

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
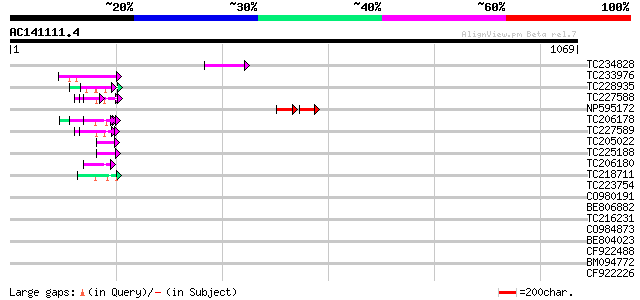
Query= AC141111.4 - phase: 0 /pseudo
(1069 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC234828 60 4e-09
TC233976 51 2e-06
TC228935 similar to UP|Q9FHC2 (Q9FHC2) Arabidopsis thaliana geno... 49 2e-05
TC227588 similar to PIR|T00837|T00837 glycine-rich protein T13L1... 48 3e-05
NP595172 polyprotein [Glycine max] 37 3e-05
TC206178 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, part... 47 6e-05
TC227589 46 8e-05
TC205022 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, pa... 45 1e-04
TC225188 similar to UP|Q9FYB7 (Q9FYB7) Splicing factor-like prot... 45 2e-04
TC206180 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, part... 44 4e-04
TC218711 similar to UP|Q90Z60 (Q90Z60) Rev-Erb beta (Fragment), ... 43 6e-04
TC223754 similar to UP|Q86EQ4 (Q86EQ4) Clone ZZD1536 mRNA sequen... 42 0.001
CO980191 40 0.007
BE806882 similar to GP|27764548|gb polyprotein {Glycine max}, pa... 39 0.016
TC216231 similar to UP|Q41188 (Q41188) Glycine-rich protein 2 (G... 35 0.18
CO984873 34 0.30
BE804023 33 0.51
CF922488 32 1.1
BM094772 32 1.1
CF922226 32 2.0
>TC234828
Length = 857
Score = 60.5 bits (145), Expect = 4e-09
Identities = 38/85 (44%), Positives = 44/85 (51%)
Frame = +1
Query: 368 CHWRIKL*LTDCRL*MNFQKFFLMRFQMCHQRGRLNFQLTLFQERSRCRWHLTVCRRPS* 427
C W+ L L R NFQKFF M F CH R + N LT R +CR L C R +*
Sbjct: 598 CMWKRTLKLVVYRWCQNFQKFFRMMFVNCHLREKWNSLLTWCLGRIQCRLRLIECLRWN* 777
Query: 428 LS*RNSWRTCLIRNL*NQVFHRGER 452
*R+ +RT NL Q HRGER
Sbjct: 778 QR*RHKYRTF*ANNLFVQAHHRGER 852
Score = 29.3 bits (64), Expect = 9.7
Identities = 16/53 (30%), Positives = 26/53 (48%)
Frame = +3
Query: 187 NFNEEGHISLQCTQPKKVRTGGKVFALTGTQTVNED*LIRGTCFFNIVLL*LL 239
N G++S + + +VFA++G++ D LIRG C LL +L
Sbjct: 66 NSTRSGNVSNNNISGGRPKVPSRVFAMSGSEAAASDDLIRGKCLIADKLLDVL 224
>TC233976
Length = 763
Score = 51.2 bits (121), Expect = 2e-06
Identities = 32/128 (25%), Positives = 54/128 (42%), Gaps = 9/128 (7%)
Frame = +1
Query: 92 EEDTKAHYKVMSERRGKGQL----SRPKPYSAPPDK-----GK*RLKDERRPKMRDAPTD 142
+++ A YK + GK + SR KPYSAP + G+ +
Sbjct: 25 QQEKVAFYKNANASHGKEKKPMTHSRAKPYSAPLEYENHYGGQRTSGGHHLAGGSSQLVN 204
Query: 143 IVCFKCGEKGHKSNVCDRDEKKCFRCGKKGHTLADCKRGDIVCYNFNEEGHISLQCTQPK 202
V G G + +C +CG+ GH +C ++ C+N+ +GH+S C P+
Sbjct: 205 RVSQPAGRGGSGAPAIVTTPLRCRKCGRLGHNAHECTYREVTCFNYQGKGHLSTNCPHPR 384
Query: 203 KVRTGGKV 210
K + G +
Sbjct: 385 KEKRSGSL 408
>TC228935 similar to UP|Q9FHC2 (Q9FHC2) Arabidopsis thaliana genomic DNA,
chromosome 5, TAC clone:K24M7, partial (36%)
Length = 1224
Score = 48.5 bits (114), Expect = 2e-05
Identities = 23/80 (28%), Positives = 38/80 (46%), Gaps = 13/80 (16%)
Frame = +1
Query: 134 PKMRDAPTDIVCFKCGEKGHKSNVCD------RDEKKCFRCGKKGHTLADC----KRGD- 182
P+ + + +C +C +GH++ C +D K C+ CG+ GH L C + G
Sbjct: 313 PEKAEWEKNKICLRCRRRGHRAKNCPEVLDGAKDAKYCYNCGENGHALTQCLHPLQEGGT 492
Query: 183 --IVCYNFNEEGHISLQCTQ 200
C+ N+ GH+S C Q
Sbjct: 493 KFAECFVCNQRGHLSKNCPQ 552
Score = 44.7 bits (104), Expect = 2e-04
Identities = 36/129 (27%), Positives = 47/129 (35%), Gaps = 30/129 (23%)
Frame = +1
Query: 114 PKPYSAPP---DKGK*RLKDERRPKMRDAPTD----------------IVCFKCGEKGHK 154
PKP PP DK K + K+ + R P CF C H
Sbjct: 121 PKPKPTPPKDPDKKKKKNKNSAFKRKRPEPKPGSRKRHLLRVPGMKPGESCFICKAMDHI 300
Query: 155 SNVCDRD-----EKKCFRCGKKGHTLADCK------RGDIVCYNFNEEGHISLQCTQPKK 203
+ +C K C RC ++GH +C + CYN E GH QC P
Sbjct: 301 AKLCPEKAEWEKNKICLRCRRRGHRAKNCPEVLDGAKDAKYCYNCGENGHALTQCLHP-- 474
Query: 204 VRTGGKVFA 212
++ GG FA
Sbjct: 475 LQEGGTKFA 501
>TC227588 similar to PIR|T00837|T00837 glycine-rich protein T13L16.11 -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(10%)
Length = 1300
Score = 47.8 bits (112), Expect = 3e-05
Identities = 29/97 (29%), Positives = 40/97 (40%), Gaps = 11/97 (11%)
Frame = +1
Query: 123 KGK*RLKDERRPKMRDAPTDIVCFKCGEKGHKSNVCDRD-------EKKCFRCGKKGHTL 175
KG R KD + + +C KCG GH C D E +C+ C + GH
Sbjct: 199 KGGHRAKDCPEKHTSTSKSIAICLKCGNSGHDIFSCRNDYSQDDLKEIQCYVCKRLGHLC 378
Query: 176 A----DCKRGDIVCYNFNEEGHISLQCTQPKKVRTGG 208
D G+I CY + GH L C++ + T G
Sbjct: 379 CVNTDDATPGEISCYKCGQLGHTGLACSRLRDEITSG 489
Score = 44.3 bits (103), Expect = 3e-04
Identities = 27/86 (31%), Positives = 41/86 (47%), Gaps = 4/86 (4%)
Frame = +1
Query: 132 RRPKMRDAPTDI--VCFKCGEKGHKSNVCD--RDEKKCFRCGKKGHTLADCKRGDIVCYN 187
R P+ D P + CF CGE+GH + C + +K C+ CG GH C + C+
Sbjct: 16 RGPRYFDPPDNSWGACFNCGEEGHAAVNCSAVKRKKPCYVCGCLGHNARQCSKVQ-DCFI 192
Query: 188 FNEEGHISLQCTQPKKVRTGGKVFAL 213
+ GH + C P+K + K A+
Sbjct: 193 CKKGGHRAKDC--PEKHTSTSKSIAI 264
Score = 43.1 bits (100), Expect = 6e-04
Identities = 19/48 (39%), Positives = 26/48 (53%), Gaps = 9/48 (18%)
Frame = +1
Query: 140 PTDIVCFKCGEKGHKSNVCD--RDE-------KKCFRCGKKGHTLADC 178
P +I C+KCG+ GH C RDE CF+CG++GH +C
Sbjct: 403 PGEISCYKCGQLGHTGLACSRLRDEITSGATPSSCFKCGEEGHFAREC 546
Score = 42.4 bits (98), Expect = 0.001
Identities = 27/85 (31%), Positives = 37/85 (42%), Gaps = 13/85 (15%)
Frame = +1
Query: 132 RRPKMRDAPTDIVCFKCGEKGHKSNVCDRD----EKKCFRCGKKGHTLADCKR--GDIV- 184
R +D +I C+ C GH V D E C++CG+ GHT C R +I
Sbjct: 307 RNDYSQDDLKEIQCYVCKRLGHLCCVNTDDATPGEISCYKCGQLGHTGLACSRLRDEITS 486
Query: 185 ------CYNFNEEGHISLQCTQPKK 203
C+ EEGH + +CT K
Sbjct: 487 GATPSSCFKCGEEGHFARECTSSIK 561
Score = 33.1 bits (74), Expect = 0.67
Identities = 18/48 (37%), Positives = 23/48 (47%)
Frame = +1
Query: 127 RLKDERRPKMRDAPTDIVCFKCGEKGHKSNVCDRDEKKCFRCGKKGHT 174
RL+DE + T CFKCGE+GH + C K R + HT
Sbjct: 463 RLRDE----ITSGATPSSCFKCGEEGHFARECTSSIKSGKRNWESSHT 594
>NP595172 polyprotein [Glycine max]
Length = 4659
Score = 37.4 bits (85), Expect(2) = 3e-05
Identities = 19/38 (50%), Positives = 28/38 (73%)
Frame = +1
Query: 546 CLWSI*TEFSMLDKFVVVFIDDILIYSETEEEHAEHLK 583
CL + +F+ L KFV+VF DDILIYS + ++H +HL+
Sbjct: 2194 CLMNKIFQFA-LRKFVLVFFDDILIYSASWKDHLKHLE 2304
Score = 29.3 bits (64), Expect(2) = 3e-05
Identities = 19/39 (48%), Positives = 24/39 (60%)
Frame = +2
Query: 504 I*GQVTIRLR*RMRICRRRPLGHVLVTMNIR*CLSVLLM 542
I*GQ TI+ +RI R+ L H++ MN *C VLLM
Sbjct: 2060 I*GQDTIKFWYSLRIERKLLLEHIMAIMNG**CHLVLLM 2176
>TC206178 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, partial (69%)
Length = 1138
Score = 46.6 bits (109), Expect = 6e-05
Identities = 30/118 (25%), Positives = 47/118 (39%), Gaps = 2/118 (1%)
Frame = +2
Query: 94 DTKAHYKVMSERRGKGQLSRPKPYSAPPDKGK*RLKDE--RRPKMRDAPTDIVCFKCGEK 151
+ K +K+ S+ R + + P + +D RR R D +C C
Sbjct: 185 ERKKKWKMSSDSRSRSRSRSRSPMDRKIRSDRFSYRDAPYRRDSRRGFSRDNLCKNCKRP 364
Query: 152 GHKSNVCDRDEKKCFRCGKKGHTLADCKRGDIVCYNFNEEGHISLQCTQPKKVRTGGK 209
GH + C + C CG GH ++C + C+N E GH++ C T GK
Sbjct: 365 GHYARECP-NVAICHNCGLPGHIASECTTKSL-CWNCKEPGHMASSCPNEGICHTCGK 532
Score = 46.2 bits (108), Expect = 8e-05
Identities = 34/119 (28%), Positives = 49/119 (40%), Gaps = 29/119 (24%)
Frame = +2
Query: 113 RPKPYSAPPDKGK*R---LKDERRPK--MRDAPTDIVCFKCGEKGHKSNVCDR------- 160
R PY +G R K+ +RP R+ P +C CG GH ++ C
Sbjct: 290 RDAPYRRDSRRGFSRDNLCKNCKRPGHYARECPNVAICHNCGLPGHIASECTTKSLCWNC 469
Query: 161 -----------DEKKCFRCGKKGHTLADCKR-----GDI-VCYNFNEEGHISLQCTQPK 202
+E C CGK GH +C GD+ +C N ++GHI+ +CT K
Sbjct: 470 KEPGHMASSCPNEGICHTCGKAGHRARECSAPPMPPGDLRLCNNCYKQGHIAAECTNEK 646
Score = 45.1 bits (105), Expect = 2e-04
Identities = 20/65 (30%), Positives = 32/65 (48%), Gaps = 6/65 (9%)
Frame = +2
Query: 140 PTDIVCFKCGEKGHKSNVCDR------DEKKCFRCGKKGHTLADCKRGDIVCYNFNEEGH 193
P + +C CG+ GH++ C D + C C K+GH A+C + C N + GH
Sbjct: 500 PNEGICHTCGKAGHRARECSAPPMPPGDLRLCNNCYKQGHIAAEC-TNEKACNNCRKTGH 676
Query: 194 ISLQC 198
++ C
Sbjct: 677 LARDC 691
Score = 45.1 bits (105), Expect = 2e-04
Identities = 26/72 (36%), Positives = 38/72 (52%), Gaps = 3/72 (4%)
Frame = +2
Query: 140 PTDI-VCFKCGEKGHKSNVCDRDEKKCFRCGKKGHTLADCKRGDIVCYNFNEEGHISLQC 198
P D+ +C C ++GH + C +EK C C K GH DC D +C N GH++ QC
Sbjct: 575 PGDLRLCNNCYKQGHIAAECT-NEKACNNCRKTGHLARDCP-NDPICNLCNVSGHVARQC 748
Query: 199 TQPKKV--RTGG 208
+ + R+GG
Sbjct: 749 PKANVLGDRSGG 784
Score = 38.9 bits (89), Expect = 0.012
Identities = 22/87 (25%), Positives = 32/87 (36%), Gaps = 25/87 (28%)
Frame = +2
Query: 137 RDAPTDIVCFKCGEKGHKSNVC-------DRD------------------EKKCFRCGKK 171
RD P D +C C GH + C DR + C C +
Sbjct: 683 RDCPNDPICNLCNVSGHVARQCPKANVLGDRSGGGGGGGGARGGGGGGYRDVVCRNCQQL 862
Query: 172 GHTLADCKRGDIVCYNFNEEGHISLQC 198
GH DC ++C+N GH++ +C
Sbjct: 863 GHMSRDCMGPLMICHNCGGRGHLAYEC 943
Score = 37.0 bits (84), Expect = 0.046
Identities = 15/40 (37%), Positives = 19/40 (47%)
Frame = +2
Query: 142 DIVCFKCGEKGHKSNVCDRDEKKCFRCGKKGHTLADCKRG 181
D+VC C + GH S C C CG +GH +C G
Sbjct: 833 DVVCRNCQQLGHMSRDCMGPLMICHNCGGRGHLAYECPSG 952
>TC227589
Length = 547
Score = 46.2 bits (108), Expect = 8e-05
Identities = 27/89 (30%), Positives = 37/89 (41%), Gaps = 11/89 (12%)
Frame = +2
Query: 123 KGK*RLKDERRPKMRDAPTDIVCFKCGEKGHKSNVCDRD-------EKKCFRCGKKGHTL 175
KG R KD + + +C KCG GH C D E +C+ C + GH
Sbjct: 185 KGGHRAKDCLEKHTSRSKSVAICLKCGNSGHDMFSCRNDYSPDDLKEIQCYVCKRVGHLC 364
Query: 176 A----DCKRGDIVCYNFNEEGHISLQCTQ 200
D G+I CY + GH L C++
Sbjct: 365 CVNTDDATPGEISCYKCGQLGHTGLACSK 451
Score = 43.5 bits (101), Expect = 5e-04
Identities = 24/79 (30%), Positives = 35/79 (43%), Gaps = 4/79 (5%)
Frame = +2
Query: 132 RRPKMRDAPTDI--VCFKCGEKGHKSNVCD--RDEKKCFRCGKKGHTLADCKRGDIVCYN 187
R P+ D P CF CGE GH + C + +K C+ CG GH C + C+
Sbjct: 2 RGPRYFDPPDSSWGACFNCGEDGHAAVNCSAAKRKKPCYVCGGLGHNARQCTKAQ-DCFI 178
Query: 188 FNEEGHISLQCTQPKKVRT 206
+ GH + C + R+
Sbjct: 179 CKKGGHRAKDCLEKHTSRS 235
Score = 39.3 bits (90), Expect = 0.009
Identities = 25/73 (34%), Positives = 33/73 (44%), Gaps = 13/73 (17%)
Frame = +2
Query: 142 DIVCFKCGEKGHKSNVCDRD----EKKCFRCGKKGHT-LADCKRGDIV--------CYNF 188
+I C+ C GH V D E C++CG+ GHT LA K D + C
Sbjct: 323 EIQCYVCKRVGHLCCVNTDDATPGEISCYKCGQLGHTGLACSKLPDEITSAATPSSCCKC 502
Query: 189 NEEGHISLQCTQP 201
E GH + +CT P
Sbjct: 503 GEAGHFAQECTSP 541
Score = 38.1 bits (87), Expect = 0.021
Identities = 17/48 (35%), Positives = 24/48 (49%), Gaps = 9/48 (18%)
Frame = +2
Query: 140 PTDIVCFKCGEKGHKSNVCDR--DE-------KKCFRCGKKGHTLADC 178
P +I C+KCG+ GH C + DE C +CG+ GH +C
Sbjct: 389 PGEISCYKCGQLGHTGLACSKLPDEITSAATPSSCCKCGEAGHFAQEC 532
>TC205022 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, partial (69%)
Length = 1623
Score = 45.4 bits (106), Expect = 1e-04
Identities = 22/46 (47%), Positives = 24/46 (51%), Gaps = 3/46 (6%)
Frame = +2
Query: 164 KCFRCGKKGHTLADCKRGD--IVCYNFNEEGHISLQC-TQPKKVRT 206
+CF CG GH DCK GD CY E GHI C PKK+ T
Sbjct: 311 RCFNCGLDGHWARDCKAGDWKNKCYRCGERGHIERNCKNSPKKLST 448
Score = 40.8 bits (94), Expect = 0.003
Identities = 16/37 (43%), Positives = 22/37 (59%), Gaps = 2/37 (5%)
Frame = +2
Query: 145 CFKCGEKGHKSNVCDRDE--KKCFRCGKKGHTLADCK 179
CF CG GH + C + KC+RCG++GH +CK
Sbjct: 314 CFNCGLDGHWARDCKAGDWKNKCYRCGERGHIERNCK 424
>TC225188 similar to UP|Q9FYB7 (Q9FYB7) Splicing factor-like protein, partial
(62%)
Length = 1131
Score = 45.1 bits (105), Expect = 2e-04
Identities = 22/49 (44%), Positives = 25/49 (50%), Gaps = 3/49 (6%)
Frame = +1
Query: 164 KCFRCGKKGHTLADCKRGD--IVCYNFNEEGHISLQC-TQPKKVRTGGK 209
+CF CG GH DCK GD CY E GHI C PKK+ G+
Sbjct: 361 RCFNCGIDGHWARDCKAGDWKNKCYRCGERGHIEKNCKNSPKKLSQRGR 507
Score = 40.8 bits (94), Expect = 0.003
Identities = 16/37 (43%), Positives = 22/37 (59%), Gaps = 2/37 (5%)
Frame = +1
Query: 145 CFKCGEKGHKSNVCDRDE--KKCFRCGKKGHTLADCK 179
CF CG GH + C + KC+RCG++GH +CK
Sbjct: 364 CFNCGIDGHWARDCKAGDWKNKCYRCGERGHIEKNCK 474
Score = 30.0 bits (66), Expect = 5.7
Identities = 10/27 (37%), Positives = 16/27 (59%)
Frame = +1
Query: 145 CFKCGEKGHKSNVCDRDEKKCFRCGKK 171
C++CGE+GH C KK + G++
Sbjct: 430 CYRCGERGHIEKNCKNSPKKLSQRGRR 510
>TC206180 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, partial (42%)
Length = 655
Score = 43.9 bits (102), Expect = 4e-04
Identities = 23/60 (38%), Positives = 32/60 (53%), Gaps = 1/60 (1%)
Frame = +3
Query: 140 PTDI-VCFKCGEKGHKSNVCDRDEKKCFRCGKKGHTLADCKRGDIVCYNFNEEGHISLQC 198
P D+ +C C ++GH + C +EK C C K GH DC D +C N GH++ QC
Sbjct: 60 PGDLRLCNNCYKQGHIAAECT-NEKACNNCRKTGHLARDCP-NDPICNLCNVSGHVARQC 233
Score = 39.7 bits (91), Expect = 0.007
Identities = 20/84 (23%), Positives = 31/84 (36%), Gaps = 22/84 (26%)
Frame = +3
Query: 137 RDAPTDIVCFKCGEKGHKSNVCDRD----------------------EKKCFRCGKKGHT 174
RD P D +C C GH + C + + C C + GH
Sbjct: 168 RDCPNDPICNLCNVSGHVARQCPKANVLGDXXGGGGGARGGGGGGYRDVVCRNCQQLGHM 347
Query: 175 LADCKRGDIVCYNFNEEGHISLQC 198
DC ++C+N GH++ +C
Sbjct: 348 SRDCMGPLMICHNCGGRGHLAYEC 419
Score = 39.3 bits (90), Expect = 0.009
Identities = 18/57 (31%), Positives = 28/57 (48%), Gaps = 6/57 (10%)
Frame = +3
Query: 148 CGEKGHKSNVCDR------DEKKCFRCGKKGHTLADCKRGDIVCYNFNEEGHISLQC 198
CG+ GH++ C D + C C K+GH A+C + C N + GH++ C
Sbjct: 9 CGKAGHRARECSAPPMPPGDLRLCNNCYKQGHIAAEC-TNEKACNNCRKTGHLARDC 176
Score = 37.0 bits (84), Expect = 0.046
Identities = 15/40 (37%), Positives = 19/40 (47%)
Frame = +3
Query: 142 DIVCFKCGEKGHKSNVCDRDEKKCFRCGKKGHTLADCKRG 181
D+VC C + GH S C C CG +GH +C G
Sbjct: 309 DVVCRNCQQLGHMSRDCMGPLMICHNCGGRGHLAYECPSG 428
Score = 34.3 bits (77), Expect = 0.30
Identities = 16/41 (39%), Positives = 23/41 (56%), Gaps = 6/41 (14%)
Frame = +3
Query: 168 CGKKGHTLADCKR-----GDI-VCYNFNEEGHISLQCTQPK 202
CGK GH +C GD+ +C N ++GHI+ +CT K
Sbjct: 9 CGKAGHRARECSAPPMPPGDLRLCNNCYKQGHIAAECTNEK 131
>TC218711 similar to UP|Q90Z60 (Q90Z60) Rev-Erb beta (Fragment), partial (6%)
Length = 1204
Score = 43.1 bits (100), Expect = 6e-04
Identities = 28/104 (26%), Positives = 41/104 (38%), Gaps = 21/104 (20%)
Frame = -3
Query: 128 LKDERRPKMRDAPTDIVCFKCGEKGHKSNVCDR------DEKKCFRCGKKGHTLADCKRG 181
L+ + P + + C C + GH+ C R DE+ CF CG+ GH+L C
Sbjct: 572 LRVTKSPFKHHGESSLRCRACRQPGHRFQQCQRLKCLSMDEEVCFFCGEIGHSLGKCDVS 393
Query: 182 D---------IVCYNFNEEGHISLQCTQ------PKKVRTGGKV 210
++CY GH S C Q PK + G +
Sbjct: 392 QAGGGRFAKCLLCYG---HGHFSYNCPQNGHGIDPKVLAVNGAI 270
>TC223754 similar to UP|Q86EQ4 (Q86EQ4) Clone ZZD1536 mRNA sequence, partial
(22%)
Length = 742
Score = 42.4 bits (98), Expect = 0.001
Identities = 22/80 (27%), Positives = 31/80 (38%), Gaps = 24/80 (30%)
Frame = +2
Query: 145 CFKCGEKGHKSNVCDRDEKK---------CFRCGKKGHTLADCKR--------------- 180
CF+CG GH + C + CFRCG+ GH DC
Sbjct: 230 CFRCGGFGHMARDCATGKGNIGGGGSGGGCFRCGEVGHLARDCGMEGGRFGGGGGSGGGG 409
Query: 181 GDIVCYNFNEEGHISLQCTQ 200
G C+N + GH + +C +
Sbjct: 410 GKSTCFNCGKPGHFARECVE 469
Score = 41.6 bits (96), Expect = 0.002
Identities = 25/86 (29%), Positives = 32/86 (37%), Gaps = 22/86 (25%)
Frame = +2
Query: 145 CFKCGEKGHKSNVCDRDEKK----------CFRCGKKGHTLADCKR------------GD 182
C++CGE GH + C+R CF CG GH DC R G
Sbjct: 44 CYQCGEFGHLARDCNRSSNSGGGGGGSGGGCFNCGGFGHLARDCVRGGGGSVGIGGGGGG 223
Query: 183 IVCYNFNEEGHISLQCTQPKKVRTGG 208
C+ GH++ C K GG
Sbjct: 224 GSCFRCGGFGHMARDCATGKGNIGGG 301
>CO980191
Length = 802
Score = 39.7 bits (91), Expect = 0.007
Identities = 19/40 (47%), Positives = 24/40 (59%)
Frame = +1
Query: 4 FLNRYFPEDVRGKKEIEFLELKQGDMSVTEYAAKFVELAK 43
FL ++ ED+ K+ EFLELK G+M V Y F EL K
Sbjct: 391 FLEKHLHEDI*NHKKTEFLELKHGNMIVANYLVNFDELPK 510
>BE806882 similar to GP|27764548|gb polyprotein {Glycine max}, partial (3%)
Length = 153
Score = 38.5 bits (88), Expect = 0.016
Identities = 18/30 (60%), Positives = 22/30 (73%)
Frame = -1
Query: 557 LDKFVVVFIDDILIYSETEEEHAEHLK*VF 586
L K+V+VF DDIL+YS T EH HL+ VF
Sbjct: 96 LRKYVLVFFDDILVYSSTWHEHLCHLEVVF 7
>TC216231 similar to UP|Q41188 (Q41188) Glycine-rich protein 2 (GRP2)
(AT4g38680/F20M13_240), partial (42%)
Length = 891
Score = 35.0 bits (79), Expect = 0.18
Identities = 15/47 (31%), Positives = 20/47 (41%), Gaps = 13/47 (27%)
Frame = +3
Query: 145 CFKCGEKGHKSNVCDR-------------DEKKCFRCGKKGHTLADC 178
C+KCGE GH + C + C+ CG+ GH DC
Sbjct: 231 CYKCGETGHIARDCSQGGGGGGRYGGGGGGGGSCYNCGESGHFARDC 371
Score = 34.7 bits (78), Expect = 0.23
Identities = 15/47 (31%), Positives = 20/47 (41%), Gaps = 13/47 (27%)
Frame = +3
Query: 165 CFRCGKKGHTLADCKR-------------GDIVCYNFNEEGHISLQC 198
C++CG+ GH DC + G CYN E GH + C
Sbjct: 231 CYKCGETGHIARDCSQGGGGGGRYGGGGGGGGSCYNCGESGHFARDC 371
>CO984873
Length = 754
Score = 34.3 bits (77), Expect = 0.30
Identities = 16/42 (38%), Positives = 20/42 (47%), Gaps = 11/42 (26%)
Frame = -2
Query: 160 RDEKKCFRCGKKGHTLADCKR-----------GDIVCYNFNE 190
++E KCF C KKGH DC + D VCY N+
Sbjct: 372 KNESKCFFCNKKGHIKKDCSKFKSWLNKKNIPFDFVCYESNK 247
>BE804023
Length = 407
Score = 33.5 bits (75), Expect = 0.51
Identities = 17/60 (28%), Positives = 29/60 (48%), Gaps = 1/60 (1%)
Frame = -3
Query: 145 CFKCGEKGHKSNVC-DRDEKKCFRCGKKGHTLADCKRGDIVCYNFNEEGHISLQCTQPKK 203
C C +KGH C R + KC +C + GH + ++C N N+ + + + PK+
Sbjct: 219 C*HCRKKGHPPFRC*KRPDAKCSQCNQMGHEV-------VICKNRNQSHNEAAKVVDPKE 61
>CF922488
Length = 741
Score = 32.3 bits (72), Expect = 1.1
Identities = 13/31 (41%), Positives = 23/31 (73%)
Frame = +3
Query: 556 MLDKFVVVFIDDILIYSETEEEHAEHLK*VF 586
M+ K + V++DD+++ S TEEEH +L+ +F
Sbjct: 306 MMHKEIEVYMDDMIVKSRTEEEHLVNLRKLF 398
>BM094772
Length = 421
Score = 32.3 bits (72), Expect = 1.1
Identities = 14/36 (38%), Positives = 19/36 (51%), Gaps = 6/36 (16%)
Frame = +2
Query: 145 CFKCGEKGHKSNVCDR------DEKKCFRCGKKGHT 174
C C + GH+ C R DE+ CF CG+ GH+
Sbjct: 314 CRACRQPGHRFQQCQRLKCLSMDEEVCFFCGEIGHS 421
>CF922226
Length = 667
Score = 31.6 bits (70), Expect = 2.0
Identities = 21/84 (25%), Positives = 36/84 (42%), Gaps = 7/84 (8%)
Frame = -3
Query: 87 SCRIYEEDTKAHYKVMSERR-------GKGQLSRPKPYSAPPDKGK*RLKDERRPKMRDA 139
S + E T + K ++ER+ G+G +R K + + K + K E +
Sbjct: 437 SVSLDEVQTALNSKELNERKEKKSSASGEGLTARGKTFKKDSEFDKKKQKPENQKNGEGN 258
Query: 140 PTDIVCFKCGEKGHKSNVCDRDEK 163
I C+ C ++GH VC +K
Sbjct: 257 IFKIRCYHCKKEGHTRKVCPERQK 186
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.369 0.167 0.628
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 51,854,885
Number of Sequences: 63676
Number of extensions: 830917
Number of successful extensions: 10974
Number of sequences better than 10.0: 50
Number of HSP's better than 10.0 without gapping: 4263
Number of HSP's successfully gapped in prelim test: 390
Number of HSP's that attempted gapping in prelim test: 6068
Number of HSP's gapped (non-prelim): 5378
length of query: 1069
length of database: 12,639,632
effective HSP length: 107
effective length of query: 962
effective length of database: 5,826,300
effective search space: 5604900600
effective search space used: 5604900600
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.7 bits)
S2: 64 (29.3 bits)
Medicago: description of AC141111.4


