

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC141111.5 - phase: 0 /pseudo
(1478 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
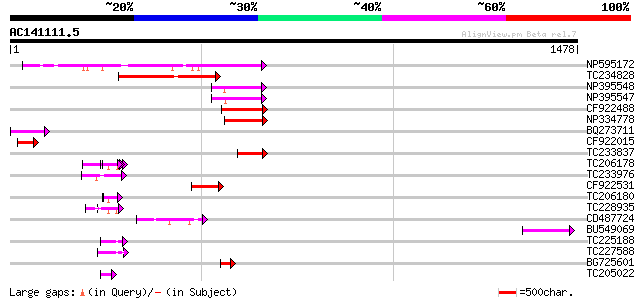
Score E
Sequences producing significant alignments: (bits) Value
NP595172 polyprotein [Glycine max] 227 3e-59
TC234828 218 2e-56
NP395548 reverse transcriptase [Glycine max] 107 4e-23
NP395547 reverse transcriptase [Glycine max] 101 3e-21
CF922488 96 1e-19
NP334778 reverse transcriptase [Glycine max] 92 2e-18
BQ273711 86 9e-17
CF922015 82 2e-15
TC233837 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 61 4e-09
TC206178 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, part... 55 3e-07
TC233976 52 2e-06
CF922531 51 3e-06
TC206180 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, part... 49 2e-05
TC228935 similar to UP|Q9FHC2 (Q9FHC2) Arabidopsis thaliana geno... 48 3e-05
CD487724 48 3e-05
BU549069 47 6e-05
TC225188 similar to UP|Q9FYB7 (Q9FYB7) Splicing factor-like prot... 45 2e-04
TC227588 similar to PIR|T00837|T00837 glycine-rich protein T13L1... 45 2e-04
BG725601 similar to PIR|H86337|H86 protein F5M15.26 [imported] -... 44 7e-04
TC205022 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, pa... 44 7e-04
>NP595172 polyprotein [Glycine max]
Length = 4659
Score = 227 bits (578), Expect = 3e-59
Identities = 192/689 (27%), Positives = 309/689 (43%), Gaps = 53/689 (7%)
Frame = +1
Query: 34 RYDPDGAQTWLKEVERIFRVMQCSEVQKVRFGTHQLAEEADDWWVSLLPNLDQDGVDVTW 93
R+D W+ + E+ F + ++ + L ++ W+ L + +W
Sbjct: 310 RFDGKNVMDWIFKAEQFFDYYATPDADRLIIASVHLDQDVVPWYQML----QKTEPFSSW 477
Query: 94 AVF*REFMRRYFPEDVRRKKEIEFLELKQGNMYVTEYAAKFVELAEFYPHYAAETVEFSK 153
F R + P + F +L Q + V EY +F L +AE +
Sbjct: 478 QAFTRALELDFGPSAYDCPRATLF-KLNQ-SATVNEYYMQFTALVNRVDGLSAEAI--LD 645
Query: 154 CIKFENGLRPDIKRAIEYQQIRVFPDLVNSCRIYEED------TKAHYKVVNE------- 200
C F +GL+ +I R ++ + R V +++EE TK +
Sbjct: 646 C--FVSGLQEEISRDVKAMEPRTLTKAVALAKLFEEKYTSPPKTKTFSNLARNFTSNTSA 819
Query: 201 ----RKTKGQQSRPKPYSAPADKGKQRMVDDRRPK--KKDAPAEIT-------CFNCGEK 247
T + PKP P + R + KK +PAEI C+ C EK
Sbjct: 820 TQKYPPTNQKNDNPKPNLPPLLPTPSTKPFNLRNQNIKKISPAEIQLRREKNLCYFCDEK 999
Query: 248 GHKSNVCPEEIKKCVRCGKKGHIVADCKRNDIVCFNFNEEGHIGSQCKQPKKSPTTGRVF 307
++ CP ++ + D + N +++ H
Sbjct: 1000FSPAHKCPNRQVMLLQLEETDEDQTDEQVMVTEEANMDDDTHH----------------L 1131
Query: 308 ALDGTQTENEDRLIRCTCYINNTPLVAIIDTGATHCFIAFDCVSALGLDLFDMNGEMVVE 367
+L+ + N IR T + + ++D G++ FI L L + V+
Sbjct: 1132SLNAMRGSNGVGTIRFTGQVGGIAVKILVDGGSSDNFIQPRVAQVLKLPVEPAPNLRVLV 1311
Query: 368 TPAKGLVTTSLVCLRCLLSMFGRDFEMDLVCLPLSGMDVILGMNWLEYNHVHI-NY---- 422
+ L +V + L + G++ ++ + L +SG DVILG WL H+ +Y
Sbjct: 1312GNGQILSAEGIV-QQLPLHIQGQEVKVPVYLLQISGADVILGSTWLATLGPHVADYAALT 1488
Query: 423 --FTKSVYFSSVEEESGAEFLSTKQLKQMERDGILMYPLMASLSFENQAVIDKLQ----- 475
F ++ F +++ E +E + QL R L + S E I +Q
Sbjct: 1489LKFFQNDKFITLQGEGNSE-ATQAQLHHFRR-------LQNTKSIEECFAIQLIQKEVPE 1644
Query: 476 -VVCDFPEVFPDEIP--------------DVPLEREVEFSIDLILGTKPVSMAPYRMSAS 520
+ D P E+ +P +RE + +I L G+ PV + PYR +
Sbjct: 1645DTLKDLPTNIDPELAILLHTYAQVFAVPASLPPQREQDHAIPLKQGSGPVKVRPYRYPHT 1824
Query: 521 ELAKLKKQLEDLLEKKFVRPSVSPWGAPVLLVKKKDGSMRLCIDYRQLNKVTIKNRYPLP 580
+ +++K ++++L + ++PS SP+ P+LLVKKKDGS R C DYR LN +T+K+ +P+P
Sbjct: 1825QKDQIEKMIQEMLVQGIIQPSNSPFSLPILLVKKKDGSWRFCTDYRALNAITVKDSFPMP 2004
Query: 581 RIDDLMDQLVGAKVFSKIDLRSGYHQIKVKDEDMQKTAFSTRYGHYEYKVMPFGVTNAPG 640
+D+L+D+L GA+ FSK+DLRSGYHQI V+ ED +KTAF T +GHYE+ VMPFG+TNAP
Sbjct: 2005TVDELLDELHGAQYFSKLDLRSGYHQILVQPEDREKTAFRTHHGHYEWLVMPFGLTNAPA 2184
Query: 641 VFMEYMNRIFHAFLDRFVVVFIDDILIYS 669
F MN+IF L +FV+VF DDILIYS
Sbjct: 2185TFQCLMNKIFQFALRKFVLVFFDDILIYS 2271
>TC234828
Length = 857
Score = 218 bits (555), Expect = 2e-56
Identities = 116/270 (42%), Positives = 166/270 (60%), Gaps = 4/270 (1%)
Frame = +3
Query: 283 NFNEEGHIGSQCKQPKKSPTTGRVFALDGTQTENEDRLIRCTCYINNTPLVAIIDTGATH 342
N G++ + + RVFA+ G++ D LIR C I + L + D+GATH
Sbjct: 66 NSTRSGNVSNNNISGGRPKVPSRVFAMSGSEAAASDDLIRGKCLIADKLLDVLYDSGATH 245
Query: 343 CFIAFDCVSALGLDLFDMNGEMVVETPAKGLVTTSLVCLRCLLSMFGRDFEMDLVCLPLS 402
FI+ CV LGL ++ +MVV TP VTTS VCL+C + + GR F DL+CLPL+
Sbjct: 246 SFISHACVERLGLCATELPYDMVVSTPTSEPVTTSRVCLKCPIIVEGRSFMADLICLPLA 425
Query: 403 GMDVILGMNWLEYNHVHINYFTKSVYFSSVEEESGAEFLSTKQLKQ----MERDGILMYP 458
+DVILGM+WL NH+ ++ K + F G + + ++ LK+ E + + Y
Sbjct: 426 HLDVILGMDWLSTNHIFLDCKEKMLVF-------GGDVVPSEPLKEDAANEETEDVRTYM 584
Query: 459 LMASLSFENQAVIDKLQVVCDFPEVFPDEIPDVPLEREVEFSIDLILGTKPVSMAPYRMS 518
++ S+ E A + + VV +FPEVFPD++ ++P EREVEF ID++ G PVS+APYRMS
Sbjct: 585 VLFSMYVEEDAEVSCIPVVSEFPEVFPDDVCELPPEREVEFIIDVVPGANPVSIAPYRMS 764
Query: 519 ASELAKLKKQLEDLLEKKFVRPSVSPWGAP 548
ELA++K Q++DLL K+FVRPS SPWGAP
Sbjct: 765 PVELAEVKAQVQDLLSKQFVRPSASPWGAP 854
>NP395548 reverse transcriptase [Glycine max]
Length = 762
Score = 107 bits (267), Expect = 4e-23
Identities = 58/163 (35%), Positives = 94/163 (57%), Gaps = 19/163 (11%)
Frame = +1
Query: 525 LKKQLEDLLEKKFVRP-SVSPWGAPVLLVKKKDG------------------SMRLCIDY 565
++K++ LLE + P S S W +PVL+V KK+G S +LCIDY
Sbjct: 1 VRKEVLKLLEVGLIYPISDSAWVSPVLVVSKKEGMTVIRNEKNDLIPTRTVTSWKLCIDY 180
Query: 566 RQLNKVTIKNRYPLPRIDDLMDQLVGAKVFSKIDLRSGYHQIKVKDEDMQKTAFSTRYGH 625
R+LN+ T K+ +PLP +D ++++L G + +D GY+QI V +D +K AF+ +G
Sbjct: 181 RKLNEATRKDHFPLPFMDQMLERLAGHAYYCFLDAYFGYNQIVVDPKDQEKMAFTCPFGV 360
Query: 626 YEYKVMPFGVTNAPGVFMEYMNRIFHAFLDRFVVVFIDDILIY 668
+ Y+ +PFG+ NAP F M IF +++ + VF+DD ++
Sbjct: 361 FAYRRIPFGLCNAPTTFQMCMLAIFADIVEKSIEVFMDDFSVF 489
>NP395547 reverse transcriptase [Glycine max]
Length = 762
Score = 101 bits (251), Expect = 3e-21
Identities = 57/163 (34%), Positives = 88/163 (53%), Gaps = 19/163 (11%)
Frame = +1
Query: 525 LKKQLEDLLEKKFVRP-SVSPWGAPVLLVKKKDGSM------------------RLCIDY 565
++K++ LLE + P S S W +PV +V KK G R+CIDY
Sbjct: 1 VRKEVFKLLEAGLIYPISDSSWVSPVQVVPKKGGMTVVKNDRNELIPTRRVTRWRMCIDY 180
Query: 566 RQLNKVTIKNRYPLPRIDDLMDQLVGAKVFSKIDLRSGYHQIKVKDEDMQKTAFSTRYGH 625
R+LN+ T K+ YPLP +D ++ +L + +D SGY+QI V +D +KTAF+ +
Sbjct: 181 RKLNEATRKDHYPLPFMDQMLKRLARQSFYRFLDGYSGYNQIAVDPQDQEKTAFTCPFSV 360
Query: 626 YEYKVMPFGVTNAPGVFMEYMNRIFHAFLDRFVVVFIDDILIY 668
+ Y+ MPFG+ NA F M IF +++ + VF+DD +
Sbjct: 361 FAYRRMPFGLCNASTTFQRCMMAIFDDMVEKCIEVFMDDFSFF 489
>CF922488
Length = 741
Score = 95.9 bits (237), Expect = 1e-19
Identities = 46/119 (38%), Positives = 74/119 (61%)
Frame = +3
Query: 552 VKKKDGSMRLCIDYRQLNKVTIKNRYPLPRIDDLMDQLVGAKVFSKIDLRSGYHQIKVKD 611
V K+DG + +C+DYR LN + K+++PLP I+ L+D FS +D SGY+QIK+
Sbjct: 3 VLKEDGKV*MCVDYRDLN*ASPKDKFPLPHINVLVDNTTSFSQFSFMDGFSGYNQIKIAP 182
Query: 612 EDMQKTAFSTRYGHYEYKVMPFGVTNAPGVFMEYMNRIFHAFLDRFVVVFIDDILIYSK 670
EDM+KT F T +G + YK M FG+ N + M +F + + + V++DD+++ S+
Sbjct: 183 EDMEKTTFITLWGTFCYKAMSFGLKNVGATYQRAMVALF*DMMHKEIEVYMDDMIVKSR 359
>NP334778 reverse transcriptase [Glycine max]
Length = 431
Score = 92.0 bits (227), Expect = 2e-18
Identities = 43/111 (38%), Positives = 69/111 (61%)
Frame = +3
Query: 560 RLCIDYRQLNKVTIKNRYPLPRIDDLMDQLVGAKVFSKIDLRSGYHQIKVKDEDMQKTAF 619
R+C+DYR LN+ + K+ +PLP ID LM + +FS +D SGY+QIK+ EDM+KT F
Sbjct: 3 RMCVDYRDLNRASPKDNFPLPHIDILMANMASFALFSFMDGFSGYNQIKMAPEDMEKTTF 182
Query: 620 STRYGHYEYKVMPFGVTNAPGVFMEYMNRIFHAFLDRFVVVFIDDILIYSK 670
T +G + YKVM FG+ N + M +F + + + ++D+++ S+
Sbjct: 183 ITLWGTFCYKVMSFGLKNFGATYHRAMVALFQDMMHKEIEAYVDEMIAKSR 335
>BQ273711
Length = 409
Score = 86.3 bits (212), Expect = 9e-17
Identities = 41/104 (39%), Positives = 56/104 (53%)
Frame = -2
Query: 1 AVAQAVGQQPAAGNGEVRMLETFLRNHPPAFKGRYDPDGAQTWLKEVERIFRVMQCSEVQ 60
AV + + Q+ E R L F +NHPP F G YDP+GA+ WL E E+IF M C E
Sbjct: 312 AVLETLVQERDVEPAEYRGLMAFRKNHPPKFSGDYDPEGARLWLAETEKIFEAMGCLEEH 133
Query: 61 KVRFGTHQLAEEADDWWVSLLPNLDQDGVDVTWAVF*REFMRRY 104
KV + T L EA++WW + P+ G + W F +F+ Y
Sbjct: 132 KVLYATFMLQGEAENWWKFVKPSFVAPGGVIPWNAFKDKFLENY 1
>CF922015
Length = 172
Score = 81.6 bits (200), Expect = 2e-15
Identities = 34/54 (62%), Positives = 44/54 (80%)
Frame = -3
Query: 20 LETFLRNHPPAFKGRYDPDGAQTWLKEVERIFRVMQCSEVQKVRFGTHQLAEEA 73
L+ F RN+PP FKG YDP+GA+ WL+E+E+IFRVM+C + QKV F TH LA+EA
Sbjct: 164 LDRFQRNNPPTFKGGYDPEGAEAWLREIEKIFRVMECQDHQKVLFATHMLADEA 3
>TC233837 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (6%)
Length = 402
Score = 60.8 bits (146), Expect = 4e-09
Identities = 28/76 (36%), Positives = 47/76 (61%)
Frame = +2
Query: 595 FSKIDLRSGYHQIKVKDEDMQKTAFSTRYGHYEYKVMPFGVTNAPGVFMEYMNRIFHAFL 654
FS +D SGY+QI + ED++KT F T +G + Y+VM FG+ N + M +FH +
Sbjct: 2 FSFMDGFSGYNQI*MAREDVEKTTFVTLWGTFSYRVMAFGLKNTGATYQRAMVALFHDMM 181
Query: 655 DRFVVVFIDDILIYSK 670
+ + V++DD++ S+
Sbjct: 182 HKEIEVYVDDMIAKSR 229
>TC206178 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, partial (69%)
Length = 1138
Score = 54.7 bits (130), Expect = 3e-07
Identities = 30/118 (25%), Positives = 52/118 (43%), Gaps = 2/118 (1%)
Frame = +2
Query: 190 DTKAHYKVVNERKTKGQQSRPKPYSAP--ADKGKQRMVDDRRPKKKDAPAEITCFNCGEK 247
+ K +K+ ++ +++ + P +D+ R RR ++ + C NC
Sbjct: 185 ERKKKWKMSSDSRSRSRSRSRSPMDRKIRSDRFSYRDAPYRRDSRRGFSRDNLCKNCKRP 364
Query: 248 GHKSNVCPEEIKKCVRCGKKGHIVADCKRNDIVCFNFNEEGHIGSQCKQPKKSPTTGR 305
GH + CP + C CG GHI ++C + C+N E GH+ S C T G+
Sbjct: 365 GHYARECP-NVAICHNCGLPGHIASECTTKSL-CWNCKEPGHMASSCPNEGICHTCGK 532
Score = 51.2 bits (121), Expect = 3e-06
Identities = 23/65 (35%), Positives = 34/65 (51%), Gaps = 6/65 (9%)
Frame = +2
Query: 236 PAEITCFNCGEKGHKSNVC------PEEIKKCVRCGKKGHIVADCKRNDIVCFNFNEEGH 289
P E C CG+ GH++ C P +++ C C K+GHI A+C N+ C N + GH
Sbjct: 500 PNEGICHTCGKAGHRARECSAPPMPPGDLRLCNNCYKQGHIAAEC-TNEKACNNCRKTGH 676
Query: 290 IGSQC 294
+ C
Sbjct: 677 LARDC 691
Score = 48.5 bits (114), Expect = 2e-05
Identities = 23/64 (35%), Positives = 33/64 (50%), Gaps = 6/64 (9%)
Frame = +2
Query: 241 CFNCGEKGHKSNVCPEEIKKCVRCGKKGHIVADCKRNDI------VCFNFNEEGHIGSQC 294
C+NC E GH ++ CP E C CGK GH +C + +C N ++GHI ++C
Sbjct: 458 CWNCKEPGHMASSCPNE-GICHTCGKAGHRARECSAPPMPPGDLRLCNNCYKQGHIAAEC 634
Query: 295 KQPK 298
K
Sbjct: 635 TNEK 646
Score = 48.5 bits (114), Expect = 2e-05
Identities = 23/54 (42%), Positives = 29/54 (53%)
Frame = +2
Query: 241 CFNCGEKGHKSNVCPEEIKKCVRCGKKGHIVADCKRNDIVCFNFNEEGHIGSQC 294
C NC ++GH + C E K C C K GH+ DC ND +C N GH+ QC
Sbjct: 593 CNNCYKQGHIAAECTNE-KACNNCRKTGHLARDCP-NDPICNLCNVSGHVARQC 748
Score = 37.7 bits (86), Expect = 0.038
Identities = 13/37 (35%), Positives = 20/37 (53%)
Frame = +2
Query: 238 EITCFNCGEKGHKSNVCPEEIKKCVRCGKKGHIVADC 274
++ C NC + GH S C + C CG +GH+ +C
Sbjct: 833 DVVCRNCQQLGHMSRDCMGPLMICHNCGGRGHLAYEC 943
Score = 36.6 bits (83), Expect = 0.084
Identities = 21/85 (24%), Positives = 29/85 (33%), Gaps = 25/85 (29%)
Frame = +2
Query: 238 EITCFNCGEKGHKSNVCPEEIKKCVRCGKKGHIVADCKR--------------------- 276
E C NC + GH + CP + C C GH+ C +
Sbjct: 641 EKACNNCRKTGHLARDCPND-PICNLCNVSGHVARQCPKANVLGDRSGGGGGGGGARGGG 817
Query: 277 ----NDIVCFNFNEEGHIGSQCKQP 297
D+VC N + GH+ C P
Sbjct: 818 GGGYRDVVCRNCQQLGHMSRDCMGP 892
Score = 33.5 bits (75), Expect = 0.71
Identities = 19/87 (21%), Positives = 29/87 (32%), Gaps = 25/87 (28%)
Frame = +2
Query: 233 KDAPAEITCFNCGEKGHKSNVCPE-------------------------EIKKCVRCGKK 267
+D P + C C GH + CP+ C C +
Sbjct: 683 RDCPNDPICNLCNVSGHVARQCPKANVLGDRSGGGGGGGGARGGGGGGYRDVVCRNCQQL 862
Query: 268 GHIVADCKRNDIVCFNFNEEGHIGSQC 294
GH+ DC ++C N GH+ +C
Sbjct: 863 GHMSRDCMGPLMICHNCGGRGHLAYEC 943
>TC233976
Length = 763
Score = 52.0 bits (123), Expect = 2e-06
Identities = 35/133 (26%), Positives = 56/133 (41%), Gaps = 16/133 (12%)
Frame = +1
Query: 188 EEDTKAHYKVVNERKTKGQQ----SRPKPYSAPADK----GKQRMV--------DDRRPK 231
+++ A YK N K ++ SR KPYSAP + G QR +
Sbjct: 25 QQEKVAFYKNANASHGKEKKPMTHSRAKPYSAPLEYENHYGGQRTSGGHHLAGGSSQLVN 204
Query: 232 KKDAPAEITCFNCGEKGHKSNVCPEEIKKCVRCGKKGHIVADCKRNDIVCFNFNEEGHIG 291
+ PA G G + +C +CG+ GH +C ++ CFN+ +GH+
Sbjct: 205 RVSQPA-------GRGGSGAPAIVTTPLRCRKCGRLGHNAHECTYREVTCFNYQGKGHLS 363
Query: 292 SQCKQPKKSPTTG 304
+ C P+K +G
Sbjct: 364 TNCPHPRKEKRSG 402
>CF922531
Length = 602
Score = 51.2 bits (121), Expect = 3e-06
Identities = 30/83 (36%), Positives = 50/83 (60%), Gaps = 1/83 (1%)
Frame = -2
Query: 475 QVVCDFPEVFPDEIP-DVPLEREVEFSIDLILGTKPVSMAPYRMSASELAKLKKQLEDLL 533
+++ +F ++FP EIP +P R +E IDL+ + YR + E +++ Q+++LL
Sbjct: 250 ELLHEFGDIFPKEIPLGLPPLRGIEHQIDLVPRASLPNRPTYRTNPQETKEIESQVKELL 71
Query: 534 EKKFVRPSVSPWGAPVLLVKKKD 556
EK +V+ S+S VLLV KKD
Sbjct: 70 EKGWVQESLSLCVVLVLLVPKKD 2
>TC206180 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, partial (42%)
Length = 655
Score = 48.5 bits (114), Expect = 2e-05
Identities = 23/54 (42%), Positives = 29/54 (53%)
Frame = +3
Query: 241 CFNCGEKGHKSNVCPEEIKKCVRCGKKGHIVADCKRNDIVCFNFNEEGHIGSQC 294
C NC ++GH + C E K C C K GH+ DC ND +C N GH+ QC
Sbjct: 78 CNNCYKQGHIAAECTNE-KACNNCRKTGHLARDCP-NDPICNLCNVSGHVARQC 233
Score = 46.2 bits (108), Expect = 1e-04
Identities = 20/57 (35%), Positives = 31/57 (54%), Gaps = 6/57 (10%)
Frame = +3
Query: 244 CGEKGHKSNVC------PEEIKKCVRCGKKGHIVADCKRNDIVCFNFNEEGHIGSQC 294
CG+ GH++ C P +++ C C K+GHI A+C N+ C N + GH+ C
Sbjct: 9 CGKAGHRARECSAPPMPPGDLRLCNNCYKQGHIAAEC-TNEKACNNCRKTGHLARDC 176
Score = 37.7 bits (86), Expect = 0.038
Identities = 13/37 (35%), Positives = 20/37 (53%)
Frame = +3
Query: 238 EITCFNCGEKGHKSNVCPEEIKKCVRCGKKGHIVADC 274
++ C NC + GH S C + C CG +GH+ +C
Sbjct: 309 DVVCRNCQQLGHMSRDCMGPLMICHNCGGRGHLAYEC 419
Score = 37.7 bits (86), Expect = 0.038
Identities = 21/82 (25%), Positives = 29/82 (34%), Gaps = 22/82 (26%)
Frame = +3
Query: 238 EITCFNCGEKGHKSNVCPEEIKKCVRCGKKGHIVADCKR--------------------- 276
E C NC + GH + CP + C C GH+ C +
Sbjct: 126 EKACNNCRKTGHLARDCPND-PICNLCNVSGHVARQCPKANVLGDXXGGGGGARGGGGGG 302
Query: 277 -NDIVCFNFNEEGHIGSQCKQP 297
D+VC N + GH+ C P
Sbjct: 303 YRDVVCRNCQQLGHMSRDCMGP 368
Score = 34.7 bits (78), Expect = 0.32
Identities = 19/84 (22%), Positives = 29/84 (33%), Gaps = 22/84 (26%)
Frame = +3
Query: 233 KDAPAEITCFNCGEKGHKSNVCPE----------------------EIKKCVRCGKKGHI 270
+D P + C C GH + CP+ C C + GH+
Sbjct: 168 RDCPNDPICNLCNVSGHVARQCPKANVLGDXXGGGGGARGGGGGGYRDVVCRNCQQLGHM 347
Query: 271 VADCKRNDIVCFNFNEEGHIGSQC 294
DC ++C N GH+ +C
Sbjct: 348 SRDCMGPLMICHNCGGRGHLAYEC 419
>TC228935 similar to UP|Q9FHC2 (Q9FHC2) Arabidopsis thaliana genomic DNA,
chromosome 5, TAC clone:K24M7, partial (36%)
Length = 1224
Score = 48.1 bits (113), Expect = 3e-05
Identities = 23/80 (28%), Positives = 33/80 (40%), Gaps = 13/80 (16%)
Frame = +1
Query: 230 PKKKDAPAEITCFNCGEKGHKSNVCPE------EIKKCVRCGKKGHIVADCKR------- 276
P+K + C C +GH++ CPE + K C CG+ GH + C
Sbjct: 313 PEKAEWEKNKICLRCRRRGHRAKNCPEVLDGAKDAKYCYNCGENGHALTQCLHPLQEGGT 492
Query: 277 NDIVCFNFNEEGHIGSQCKQ 296
CF N+ GH+ C Q
Sbjct: 493 KFAECFVCNQRGHLSKNCPQ 552
Score = 47.8 bits (112), Expect = 4e-05
Identities = 33/113 (29%), Positives = 53/113 (46%), Gaps = 14/113 (12%)
Frame = +1
Query: 199 NERKTKGQQS---RPKPYSAPADKGKQRMVDDRRPKKKDAPAEITCFNCGEKGHKSNVCP 255
+++K K + S R +P P + + + R P K P E +CF C H + +CP
Sbjct: 154 DKKKKKNKNSAFKRKRPEPKPGSRKRHLL---RVPGMK--PGE-SCFICKAMDHIAKLCP 315
Query: 256 EEI-----KKCVRCGKKGHIVADCK------RNDIVCFNFNEEGHIGSQCKQP 297
E+ K C+RC ++GH +C ++ C+N E GH +QC P
Sbjct: 316 EKAEWEKNKICLRCRRRGHRAKNCPEVLDGAKDAKYCYNCGENGHALTQCLHP 474
>CD487724
Length = 676
Score = 48.1 bits (113), Expect = 3e-05
Identities = 49/207 (23%), Positives = 91/207 (43%), Gaps = 22/207 (10%)
Frame = +3
Query: 331 PLVAIIDTGATHCFIAFDCVSALGLDLFDMNGEMVVETPAKGLVTTSLVCLRCLLSMFGR 390
PLV ++D G+TH F+ VS LGL V+ L T+ +C +S+
Sbjct: 21 PLVYLVDGGSTHNFVQQPLVSQLGLPCRSTPPLRVMVGNGHHLKCTT-ICEAIPISIQNI 197
Query: 391 DFEMDLVCLPLSGMDVILGMNWL--------EYNHVHINYFTK-SVYFSSVEEESGAEFL 441
+F + L LP+ G +++LG+ WL +YN + + +F + + E E+ L
Sbjct: 198 EFLVHLYVLPIVGANIVLGVQWLKTLGPILVDYNSLSMQFFYQHRLVQLKGESEAQLGLL 377
Query: 442 STKQLKQMERDGILMYPLMASLSFEN-------------QAVIDKLQVVCDFPEVFPDEI 488
+ QL+++ + + ++ EN Q ++D+ + P+
Sbjct: 378 NHHQLRRLHQTHEPVTYFHIAILTENTSPTSSPPLPQPIQHLLDQFSALFQ*PQ------ 539
Query: 489 PDVPLEREVEFSIDLILGTKPVSMAPY 515
+P RE + I L+ ++PV+M Y
Sbjct: 540 -GLPPARETDHHIHLLP*SEPVNMRLY 617
>BU549069
Length = 615
Score = 47.0 bits (110), Expect = 6e-05
Identities = 41/134 (30%), Positives = 67/134 (49%)
Frame = -3
Query: 1338 ESHSCDWCWMCFEVKEVDSEIFRSVSDIEKIWNGGVSSGITTASFELARCFPCVATSELC 1397
ESHS DW W E+ + + ++RS + K + G+ + IT +F ++C CV+T +
Sbjct: 613 ESHSKDWGWSSTEIPKTHTSLYRSFPNS*KSXSCGIPNCITPITF*SSQCLSCVSTPYVY 434
Query: 1398 SGSISCDPE**CAS*RQPYGRDFTGED**SKSEDVERQGDTPC*SR*GWSDW*KLDMGA* 1457
SISC * S R+ + T ED K++ +++G++ * G + +G
Sbjct: 433 P*SISCGQIG*RTSKRELDI*NITIEDRG*KNKAPKKEGESIG*GDLGRYIRRRCHVGIR 254
Query: 1458 E*DAGVLSRIVCLR 1471
E DA LS V +R
Sbjct: 253 ESDASSLSIFV*VR 212
>TC225188 similar to UP|Q9FYB7 (Q9FYB7) Splicing factor-like protein, partial
(62%)
Length = 1131
Score = 45.4 bits (106), Expect = 2e-04
Identities = 25/72 (34%), Positives = 35/72 (47%), Gaps = 2/72 (2%)
Frame = +1
Query: 236 PAEITCFNCGEKGHKSNVCP--EEIKKCVRCGKKGHIVADCKRNDIVCFNFNEEGHIGSQ 293
P CFNCG GH + C + KC RCG++GHI +CK + ++ G S+
Sbjct: 349 PGSGRCFNCGIDGHWARDCKAGDWKNKCYRCGERGHIEKNCKNSP---KKLSQRGRRLSR 519
Query: 294 CKQPKKSPTTGR 305
+SP GR
Sbjct: 520 SPVRSRSPRRGR 555
>TC227588 similar to PIR|T00837|T00837 glycine-rich protein T13L16.11 -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(10%)
Length = 1300
Score = 45.4 bits (106), Expect = 2e-04
Identities = 29/86 (33%), Positives = 39/86 (44%), Gaps = 4/86 (4%)
Frame = +1
Query: 228 RRPKKKDAPAEI--TCFNCGEKGHKSNVCP--EEIKKCVRCGKKGHIVADCKRNDIVCFN 283
R P+ D P CFNCGE+GH + C + K C CG GH C + CF
Sbjct: 16 RGPRYFDPPDNSWGACFNCGEEGHAAVNCSAVKRKKPCYVCGCLGHNARQCSKVQ-DCFI 192
Query: 284 FNEEGHIGSQCKQPKKSPTTGRVFAL 309
+ GH C P+K +T + A+
Sbjct: 193 CKKGGHRAKDC--PEKHTSTSKSIAI 264
Score = 41.6 bits (96), Expect = 0.003
Identities = 28/87 (32%), Positives = 36/87 (41%), Gaps = 14/87 (16%)
Frame = +1
Query: 228 RRPKKKDAPAEITCFNCGEKGH-----KSNVCPEEIKKCVRCGKKGHIVADCK--RNDIV 280
R +D EI C+ C GH + P EI C +CG+ GH C R++I
Sbjct: 307 RNDYSQDDLKEIQCYVCKRLGHLCCVNTDDATPGEIS-CYKCGQLGHTGLACSRLRDEIT 483
Query: 281 -------CFNFNEEGHIGSQCKQPKKS 300
CF EEGH +C KS
Sbjct: 484 SGATPSSCFKCGEEGHFARECTSSIKS 564
Score = 40.4 bits (93), Expect = 0.006
Identities = 19/64 (29%), Positives = 31/64 (47%), Gaps = 14/64 (21%)
Frame = +1
Query: 241 CFNCGEKGHKSNVCPEE-------IKKCVRCGKKGHIVADCKRN-------DIVCFNFNE 286
CF C + GH++ CPE+ I C++CG GH + C+ + +I C+
Sbjct: 184 CFICKKGGHRAKDCPEKHTSTSKSIAICLKCGNSGHDIFSCRNDYSQDDLKEIQCYVCKR 363
Query: 287 EGHI 290
GH+
Sbjct: 364 LGHL 375
Score = 38.1 bits (87), Expect = 0.029
Identities = 23/97 (23%), Positives = 40/97 (40%), Gaps = 11/97 (11%)
Frame = +1
Query: 219 KGKQRMVDDRRPKKKDAPAEITCFNCGEKGH-----KSNVCPEEIK--KCVRCGKKGHIV 271
KG R D + + C CG GH +++ +++K +C C + GH+
Sbjct: 199 KGGHRAKDCPEKHTSTSKSIAICLKCGNSGHDIFSCRNDYSQDDLKEIQCYVCKRLGHLC 378
Query: 272 A----DCKRNDIVCFNFNEEGHIGSQCKQPKKSPTTG 304
D +I C+ + GH G C + + T+G
Sbjct: 379 CVNTDDATPGEISCYKCGQLGHTGLACSRLRDEITSG 489
Score = 35.8 bits (81), Expect = 0.14
Identities = 23/71 (32%), Positives = 33/71 (46%), Gaps = 2/71 (2%)
Frame = +1
Query: 240 TCFNCGEKGHKSNVCPEEIKKCVRCGKKGHI--VADCKRNDIVCFNFNEEGHIGSQCKQP 297
+CF CGE+GH + C IK R + H K ND + +G++ K
Sbjct: 502 SCFKCGEEGHFARECTSSIKSGKRNWESSHTKDKRSQKENDYMGNRSASNDMVGARRK-- 675
Query: 298 KKSPTTGRVFA 308
K+SPT R F+
Sbjct: 676 KRSPTEERGFS 708
>BG725601 similar to PIR|H86337|H86 protein F5M15.26 [imported] - Arabidopsis
thaliana, partial (1%)
Length = 285
Score = 43.5 bits (101), Expect = 7e-04
Identities = 20/41 (48%), Positives = 26/41 (62%)
Frame = -3
Query: 549 VLLVKKKDGSMRLCIDYRQLNKVTIKNRYPLPRIDDLMDQL 589
V++VKK +G R+C DY LN K+ YPLP ID + D L
Sbjct: 124 VVMVKKPNGKWRICTDYIDLN*ACPKDAYPLPNIDHMTDGL 2
>TC205022 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, partial (69%)
Length = 1623
Score = 43.5 bits (101), Expect = 7e-04
Identities = 19/44 (43%), Positives = 25/44 (56%), Gaps = 2/44 (4%)
Frame = +2
Query: 236 PAEITCFNCGEKGHKSNVCP--EEIKKCVRCGKKGHIVADCKRN 277
P CFNCG GH + C + KC RCG++GHI +CK +
Sbjct: 299 PGSGRCFNCGLDGHWARDCKAGDWKNKCYRCGERGHIERNCKNS 430
Score = 40.0 bits (92), Expect = 0.008
Identities = 19/48 (39%), Positives = 22/48 (45%), Gaps = 2/48 (4%)
Frame = +2
Query: 260 KCVRCGKKGHIVADCKRND--IVCFNFNEEGHIGSQCKQPKKSPTTGR 305
+C CG GH DCK D C+ E GHI CK K +T R
Sbjct: 311 RCFNCGLDGHWARDCKAGDWKNKCYRCGERGHIERNCKNSPKKLSTRR 454
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.349 0.155 0.537
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 63,969,210
Number of Sequences: 63676
Number of extensions: 892984
Number of successful extensions: 8134
Number of sequences better than 10.0: 59
Number of HSP's better than 10.0 without gapping: 5894
Number of HSP's successfully gapped in prelim test: 182
Number of HSP's that attempted gapping in prelim test: 1916
Number of HSP's gapped (non-prelim): 6429
length of query: 1478
length of database: 12,639,632
effective HSP length: 109
effective length of query: 1369
effective length of database: 5,698,948
effective search space: 7801859812
effective search space used: 7801859812
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.8 bits)
S2: 65 (29.6 bits)
Medicago: description of AC141111.5


