPGRNHLFVPGPVNIPDQIIRAMNRNNEDYRSPAIPAMTKTLLEDVKKIFKTTTG 286
Query: 61 TPFLIPTTGTGAWESALTNTLSPGDRIVSFVIGQFSLLWVDQQQRLKFNVDVVESEWGRG 120
TPFLIPTTGTGAWESALTNTLSPGDRIVSF+IGQFSLLW+DQQQRLKFNVDVVESEWG G
Sbjct: 287 TPFLIPTTGTGAWESALTNTLSPGDRIVSFLIGQFSLLWIDQQQRLKFNVDVVESEWGHG 466
Query: 121 ADLDILESKLASD 133
A LD+LESK+AS+
Sbjct: 467 AKLDVLESKIASE 505
Score = 36.6 bits (83), Expect(2) = 2e-04
Identities = 17/19 (89%), Positives = 18/19 (94%)
Frame = +2
Query: 360 EVELLLPVHTYRTIFLSSL 378
EVELLLPVHTYRT+FL SL
Sbjct: 602 EVELLLPVHTYRTLFL*SL 658
Score = 25.8 bits (55), Expect(2) = 2e-04
Identities = 10/11 (90%), Positives = 11/11 (99%)
Frame = +1
Query: 346 FRIGHLGNLNE 356
FRIGHLG+LNE
Sbjct: 499 FRIGHLGHLNE 531
>TC214242 similar to UP|Q6V1W7 (Q6V1W7) Aminotransferase 1, partial (29%)
Length = 525
Score = 186 bits (473), Expect = 1e-47
Identities = 91/98 (92%), Positives = 95/98 (96%)
Frame = +2
Query: 1 MDYVNGPGRNHLFVPGPVNIPDQVIRAMSRNNEDYRSPAIPALTKTLLEDVKKIFKTTTG 60
MDY N PGRNHLFVPGPVNIPDQ+IRAM+RNNEDYRSPAIPA+TKTLLEDVKKIFKTTTG
Sbjct: 41 MDYFNAPGRNHLFVPGPVNIPDQIIRAMNRNNEDYRSPAIPAMTKTLLEDVKKIFKTTTG 220
Query: 61 TPFLIPTTGTGAWESALTNTLSPGDRIVSFVIGQFSLL 98
TPFLIPTTGT AWESALTNTLSPGDRIVSF+IGQFSLL
Sbjct: 221 TPFLIPTTGT-AWESALTNTLSPGDRIVSFLIGQFSLL 331
Score = 31.6 bits (70), Expect = 0.62
Identities = 14/19 (73%), Positives = 16/19 (83%)
Frame = +2
Query: 360 EVELLLPVHTYRTIFLSSL 378
+ LLLPVHTYRT+FL SL
Sbjct: 317 QFSLLLPVHTYRTLFL*SL 373
>TC214412 homologue to UP|Q6V1W5 (Q6V1W5) Aminotransferase 1, partial (24%)
Length = 809
Score = 99.0 bits (245), Expect(2) = 6e-26
Identities = 47/53 (88%), Positives = 50/53 (93%)
Frame = -1
Query: 304 FSDTVTAVVVPPYIDGAEIVRRAWKRYNLSLGLGLNKVAGKVFRIGHLGNLNE 356
+SDTVTAV+VP YID EIVRRAWKRYNLSLGLGLNKVAGKVFRIGHLG+LNE
Sbjct: 404 YSDTVTAVLVPAYIDSTEIVRRAWKRYNLSLGLGLNKVAGKVFRIGHLGHLNE 246
Score = 36.6 bits (83), Expect(2) = 6e-26
Identities = 17/19 (89%), Positives = 18/19 (94%)
Frame = -2
Query: 360 EVELLLPVHTYRTIFLSSL 378
EVELLLPVHTYRT+FL SL
Sbjct: 175 EVELLLPVHTYRTLFL*SL 119
>BF324365
Length = 184
Score = 100 bits (249), Expect = 1e-21
Identities = 47/59 (79%), Positives = 50/59 (84%)
Frame = +2
Query: 240 FYKMGTYWPYTPSIQLLYGLRAALDLIFEEGLENIIARHNRLGTATRLAVEAWGLKNCT 298
FY+ GTYW Y PSI LLYGL AALDL+ EEGLEN+IA HNRLG AT LAVEAWGLKNCT
Sbjct: 2 FYRGGTYWSYAPSIHLLYGLIAALDLMLEEGLENVIAWHNRLGEATTLAVEAWGLKNCT 178
>TC214187 UP|Q93YG3 (Q93YG3) Chlorophyll a/b binding protein type II
precursor, partial (28%)
Length = 2003
Score = 74.7 bits (182), Expect(2) = 9e-19
Identities = 35/36 (97%), Positives = 36/36 (99%)
Frame = -3
Query: 321 EIVRRAWKRYNLSLGLGLNKVAGKVFRIGHLGNLNE 356
EIVRRAWKRYNLSLGLGLNKVAGKVFRIGHLG+LNE
Sbjct: 390 EIVRRAWKRYNLSLGLGLNKVAGKVFRIGHLGHLNE 283
Score = 36.6 bits (83), Expect(2) = 9e-19
Identities = 17/19 (89%), Positives = 18/19 (94%)
Frame = -1
Query: 360 EVELLLPVHTYRTIFLSSL 378
EVELLLPVHTYRT+FL SL
Sbjct: 212 EVELLLPVHTYRTLFL*SL 156
>TC206201 similar to UP|Q6ISX8 (Q6ISX8) Pathogen-inducible
trypsin-inhibitor-like protein, partial (65%)
Length = 880
Score = 28.5 bits (62), Expect = 5.2
Identities = 15/30 (50%), Positives = 19/30 (63%)
Frame = -2
Query: 193 DVAITGSQKALSLPTGIGFVVASPKAIEAS 222
DV+ITGS A + P+ GFVV + K AS
Sbjct: 135 DVSITGSGAAPAAPSSSGFVVKAKKRRNAS 46
>TC217356 similar to UP|Q84JM5 (Q84JM5) Angustifolia, partial (57%)
Length = 1672
Score = 28.1 bits (61), Expect = 6.8
Identities = 21/64 (32%), Positives = 31/64 (47%)
Frame = +3
Query: 121 ADLDILESKLASDSAHTIKAICIVHNETATGVTNNLAKVRQLLDAYQHPALLLVDGVSSI 180
+DL L L +++ I A C+ H + + N + QLLD LL +DG +
Sbjct: 159 SDLISLHCALTNETMQIINAECLQHVKPGAFIVNTGSS--QLLDDCAVKQLL-IDGTLAG 329
Query: 181 CALD 184
CALD
Sbjct: 330 CALD 341
>TC228832 UP|O81698 (O81698) Proline-rich protein, complete
Length = 1452
Score = 27.7 bits (60), Expect = 8.9
Identities = 17/50 (34%), Positives = 23/50 (46%), Gaps = 1/50 (2%)
Frame = +1
Query: 6 GPGRNHLFVPGPVNIPDQVIRAMSRNNEDYRSPAI-PALTKTLLEDVKKI 54
GP N + GP P V M N+ R PA+ P + K LL+ V +
Sbjct: 964 GPSDNLAHLSGPPGPPYVVSGQMGAANQPLRPPALTPDMEKALLQQVMSL 1113
>TC204295 similar to UP|Q9LJE5 (Q9LJE5) Gb|AAD10646.1 (AT3g13460/MRP15_10),
partial (34%)
Length = 1745
Score = 27.7 bits (60), Expect = 8.9
Identities = 16/50 (32%), Positives = 25/50 (50%), Gaps = 1/50 (2%)
Frame = -1
Query: 19 NIPDQVIRAMSRNNEDYRSPAIPALTKTLLEDVKKIFKTT-TGTPFLIPT 67
N+P Q++R + N R AIP + +L + KI +T P +PT
Sbjct: 638 NVPQQIVRNILNNVPLKRKAAIPLVLLPILNTLVKINRTNHFSKPHKLPT 489
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.321 0.138 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,583,286
Number of Sequences: 63676
Number of extensions: 214079
Number of successful extensions: 802
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 797
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 801
length of query: 379
length of database: 12,639,632
effective HSP length: 99
effective length of query: 280
effective length of database: 6,335,708
effective search space: 1773998240
effective search space used: 1773998240
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC141111.6


